Week 03 HW: Lab Automation
WEEK 3 — LAB AUTOMATION
LAB PROTOCOL
1. Review of Materials
First, I reviewed the available documentation on HTGAA (LAB–Week 3 – Opentrons Art). Key information required to prepare the design was found at the beginning of the Google Colab notebook, including technical constraints and recommended parameters for droplet spacing and volume.

2. Design Generation
I generated the artistic design using the GUI available at:
https://opentrons-art.rcdonovan.com
Initially, I had questions regarding droplet size and spacing between points. The Google Colab notebook provided specific recommendations about these parameters.
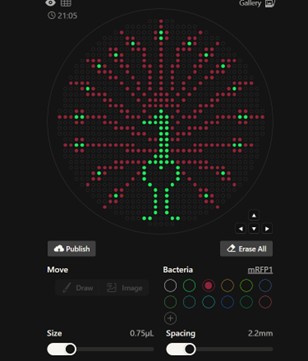
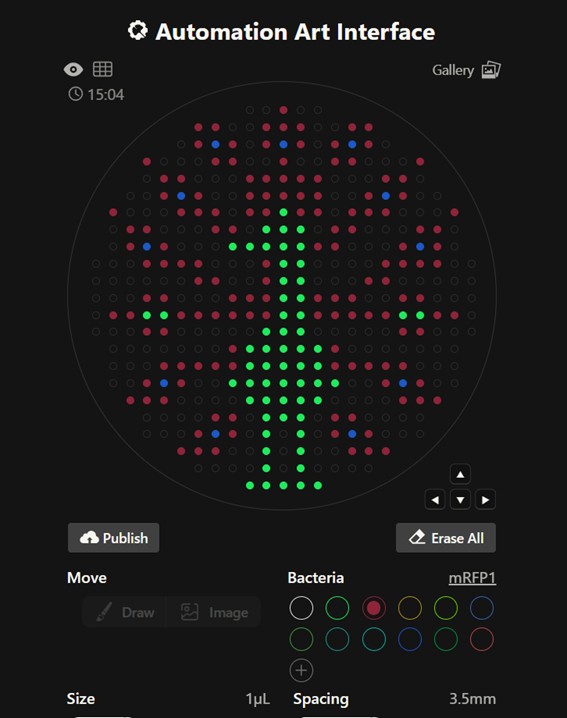
Before identifying these constraints, I created a version of the design with reduced spacing between points to increase visual detail. However, decreasing the distance between droplets increased the risk of unintended merging during robotic dispensing. After reviewing the guidelines, I adjusted the design to comply with the recommended 3.5 mm spacing.
From Donovan’s platform, I downloaded the coordinate sets to be used in the Python script. The coordinates were grouped by color and already respected the recommended spacing (3.5 mm).
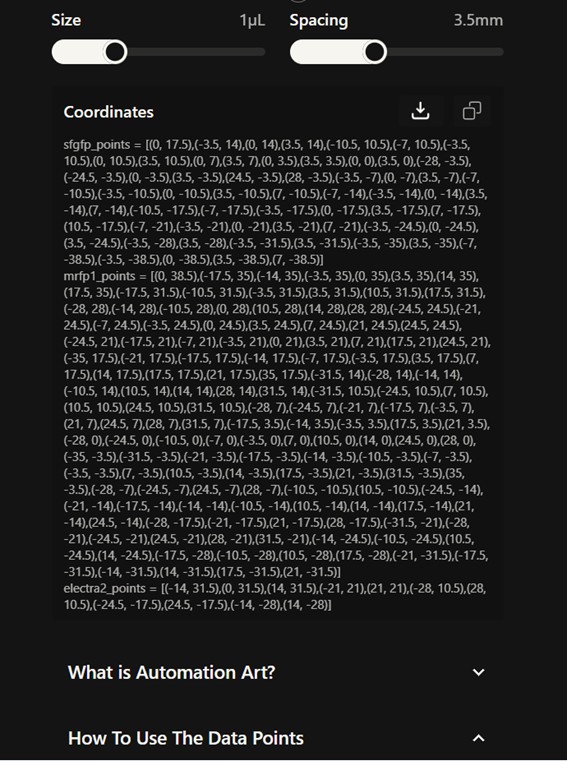

The design was registered on the platform under the following ID:
https://opentrons-art.rcdonovan.com/?id=w8392ofgw0pexpu

Figure Art design
3. Script Development and Simulation
I opened the HTGAA26 Opentrons Colab notebook and created a personal copy to develop the script. The notebook included reference examples from previous students, which were useful to understand how coordinate-based dispensing is implemented on agar plates.
The script was written in Python. Since the laboratory setup provides only two available colors for execution, I adapted the original design as follows:
- Blue droplets were replaced with green droplets using a higher volume (1.2 µL) to create visual differentiation.
- Green and red droplets were dispensed using the recommended volume (1 µL).
After completing the implementation, I executed the simulation within the Opentrons environment. The simulation ran successfully without errors, confirming correct tip usage, aspiration logic, and coordinate positioning.
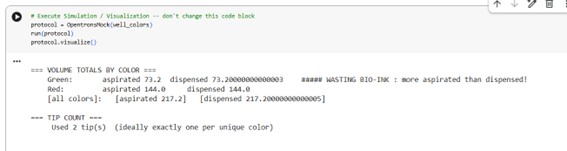

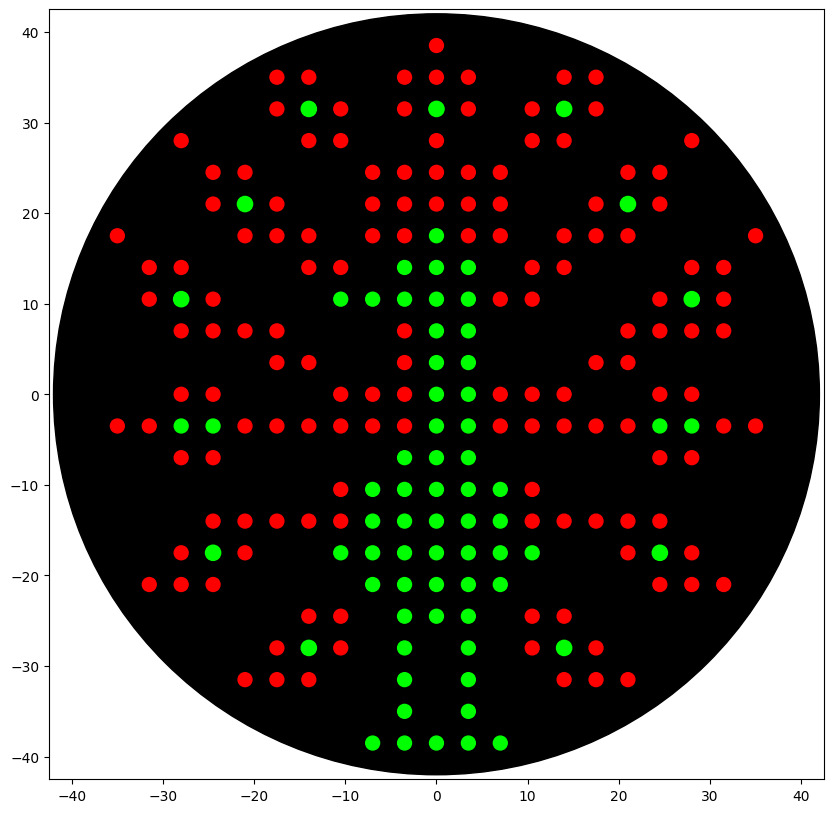
Figure Art design
Here, the script:
4. Use of AI Assistance
Artificial intelligence tools were used to support the development of the script. The structural logic was based on the provided examples in the notebook, particularly Example 7 (Microbial Earth), which clarified how grouped coordinates are iterated and dispensed.
AI assistance was used to:
- Understand how the center of the agar plate is calculated.
- Implement coordinate offset correction for centering the design.
- Structure iteration over grouped coordinate lists while maintaining color consistency.
- Optimize aspiration logic to minimize reagent waste.
The questions posed were specific and focused on resolving implementation details. AI was used as a technical support tool rather than as a replacement for understanding the protocol logic.
5. Robot Scheduling
I already booked Friday 27, 14.00
6. Submission
The Python script was submitted via the corresponding Google Form, and confirmation was received.
The script was shared using the following link:
https://colab.research.google.com/drive/15lHDPfQqFryL9ydvX3P-SeppB7QAd0d8?usp=sharing

Additionally, the metadata section generated the following direct cell link:
https://colab.research.google.com/drive/15lHDPfQqFryL9ydvX3P-SeppB7QAd0d8#scrollTo=t5Bt7VmS_JF_&line=17&uniqifier=1
POST LAB QUESTIONS
1. Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.**
Rosini, E., Battaglia, C., Miani, D., Molinari, F., Arrigoni, F., Piarulli, U., … & Pollegioni, L. (2025). Valuable compounds from pollutants: converting PET into enantiopure alanine. ACS Catalysis, 15(21), 17829-17843.
In this study, the Opentrons OT‑2 automated pipetting system was used to perform a colorimetric assay with PSP dye, which allows measurement of pH changes associated with terephthalic acid (TPA) production during PET depolymerization. The automation enabled rapid and consistent processing of multiple samples, improving the reproducibility and efficiency of depolymerizing enzyme screening. The Opentrons OT‑2 automated system was used to perform a PSP colorimetric assay to measure pH changes during PET depolymerization, enabling fast and reproducible enzyme screening.
2. Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more. You may reference this week’s recitation slide deck for lab automation details.**
No ideas already.
FINAL PROJECT IDEAS
No ideas already.