Week 6 HW: Genetic Circuits: Part I
Answer these questions about the protocol in this week’s lab:
1. What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
Components of the Phusion High-Fidelity PCR Master Mix and their purpose:
- Phusion DNA polymerase – a high-fidelity DNA polymerase that synthesizes new DNA strands with a low error rate during PCR.
- Primers (forward and reverse) – short DNA sequences that bind to the target DNA and define the region that will be amplified.
- dNTPs (dATP, dTTP, dCTP, dGTP) – the nucleotide building blocks used by the polymerase to synthesize new DNA strands.
- Reaction buffer – maintains optimal pH and ionic conditions for proper enzyme activity.
- Mg²⁺ ions – an essential cofactor required for DNA polymerase catalytic activity.
- Nuclease-free water – maintains the correct reaction volume and prevents degradation of DNA.
2. What are some factors that determine primer annealing temperature during PCR?
- Primer melting temperature (Tm) – The annealing temperature is usually set about 3–5 °C below the primer Tm to allow specific binding to the DNA template.
- Primer length – Longer primers generally have higher Tm values, which increases the annealing temperature.
- GC content – Primers with higher GC content bind more strongly (three hydrogen bonds), increasing Tm and the annealing temperature.
- Primer–template complementarity – Mismatches between the primer and the template reduce binding efficiency and may require a lower annealing temperature.
- Reaction conditions – Salt concentration and Mg²⁺ levels in the PCR mix influence DNA stability and can affect the optimal annealing temperature.
3. There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
Comparison of PCR and Restriction Enzyme Digestion
| Criterion | PCR (Polymerase Chain Reaction) | Restriction Enzyme Digestion |
|---|---|---|
| Underlying mechanism | DNA amplification using primers and a thermostable DNA polymerase through repeated thermal cycles. | Enzymatic cleavage of DNA at specific recognition sequences by restriction endonucleases. |
| Key reagents | DNA template, forward and reverse primers, DNA polymerase (e.g., Taq or Phusion), dNTPs, buffer. | Restriction enzyme(s), compatible reaction buffer, and DNA substrate. |
| Sequence constraints | Only requires primer binding regions; primers can be designed to introduce mutations or overlaps. | Requires pre-existing restriction sites within the DNA sequence. |
| Experimental flexibility | Highly adaptable; enables mutagenesis, sequence insertion, and creation of overlaps for cloning strategies such as Gibson Assembly. | Limited flexibility; modification depends on the presence and position of restriction sites. |
| Resulting DNA product | Defined amplified DNA fragment. | DNA fragments generated by site-specific cleavage. |
| Typical applications | Gene amplification, site-directed mutagenesis, preparation of fragments for advanced cloning methods. | Traditional cloning workflows where compatible restriction sites are available. |
| When it is preferred | When sequence modification or precise amplification is required. | When simple and reliable DNA cutting is sufficient for cloning. |
4. How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
- Design 20–40 bp complementary overhangs between adjacent DNA fragments.
- Include the correct 5′ overhangs in PCR primers so fragments share matching overlaps.
- Ensure fragments are in the correct 5′→3′ orientation for proper assembly.
- Avoid secondary structures (e.g., hairpins or primer dimers) in the overlap regions.
5. How does the plasmid DNA enter the E. coli cells during transformation?
- Heat shock: A sudden temperature change creates temporary pores in the membrane, allowing plasmid DNA to enter.
- Electroporation: An electrical pulse creates pores in the membrane through which plasmid DNA enters the cell.
6. Describe another assembly method in detail (such as Golden Gate Assembly). Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
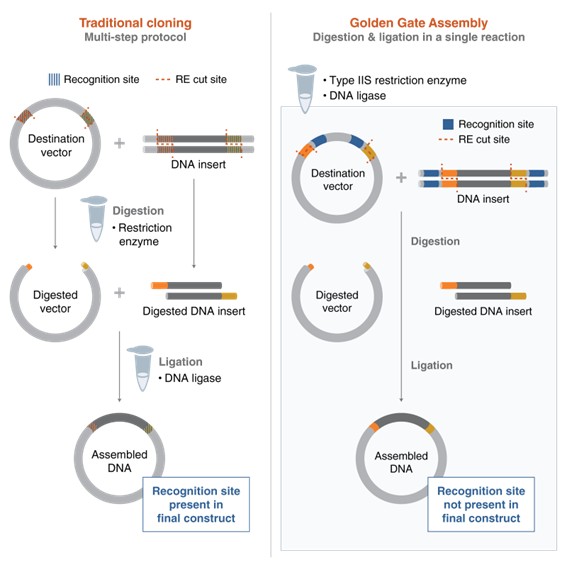
Golden Gate Assembly is a molecular cloning method that allows the assembly of multiple DNA fragments in a single reaction. It uses Type IIS restriction enzymes (such as BsaI) together with DNA ligase. These enzymes cut DNA outside of their recognition site, creating specific overhangs that can be designed to match between fragments. The complementary overhangs allow the DNA fragments to anneal in a predefined order. DNA ligase then joins the fragments together to form a continuous DNA molecule. Because the restriction sites are removed during the process, the final assembled DNA can no longer be cut again, making the reaction efficient and directional. This method allows the simultaneous and seamless assembly of multiple fragments in one tube.

DNA Fragment A [ATGCGTACGTTAGCTAGCTAGCGATCGATCGTAGCTAGCTAGCTA]
DNA Fragment B [AGCTATTTGCGGATCGATCA]
DNA Fragment C [ATCAGGCTAGCGTATCGTAA]
Golden Gate Assembly, Conceptual Diagram
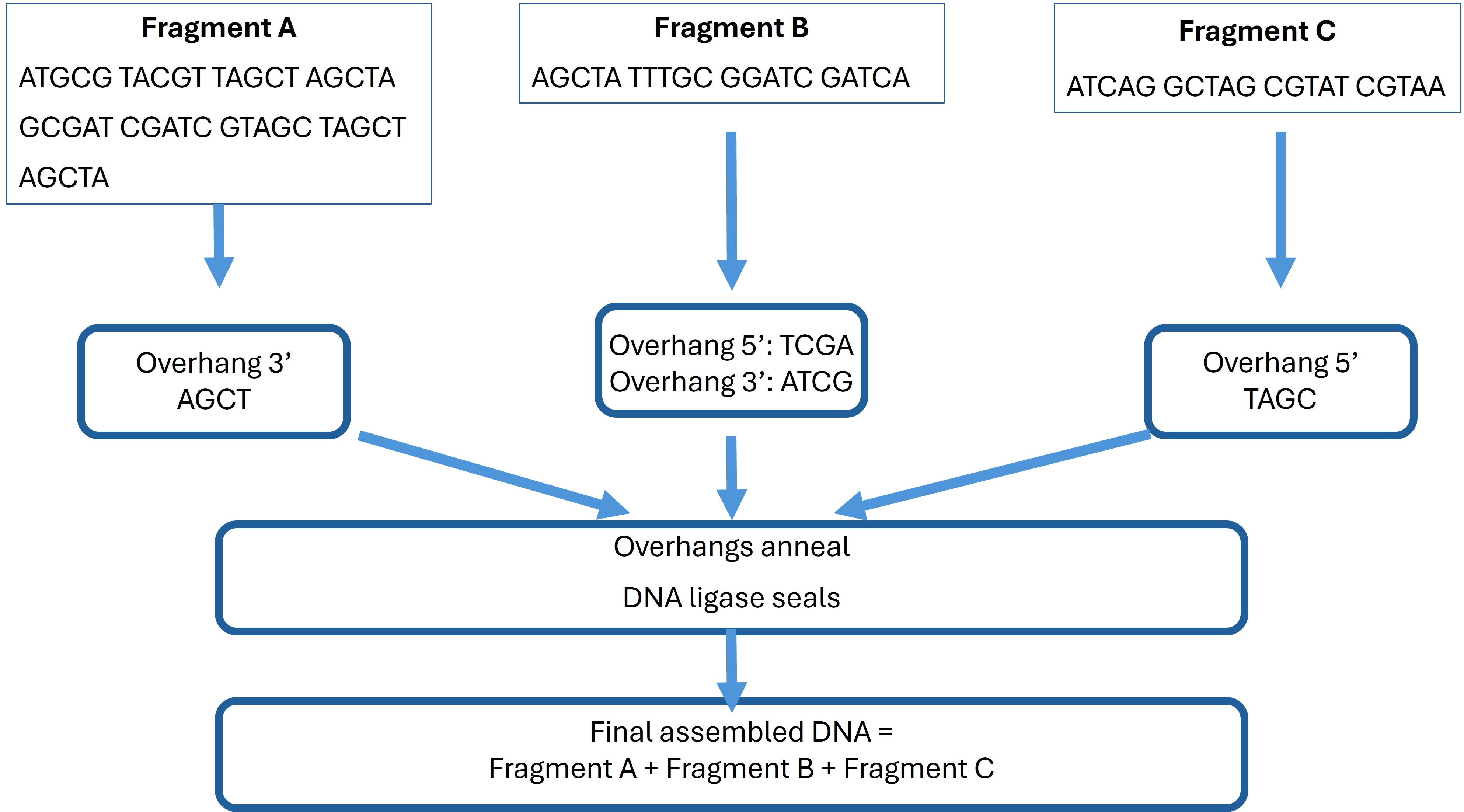 Figure 1. pUC19 backbone |
These overhangs ensure correct directional assembly of fragments A, B, and C. The sequences have been prepared and modified to include complementary overhangs for accurate Golden Gate assembly simulation in Benchling.
Golden Gate Assembly Overhangs
Fragment A → Fragment B
| Fragment A (5’→3’) | Fragment B (5’→3’) | Overhang (A → B, antiparallel) |
|---|---|---|
| ATGCGTACGTTAGCTAGCTAGCGATCGATCGTAGCTAGCTAGCTA | AGCTATTTGCGGATCGATCA | TAGC → GCTA |
Fragment B → Fragment C
| Fragment B (5’→3’) | Fragment C (5’→3’) | Overhang (B → C, antiparallel) |
|---|---|---|
| AGCTATTTGCGGATCGATCA | ATCAGGCTAGCGTATCGTAA | CGAT → GCTA |
Overhangs’ summary
| Fragment | Overhang 5’ | Overhang 3’ |
|---|---|---|
| A | — | AGCT |
| B | TCGA | ATCG |
| C | TAGC | — |
Screen shots from Benchling
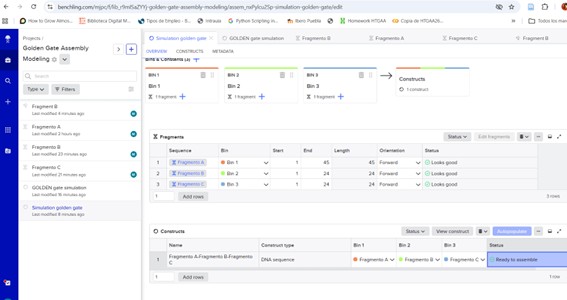 Figure 1. pUC19 backbone | 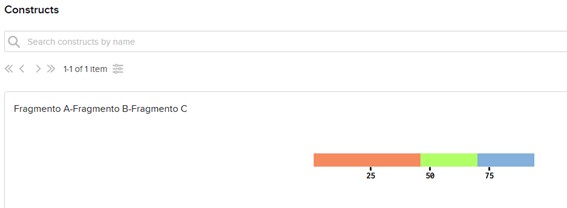 Figure 2. Recombinant plasmid |
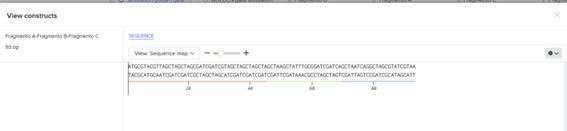
Figure 3. Final construct
Assignment: Asimov Kernel
1.Setup: Create a Repository and a blank Notebook entry to document the homework and save it to that Repository


2. Explore the devices in the Bacterial Demos Repo to understand how the parts work together by running the Simulator on various examples.
Demo - Represssilator
a. Construct
The repressilator construct consists of three genes (LacI, LambdaCI, and TetR) arranged in a cyclic inhibitory network. Each gene is placed under the control of a promoter that is repressed by the previous gene in the cycle. Specifically, LacI represses LambdaCI, LambdaCI represses TetR, and TetR represses LacI.
This design creates a closed loop of negative regulation, where each protein periodically inhibits the next, preventing the system from reaching a stable steady state.
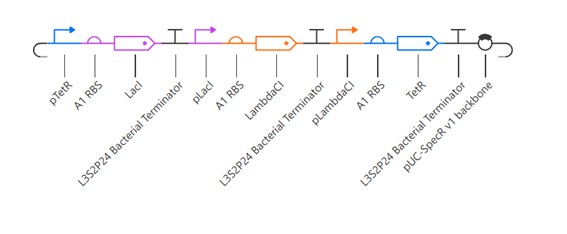

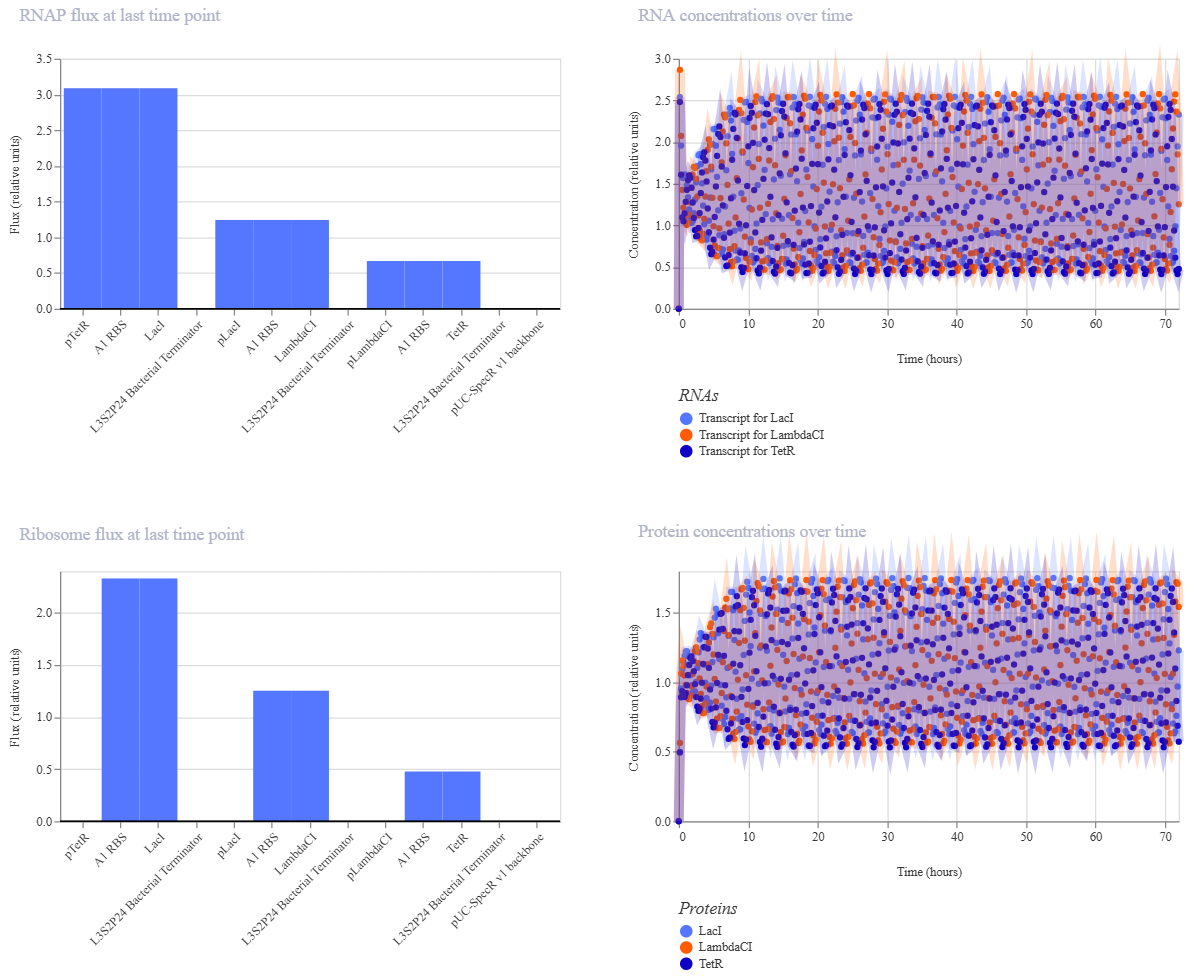
The RNAP flux graph shows the transcriptional activity of each gene at the final time point, indicating how strongly RNA polymerase is initiating transcription for each promoter. Differences in flux reflect variations in promoter activity within the circuit.
The RNA concentrations over time graph displays oscillatory behavior for all three genes (LacI, LambdaCI, and TetR), demonstrating periodic transcription dynamics characteristic of a repressilator system.
The ribosome flux graph represents the translation activity at the final time point, showing how actively ribosomes are producing proteins from each mRNA. Variations indicate differences in translation efficiency.
The protein concentrations over time graph shows clear oscillations in the levels of LacI, LambdaCI, and TetR. These periodic fluctuations confirm the cyclic repression mechanism, where each protein represses the next in sequence, generating sustained oscillatory behavior.
Demo - Comparing Promoters
a. Construct
This construct is designed to compare the strength of different promoters. It consists of three transcriptional units, each containing a promoter, a ribosome binding site (RBS), a reporter gene, and a terminator.
The promoters used (BBa_J23101, BBa_J23106, and BBa_J23117) have different strengths, allowing direct comparison of their effect on gene expression. Since all other components are identical, any differences in expression levels can be attributed to the promoter strength.
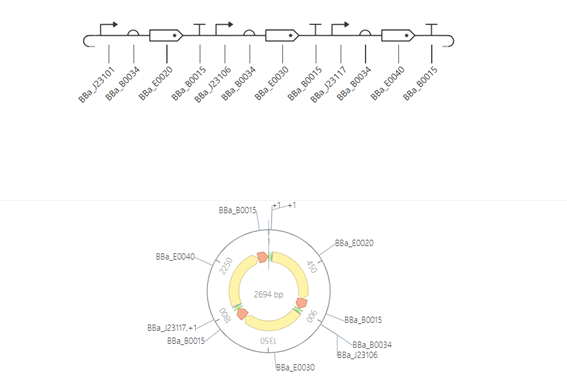

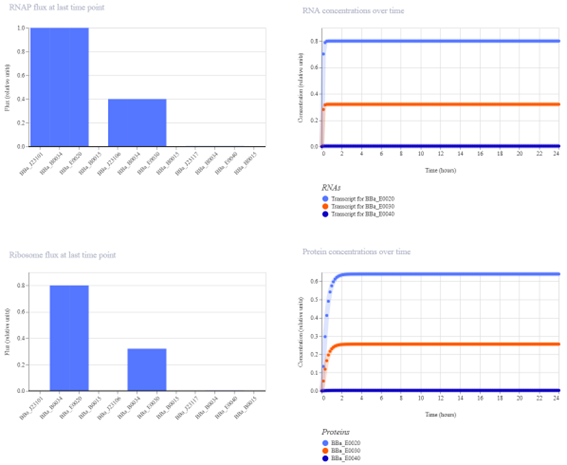
The protein concentration graph shows three horizontal lines at different levels, each corresponding to a different gene. These steady expression levels reflect the strength of the promoters controlling each gene.
The strongest promoter produces the highest protein concentration, while the weakest promoter results in the lowest expression. Since there are no regulatory interactions between the genes, the system reaches a stable steady state without oscillations.
Demo - Multiplexer/ H1 RBS
a. Construct
The multiplexer circuit consists of multiple input signals that activate different regulatory proteins. These proteins act as repressors and interact to control the expression of the output gene (GFP).
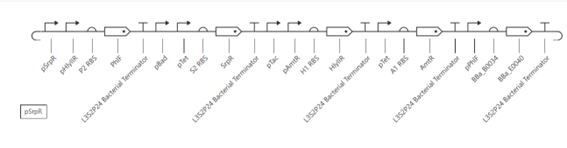

Each input activates a specific promoter, leading to the production of regulatory proteins. These proteins then inhibit or allow the expression of the output depending on their combination. As a result, the system produces GFP only under specific input conditions, demonstrating combinatorial control of gene expression.
| Stage | Element | Action | Target |
|---|---|---|---|
| Input | L-arabinose | activates | A |
| Input | aTc | activates | B |
| Input | IPTG | activates | C |
| Regulation | A | represses | D |
| Regulation | B | represses | E |
| Regulation | C | represses | F |
| Output | D, E, F | repress | GFP |
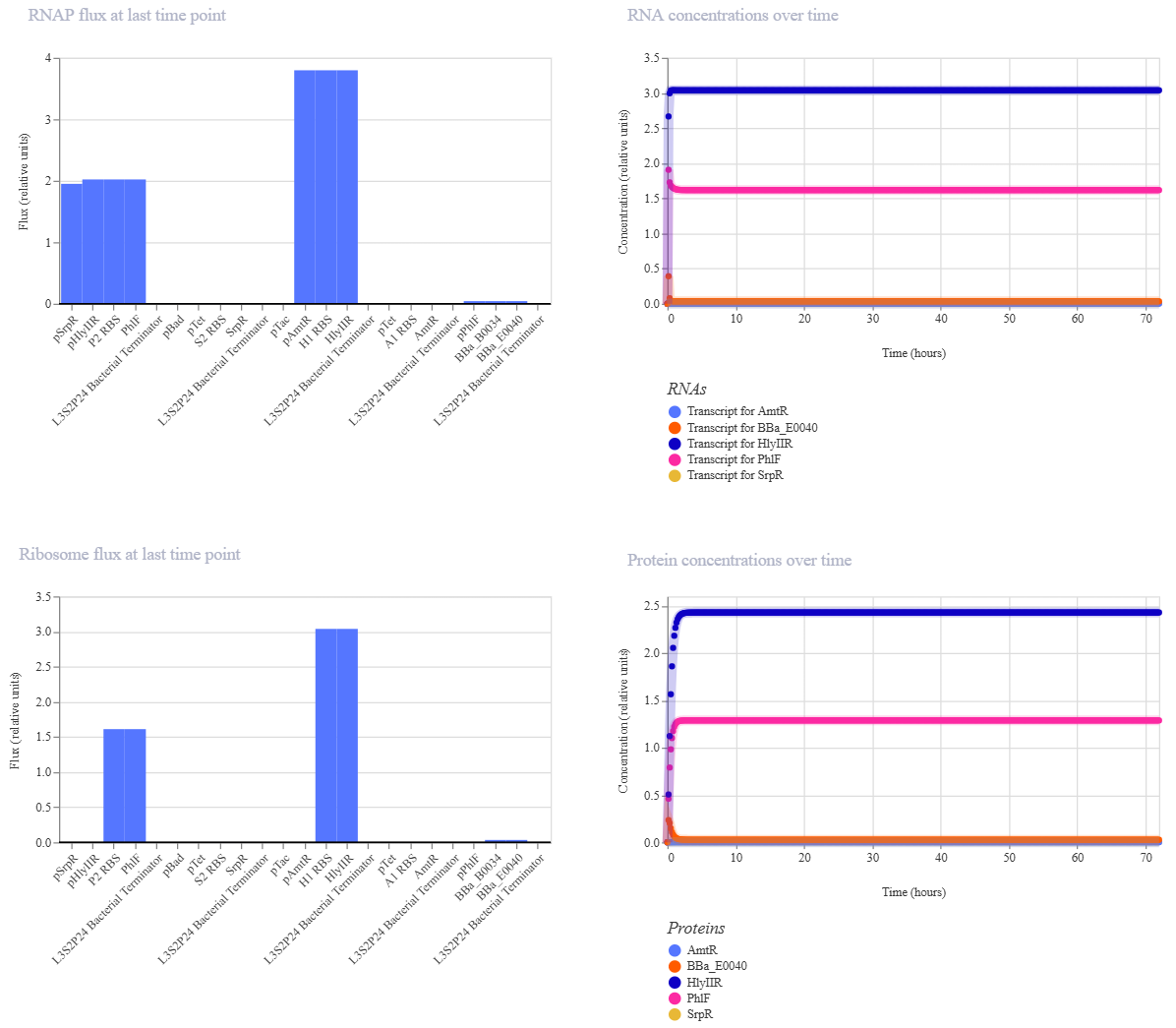
The simulation results show that certain regulatory proteins, particularly AmtR, reach high expression levels, while others remain low. The system stabilizes without oscillations, indicating a steady-state behavior.
The protein concentration graph shows that GFP (BBa_E0040) remains at very low levels, meaning that the output is effectively repressed under these conditions. This suggests that the circuit selects a regulatory pathway that inhibits GFP expression.
Demo - AND
a. Construct
This construct implements an AND logic gate using a cascade of repressors. The inputs aTc and IPTG activate the production of regulatory proteins, which together repress an intermediate repressor. As a result, the output gene (GFP) is expressed only when both inputs are present, demonstrating conditional gene expression based on multiple signals.
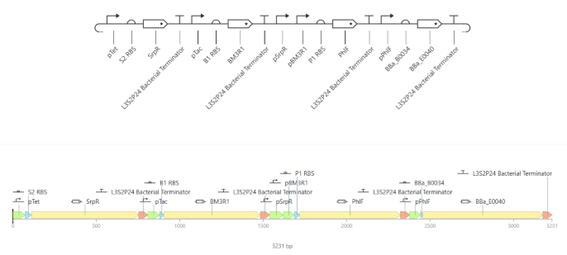

This construct contains three transcription units, each controlled by promoters of different strengths: strong, medium, and weak. Each promoter drives the expression of a different reporter gene.
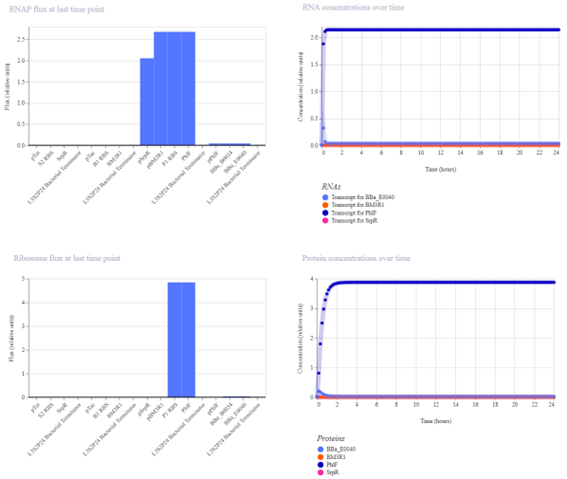
The simulation results show that only one protein reaches a high concentration, corresponding to the output gene GFP. The other proteins remain at low levels.
The protein concentration graph reflects how promoter strength affects gene expression. Strong promoters result in higher protein levels, while weaker promoters produce lower levels of expression.
This indicates that the circuit is functioning correctly as an AND gate, where GFP is expressed only under the appropriate input conditions. The system reaches a stable steady state with clear differentiation between active and inactive components.
4. Build the Repressilator: Create a blank Construct and save it to your Repository
- Recreate the Repressilator in that empty Construct using parts from the Characterized Bacterial Parts repository
- Search the parts using the Search function in the right menu
- Drag and drop the parts into the Construct
- Confirm it works by running the Simulator (“play” button)
- Compare your results with the Repressilator Construct in the Bacterial Demos repository
A repressilator is a synthetic genetic circuit composed of three genes that inhibit each other in a cyclic manner (A represses B, B represses C, and C represses A).
This feedback loop creates oscillations in gene expression over time, meaning that the levels of each protein rise and fall periodically instead of reaching a stable state. It is a classic example of a biological oscillator in synthetic biology.
a. Construct
The repressilator construct was built by assembling three transcriptional units using parts from the characterized bacterial parts repository. Each unit consists of a promoter, a ribosome binding site (RBS), a coding sequence (CDS), and a terminator.
The genes used in this construct are LacI, LambdaCI, and TetR, arranged sequentially in the construct.
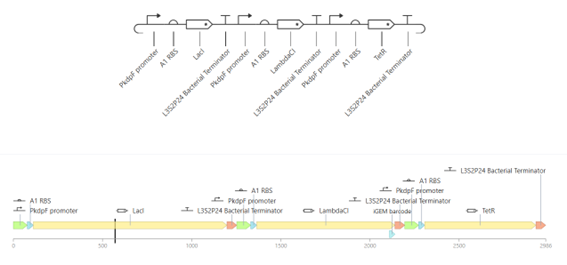

b. Expected Behavior
The repressilator is expected to exhibit oscillatory behavior in gene expression. Due to the cyclic inhibitory interactions between the three regulatory proteins, the concentration of each protein should rise and fall over time in a periodic manner.
Each gene is expressed in sequence: when one protein reaches a high level, it represses the next gene in the loop, causing its concentration to decrease, while allowing another gene to become active. This results in a continuous cycle of expression.
As a result, both RNA and protein concentrations should display sustained oscillations rather than reaching a steady state.
c. Simulation
The construct was simulated using the built-in simulator with E. coli as the chassis, a duration of 24 hours, and a timestep of 30 minutes (data for all simulations constructed).
The simulation results show that the protein concentrations increase initially and then reach a steady state without oscillations. This indicates that the system does not behave as a functional repressilator.
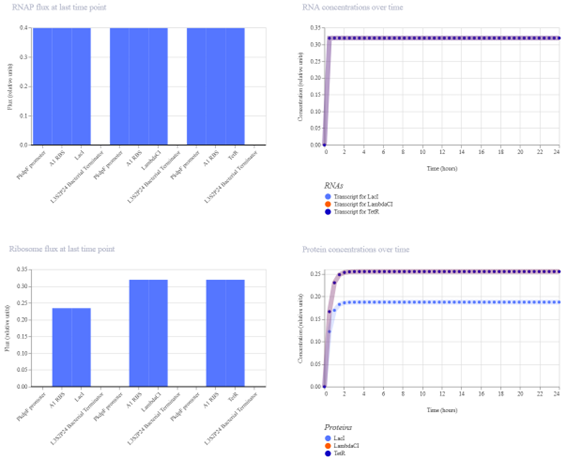

The simulation results show that protein concentrations increase initially and then reach a stable steady state without oscillations. All three proteins are expressed at constant levels over time.
d. Conclusion
This indicates that the constructed circuit does not function as a repressilator. The absence of oscillatory behavior is likely due to the use of identical promoters, which prevents proper regulatory interactions between the genes. As a result, the system behaves as a simple expression circuit rather than a dynamic oscillatory system.
In contrast, the repressilator in the Bacterial Demos repository shows clear oscillations, highlighting that specific regulatory promoters are required to achieve cyclic repression.
4. Build Your Own Constructs:
- Build three of your own Constructs using parts from the Characterized Bacterial Parts Repo
- Explain in the Notebook how you think each Construct should function
- Run the simulator and share your results
- If the results don’t match your expectations speculate why and try adjusting the simulator settings to achieve the expected outcome
Simple reporter (Construct 1)
a. Construct
This construct consists of a single transcriptional unit composed of a promoter, a ribosome binding site (RBS), a coding sequence (GFP), and a terminator. The promoter drives the expression of the GFP reporter gene, allowing direct observation of gene expression levels.
b. Expected behavior
This construct is expected to produce a constant level of GFP expression over time. Since there are no regulatory interactions or feedback mechanisms, the system should reach a stable steady state after an initial increase.
The simulation results show a single line corresponding to GFP protein concentration. The concentration increases rapidly at the beginning and then stabilizes, indicating that the system reaches a steady state.
The results match the expected behavior, as GFP expression increases and then stabilizes at a steady state. This confirms that the construct functions correctly without regulatory interactions.
Toggle (Construct 2)
a. Construct
b. Expected behavior
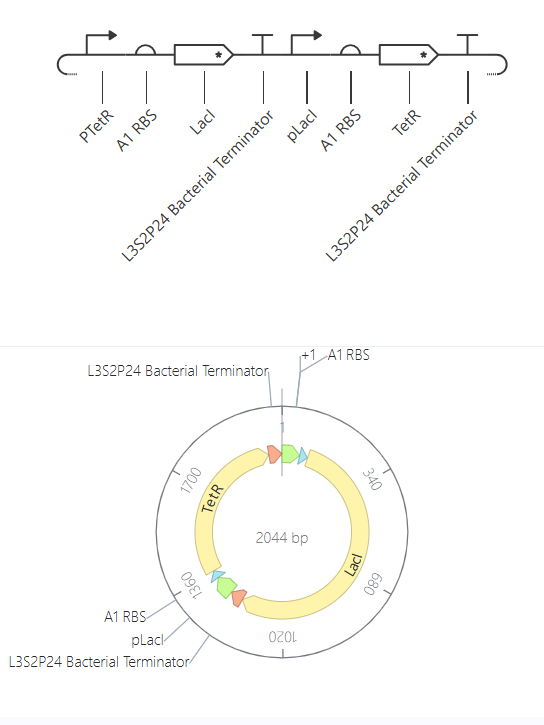
This construct is expected to behave as a toggle switch, where two genes repress each other, leading to a bistable system. Ideally, one gene should be highly expressed while the other is repressed.
The simulation results show that both LacI and TetR reach similar steady-state levels. Instead of bistability, the system converges to a balanced state where both genes are expressed at comparable levels.
The lack of bistability is likely due to the symmetry of the system. Both genes are controlled by similar promoters and conditions, and no external inputs or initial differences are introduced. As a result, the system stabilizes in an intermediate state rather than switching between two distinct states.
To improve the toggle switch behavior, asymmetry was introduced by replacing the A1 RBS with a Q1 RBS for one of the genes. This change created differences in expression levels between LacI and TetR, allowing one gene to dominate over the other. As a result, the system moved away from a symmetric steady state toward partial bistable behavior.
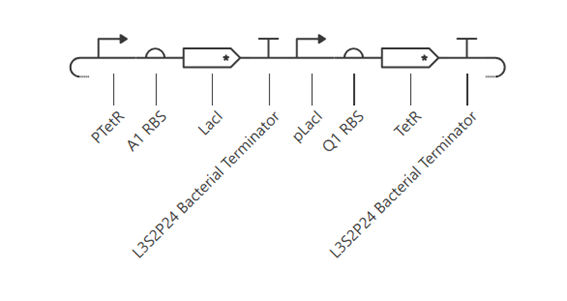

d. Improvement- Second simulation (Q1 RBS)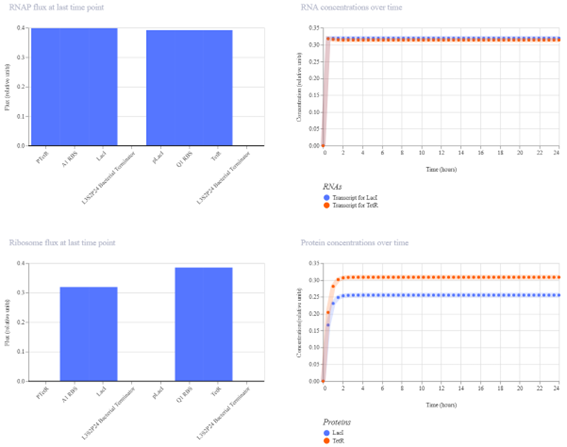

The simulation shows that TetR reaches a higher concentration than LacI, indicating that the symmetry of the system has been broken. Although the system does not exhibit perfect bistability, the results demonstrate partial toggle behavior, where one gene dominates over the other.
Cascade Circuit (Construct 3)
a. Construct
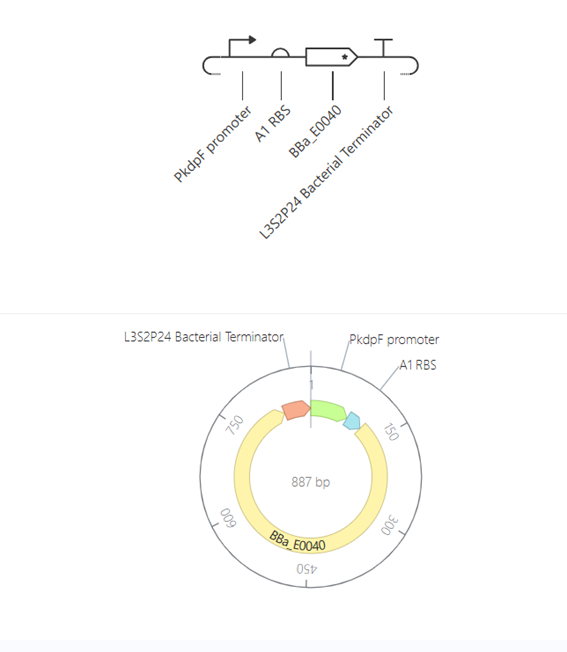
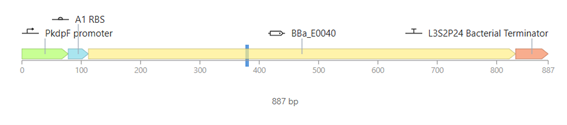
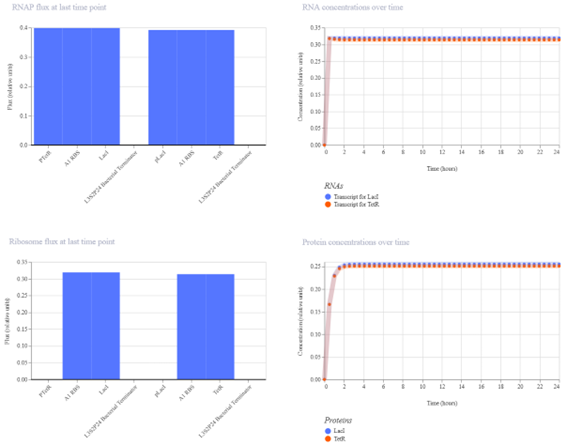
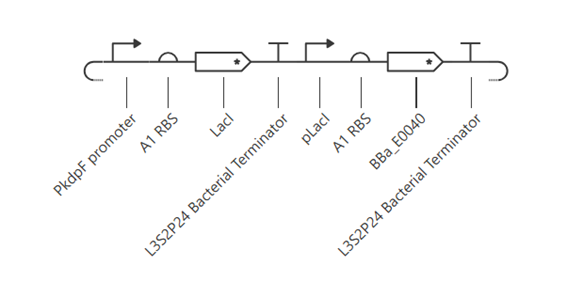
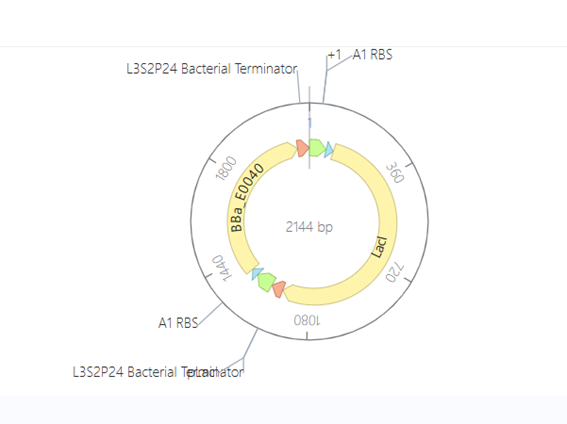
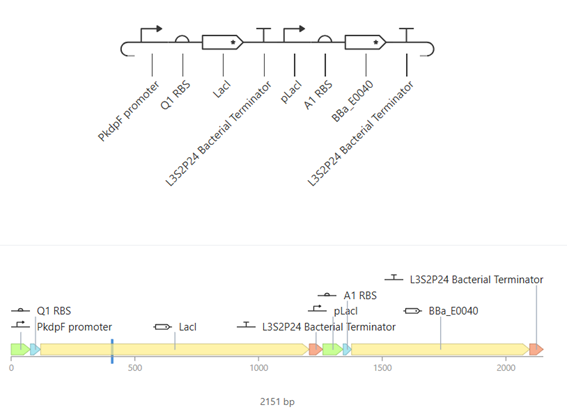
This construct represents a simple genetic cascade composed of two transcriptional units. The first unit includes a constitutive promoter (PkdpF), an RBS, and the LacI coding sequence, leading to continuous production of the LacI repressor.
The second unit contains a LacI-regulated promoter (pLacI), followed by an RBS and the GFP reporter gene (BBa_E0040). In this arrangement, LacI produced in the first unit regulates the expression of GFP in the second unit, creating a one-directional regulatory cascade.
b. Expected behavior
The construct is expected to show cascade behavior, where LacI represses GFP expression through the pLacI promoter. As LacI accumulates, GFP levels decrease, and the system reaches a steady state without oscillations.
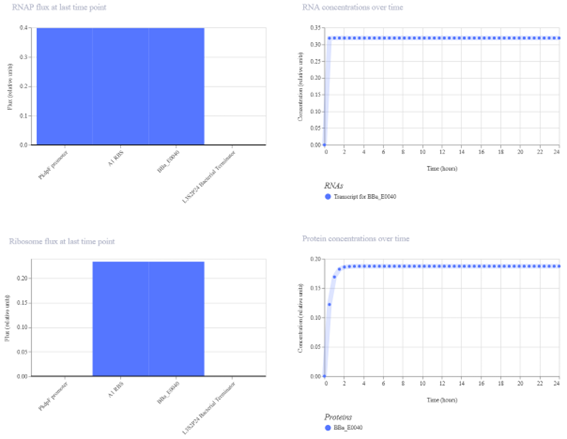
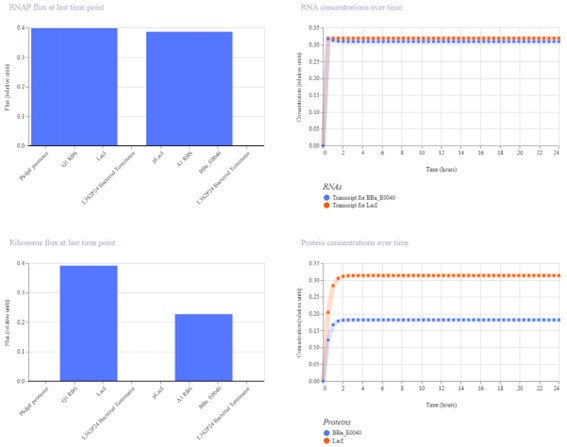
The simulation results show that both LacI and GFP increase initially and then reach a steady state. LacI reaches a slightly higher concentration than GFP, indicating a weak level of repression. The small difference between the two protein levels suggests that LacI only partially reduces GFP expression.
The results partially match the expected behavior. While LacI does repress GFP, the effect is limited, and GFP is not strongly reduced. This suggests that the regulatory interaction is present but not highly efficient.
d. Improvement – Second simulation (Q1 RBS)
To improve the cascade behavior, the RBS upstream of LacI was replaced with a stronger one (Q1 RBS). This increased LacI expression and slightly enhanced repression of GFP, although the effect remains limited.
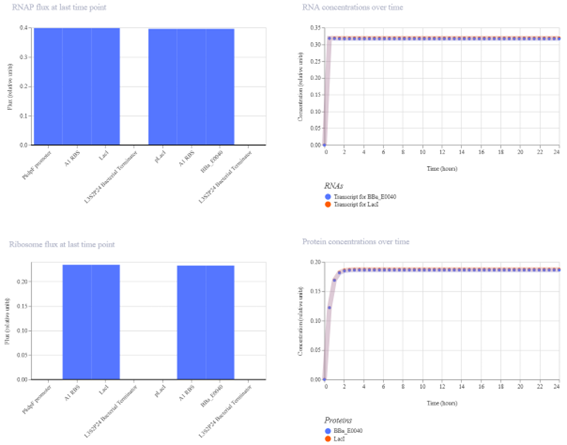
The simulation results show that both LacI and GFP increase initially and then reach a steady state. LacI reaches a significantly higher concentration than GFP, indicating a stronger level of repression. The clear separation between the two protein levels confirms that LacI effectively reduces GFP expression.
The results match the expected behavior of the cascade. As LacI accumulates, it represses GFP expression, resulting in lower GFP levels. The improvement introduced by the stronger RBS enhances this effect, making the regulatory cascade more effective.
CONCLUSION
In this work, several genetic constructs were designed and analyzed to explore different types of regulatory behavior in synthetic biology systems. The simple reporter demonstrated stable gene expression without regulation, while the repressilator highlighted the importance of specific regulatory interactions to achieve oscillatory behavior. The toggle switch showed how mutual repression can lead to bistability, and how introducing asymmetry improves its performance. Finally, the cascade circuit illustrated one-directional regulation, where adjusting expression strength enhanced control over the output gene. Overall, this work demonstrates how the design and tuning of genetic parts, such as promoters and RBS, directly influence circuit behavior, highlighting the importance of precise component selection in synthetic genetic systems.