Week 5 HW: Protein Design Part II
Part 1: Generate Binders with PepMLM
1. Begin by retrieving the human SOD1 sequence from UniProt (P00441) and introducing the A4V mutation.
Superoxide Dismutase 1 (SOD1) is a human enzyme that plays a critical role in protecting cells from oxidative stress by catalyzing the conversion of superoxide radicals into oxygen and hydrogen peroxide. It is a small, 154-amino-acid protein that typically forms a stable homodimer and contains a β-βarrel core structure with metal cofactors, copper and zinc, essential for its catalytic activity. Mutations in SOD1, such as the A4V variant, are associated with familial amyotrophic lateral sclerosis (ALS), a neurodegenerative disorder. SOD1 is widely expressed in the cytoplasm and is a key model protein for studying protein folding, aggregation, and targeted protein degradation strategies.


Mutation A4V was inserted:
Original:
MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQMutated:
MATKVVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
2. Using the PepMLM Colab linked from the HuggingFace PepMLM-650M model card:
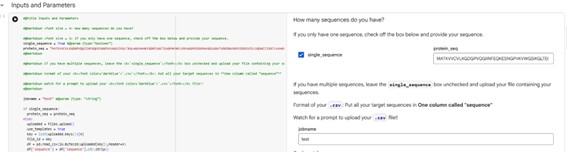

3. Generate four peptides of length 12 amino acids conditioned on the mutant SOD1 sequence.
As the original script generated four peptides containing some invalid characters, I improved the script to produce only valid peptides without those characters. You can access the script cell here:
Additionally, I included a scritp to calculate the pseudo-perplexity of a known peptide.


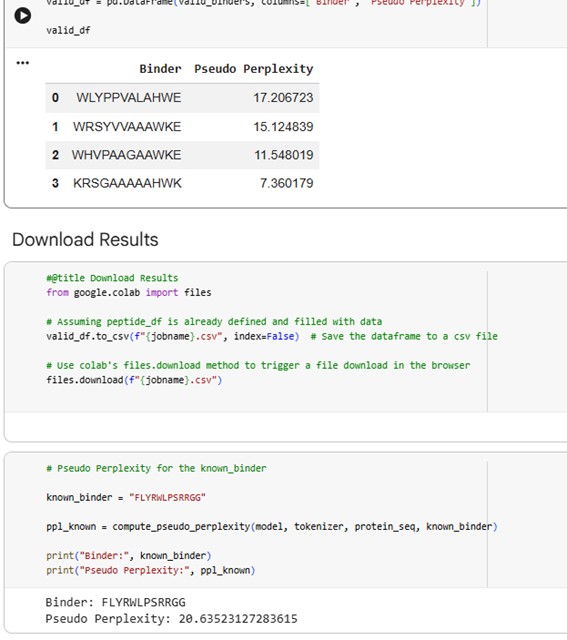

The .cvs file gives the peptides and the Pseudo Perplexity values:
Pseudo-perplexity measures the model’s confidence in a peptide sequence given the target protein context. Lower values indicate that the peptide is more consistent with patterns of protein–peptide interactions learned during training, while higher values suggest lower confidence. In the table, WRSGAAGAAWWK has the lowest pseudo-perplexity (7.12), indicating the model is most confident in this generated peptide, whereas the known binder FLYRWLPSRRGG has the highest value (20.64), reflecting comparatively lower model confidence. The other generated peptides fall in between, showing moderate confidence according to the model.
4. To your generated list, add the known SOD1-binding peptide FLYRWLPSRRGG for comparison.
- I open the test.csv file in excel, added a line and insert the SOD1-binding peptide FLYRWLPSRRGG. Record the perplexity scores that indicate PepMLM’s confidence in the binders.
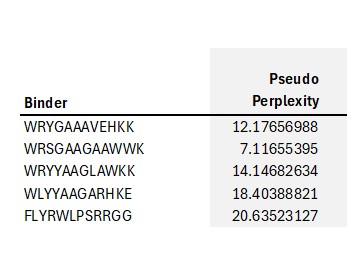

PepMLM assigns lower pseudo-perplexity scores to several generated peptides compared with the known SOD1-binding peptide FLYRWLPSRRGG. The peptide WRSGAAGAAWWK shows the lowest perplexity (7.12), indicating that the model predicts it as the most probable binder among the candidates.
Part 2: Evaluate Binders with AlphaFold3
1. Navigate to the AlphaFold Server: alphafoldserver.com
2. For each peptide, submit the mutant SOD1 sequence followed by the peptide sequence as separate chains to model the protein-peptide complex.


3. Record the ipTM score and briefly describe where the peptide appears to bind. Does it localize near the N-terminus where A4V sits? Does it engage the β-barrel region or approach the dimer interface? Does it appear surface-bound or partially buried?
The predicted binders 1–4 localize near residues 96–102 of the protein, a region that forms one of the parallel β-strands contributing to the β-sheet structure. In contrast, the known binder (binder 5) appears near residues 11–15. None of the binders are located close to the N-terminus or C-terminus of the protein. Additionally, the peptides do not appear tightly attached to the protein surface or inserted into a binding pocket, suggesting that they remain relatively exposed rather than strongly surface-bound or buried.
The predicted interaction confidence, measured by ipTM, is relatively low for all binders, with values ranging from 0.25 to 0.44. Binder 2 shows the highest interaction score (ipTM = 0.44), followed by binder 4 (0.39) and binder 1 (0.34), while binders 3 (0.28) and 5 (0.25) show lower confidence. These values indicate moderate to low confidence in the predicted peptide–protein interactions, suggesting that the binding poses may be weak or uncertain.
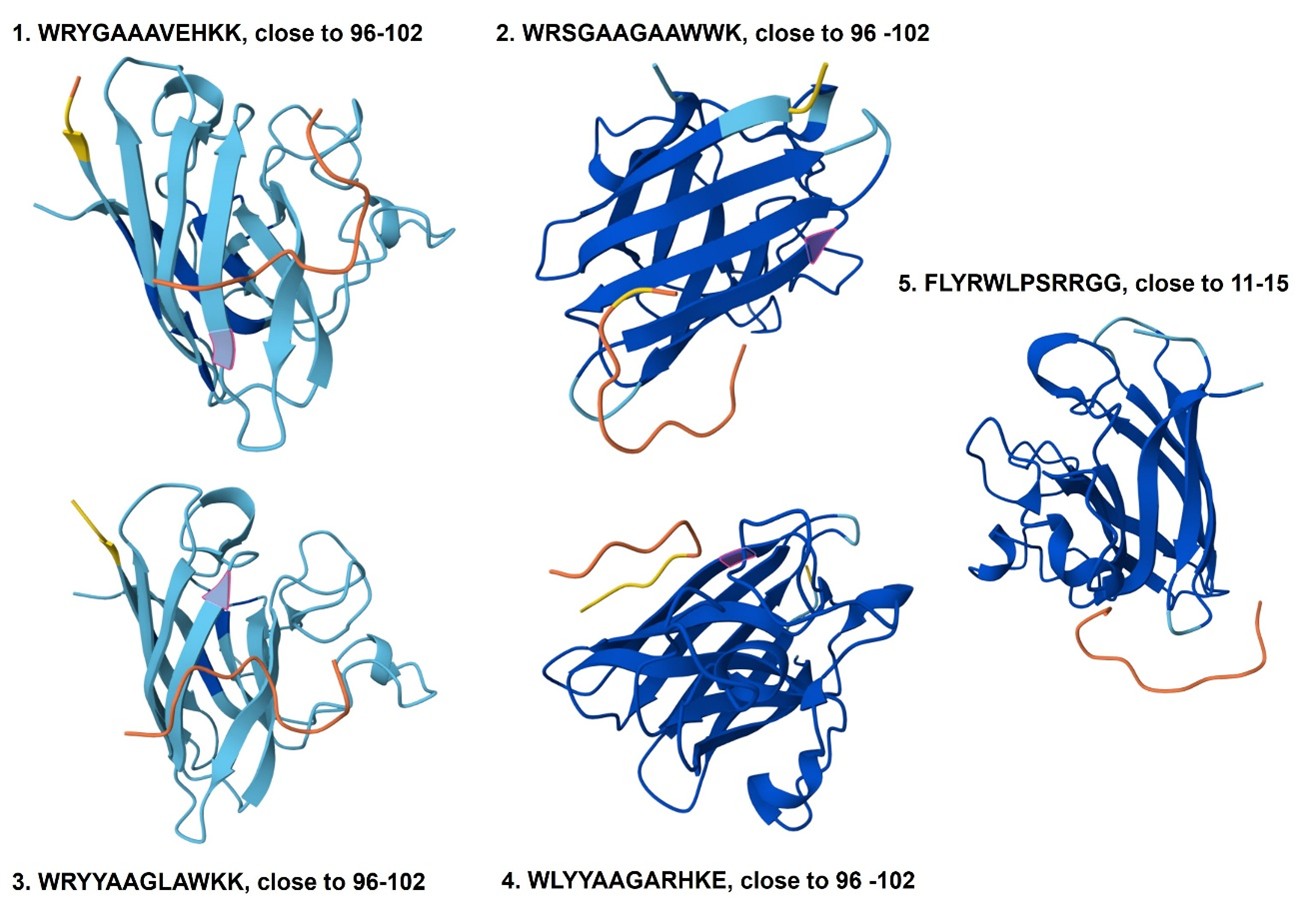

4. In a short paragraph, describe the ipTM values you observe and whether any PepMLM-generated peptide matches or exceeds the known binder.
The predicted complexes show low ipTM values (0.25–0.44), indicating weak confidence in protein–peptide interactions. The PepMLM-generated peptide WRSGAAGAAWWK has the highest ipTM score (0.44), exceeding that of the known binder FLYRWLPSRRGG (0.25), but none of the peptides appear to form a stable interaction with SOD1 in the predicted models.
Part 3: Evaluate Properties of Generated Peptides in the PeptiVerse
Structural confidence alone is insufficient for therapeutic development. Using PeptiVerse, let’s evaluate the therapeutic properties of your peptide! For each PepMLM-generated peptide:
1. Paste the peptide sequence.
2. Paste the A4V mutant SOD1 sequence in the target field.
- Check the boxes:
- Predicted binding affinity
- Solubility
- Hemolysis probability
- Net charge (pH 7)
- Molecular weight
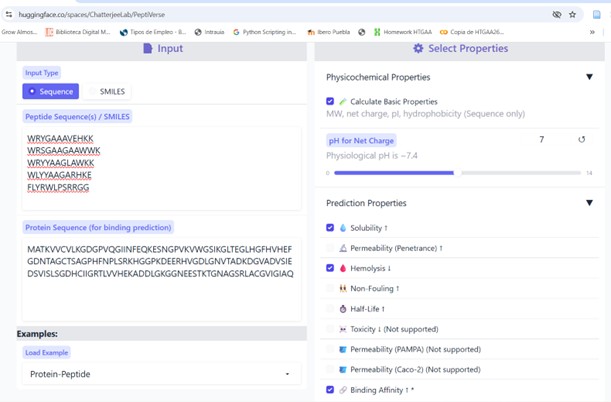

Compare these predictions to what you observed structurally with AlphaFold3. In a short paragraph, describe what you see. Do peptides with higher ipTM also show stronger predicted affinity? Are any strong binders predicted to be hemolytic or poorly soluble? Which peptide best balances predicted binding and therapeutic properties?
Choose one peptide you would advance and justify your decision briefly.
The predicted binding affinities range from pKd/pKi values of ~5.4 to 6.9, indicating weak to moderate binding overall. The peptide WRSGAAGAAWWK shows both the highest predicted affinity (pKd 6.94) and the highest ipTM value (0.44), suggesting some agreement between the structural prediction from AlphaFold3 and the affinity prediction from Peptiverse. However, most peptides did not appear to form stable interactions with SOD1 in the structural models, remaining mostly surface-proximal rather than clearly bound. All peptides show good predicted solubility (score = 1) and very low hemolysis probabilities (<0.05), indicating favorable therapeutic properties. Among the candidates, WRSGAAGAAWWK (2) best balances predicted binding strength, structural confidence, and safety properties.
See all values from alphaFold web y pepTiVerse per peptide:
| Binder | Pseudo Perplexity | ipTM | pTM | Solubility | Hemolysis | Binding Affinity (pKd/pKi) | Length (aa) | Molecular Weight (Da) | Net Charge (pH 7) | Isoelectric Point (pH) | Hydrophobicity (GRAVY) |
|---|---|---|---|---|---|---|---|---|---|---|---|
| WRYGAAAVEHKK | 12.17656988 | 0.34 | 0.75 | 1 | 0.021 | 5.436 | 12 | 1415.6 | 1.85 | 9.7 | -1 |
| WRSGAAGAAWWK | 7.11655395 | 0.44 | 0.84 | 1 | 0.032 | 6.939 | 12 | 1346.5 | 1.76 | 11 | -0.46 |
| WRYYAAGLAWKK | 14.14682634 | 0.28 | 0.73 | 1 | 0.024 | 6.735 | 12 | 1512.8 | 2.76 | 10 | -0.66 |
| WLYYAAGARHKE | 18.40388821 | 0.39 | 0.87 | 1 | 0.029 | 5.887 | 12 | 1464.6 | 0.85 | 8.5 | -0.82 |
| FLYRWLPSRRGG | 20.63523127 | 0.25 | 0.83 | 1 | 0.047 | 5.968 | 12 | 1507.7 | 2.76 | 11.71 | -0.71 |
Notes:
- ipTM score estimates the confidence of protein–peptide interactions. Values above 0.7 indicate a reliable interaction, values between 0.5 and 0.7 suggest a possible interaction, and values below 0.5 indicate that an interaction is unlikely.
- Solubility is predicted as a binary value where 0 indicates not soluble and 1 indicates soluble.
- Hemolysis probability predicts whether a peptide may damage red blood cells; 0 indicates non-hemolytic and 1 indicates hemolytic.
- Binding affinity (pKd/pKi) reflects binding strength (−log10 of Kd or Ki), where higher values indicate stronger binding. Values <5 indicate very weak binding (>100 µM), 5–6 weak (~100 µM), 6–7 moderate (~10 µM), 7–8 good (~1 µM), and >8 strong binding (<100 nM).
Part 4: Generate Optimized Peptides with moPPIt
Now, move from sampling to controlled design. moPPIt uses Multi-Objective Guided Discrete Flow Matching (MOG-DFM) to steer peptide generation toward specific residues and optimize binding and therapeutic properties simultaneously. Unlike PepMLM, which samples plausible binders conditioned on just the target sequence, moPPIt lets you choose where you want to bind and optimize multiple objectives at once.
1. Open the moPPit Colab linked from the HuggingFace moPPIt model card
2. Make a copy and switch to a GPU runtime.

3. In the notebook:
- Paste your A4V mutant SOD1 sequence.
- Choose specific residue indices on SOD1 that you want your peptide to bind (for example, residues near position 4, the dimer interface, or another surface patch).
- Set peptide length to 12 amino acids.
- Enable motif and affinity guidance (and solubility/hemolysis guidance if available). Generate peptides.
4. After generation, briefly describe how these moPPit peptides differ from your PepMLM peptides. How would you evaluate these peptides before advancing them to clinical studies?
moPPIt Generated Peptides
| Binder | Hemolysis | Solubility | Affinity | Motif |
|---|---|---|---|---|
| GGGRQKCFTLNM | 0.9349 | 0.75 | 6.5764 | 0.5096 |
| SKKQKTITCELC | 0.9677 | 0.8333 | 7.2126 | 0.6561 |
| KTCKKSFEKKQN | 0.9739 | 0.9167 | 6.2008 | 0.6707 |
| GTTCIQGKKKDE | 0.9785 | 0.9167 | 6.3731 | 0.5156 |
The peptides generated with moPPIt differ from the PepMLM peptides because their generation was guided by multiple objectives defined in the notebook, including affinity, motif targeting, solubility, hemolysis, and specificity. In contrast, PepMLM generates peptides based mainly on sequence likelihood conditioned on the target protein sequence. As a result, the moPPIt peptides show moderate predicted affinities (pKd ≈ 6.2–7.2), relatively high motif scores (≈0.51–0.67), and good solubility predictions (0.75–0.92). The PepMLM peptides also show moderate affinities but were not optimized simultaneously for these therapeutic objectives during generation.
However, when one of the generated peptides was modeled with the SOD1 structure using AlphaFold, the peptide did not appear to form a stable interaction with the protein surface. This suggests that although the predicted affinity and motif scores are moderate, the structural predictions do not clearly support strong binding.
Comparison of PepMLM and moPPIt Generated Peptides
| Method | Binder | Hemolysis | Solubility | Affinity |
|---|---|---|---|---|
| PepMLM | WRYGAAAVEHKK | 0.021 | Yes | 5.436 |
| PepMLM | WRSGAAGAAWWK | 0.032 | Yes | 6.939 |
| PepMLM | WRYYAAGLAWKK | 0.024 | Yes | 6.735 |
| PepMLM | WLYYAAGARHKE | 0.029 | Yes | 5.887 |
| PepMLM | FLYRWLPSRRGG | 0.047 | Yes | 5.968 |
| moPPIt | GGGRQKCFTLNM | 0.9349 | 0.75 | 6.5764 |
| moPPIt | SKKQKTITCELC | 0.9677 | 0.8333 | 7.2126 |
| moPPIt | KTCKKSFEKKQN | 0.9739 | 0.9167 | 6.2008 |
| moPPIt | GTTCIQGKKKDE | 0.9785 | 0.9167 | 6.3731 |
Before advancing any peptide toward clinical studies, further validation would be necessary. Additional structural modeling and docking analyses should confirm stable binding to the intended region of SOD1. Promising candidates should then be synthesized and tested experimentally through in vitro binding assays, hemolysis and cytotoxicity tests, and stability analyses to verify their predicted properties. Only peptides showing clear binding and favorable safety profiles would be considered for further development.
PENDING PENDING PENDING PENDINT
Part C: Final Project: L-Protein Mutants
High level summary: The objective of this assignment is to improve the stability and auto-folding of the lysis protein of a MS2-phage. This mechanism is key to the understanding of how phages can potentially solve antibiotic-resistance.
This homework requires computation that might take you a while to run, so please get started early.