Week 7 HW: Genetic Circuits: Part II
Week 7 — Genetic Circuits Part II: Neuromorphic Circuits
Part 1: Intracellular Artificial Neural Networks (IANNs)
1. What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions?
IANNs offer several advantages over traditional genetic circuits with Boolean input/output behavior:
- IANNs can process signals in a gradual way, not just ON/OFF like Boolean circuits, which makes them more flexible.
- They can perform more complex computations, instead of relying only on simple logic gates.
- They are easier to adjust, since you can change expression levels without redesigning the whole circuit.
- They handle biological noise better, because they don’t depend on strict thresholds.
- Overall, they are more powerful for modeling complex biological processes.
2. Describe a useful application for an IANN; include a detailed description of input/output behavior, as well as any limitations an IANN might face to achieve your goal.
A useful application for an IANN is as an intelligent disease-detection system inside cells. For example, it could be designed to detect early signs of cancer by sensing multiple biomarkers at the same time.
Input behavior:
- The inputs (X1, X2, etc.) would be DNA sequences that respond to different cellular signals, such as high levels of specific proteins, stress signals, or abnormal gene expression patterns.
- Each input would produce RNA and proteins through transcription and translation, and their levels would represent the strength of each signal.
Processing:
- Inside the IANN, these inputs are integrated using regulatory molecules such as endoribonucleases.
- These molecules act like weights in a neural network, enhancing or inhibiting the expression of downstream genes.
- This allows the system to combine multiple signals and make a more complex decision, rather than just a simple ON/OFF response.
Output behavior:
- The output could be the expression of a fluorescent protein or a therapeutic molecule.
- For example, if the combination of inputs matches a “disease state,” the cell would produce fluorescence (for detection) or release a drug (for treatment).
Limitations:
- Biological systems are noisy, so the response may not always be consistent.
- The processes of transcription and translation are relatively slow, which limits how fast the system can respond.
- It is also difficult to precisely control the “weights” of the network and scaling the system to include many layers or inputs can be complex.
- Additionally, interactions with other cellular processes may interfere with the circuit’s behavior.
3. Draw a diagram for an intracellular multilayer perceptron where layer 1 outputs an endoribonuclease that regulates a fluorescent protein output in layer 2.
First, before drawing the requested two-layer diagram, I will briefly explain the single-layer intracellular perceptron diagram in case it is useful to someone.
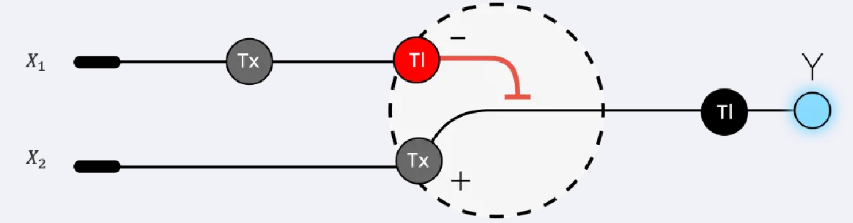

The diagram represents an intracellular single-layer perceptron where the cell processes two genetic inputs to control fluorescence output. Tx stands for transcription, the process in which DNA is copied into messenger RNA (mRNA), while Tl stands for translation, where the mRNA is used to produce a protein. Input X1 encodes the Csy4 endoribonuclease; when X1 is present, the cell transcribes and translates this gene, producing the Csy4 protein. Input X2 encodes a fluorescent protein whose mRNA can be targeted by Csy4. If only X2 is present, the fluorescent protein is produced and fluorescence is ON. However, when X1 is also present, Csy4 cuts the fluorescent mRNA, preventing translation and turning fluorescence OFF. Therefore, fluorescence is activated only when X2 is expressed without Csy4, illustrating how the cell performs a logical computation similar to a perceptron.
| X1 (Csy4) | X2 (Fluorescent gene) | Fluorescence output |
|---|---|---|
| 0 | 0 | OFF |
| 0 | 1 | ON |
| 1 | 0 | OFF |
| 1 | 1 | OFF |
In contrast to the single-layer system, the intracellular multilayer perceptron introduces an additional layer of regulation before producing the final output.
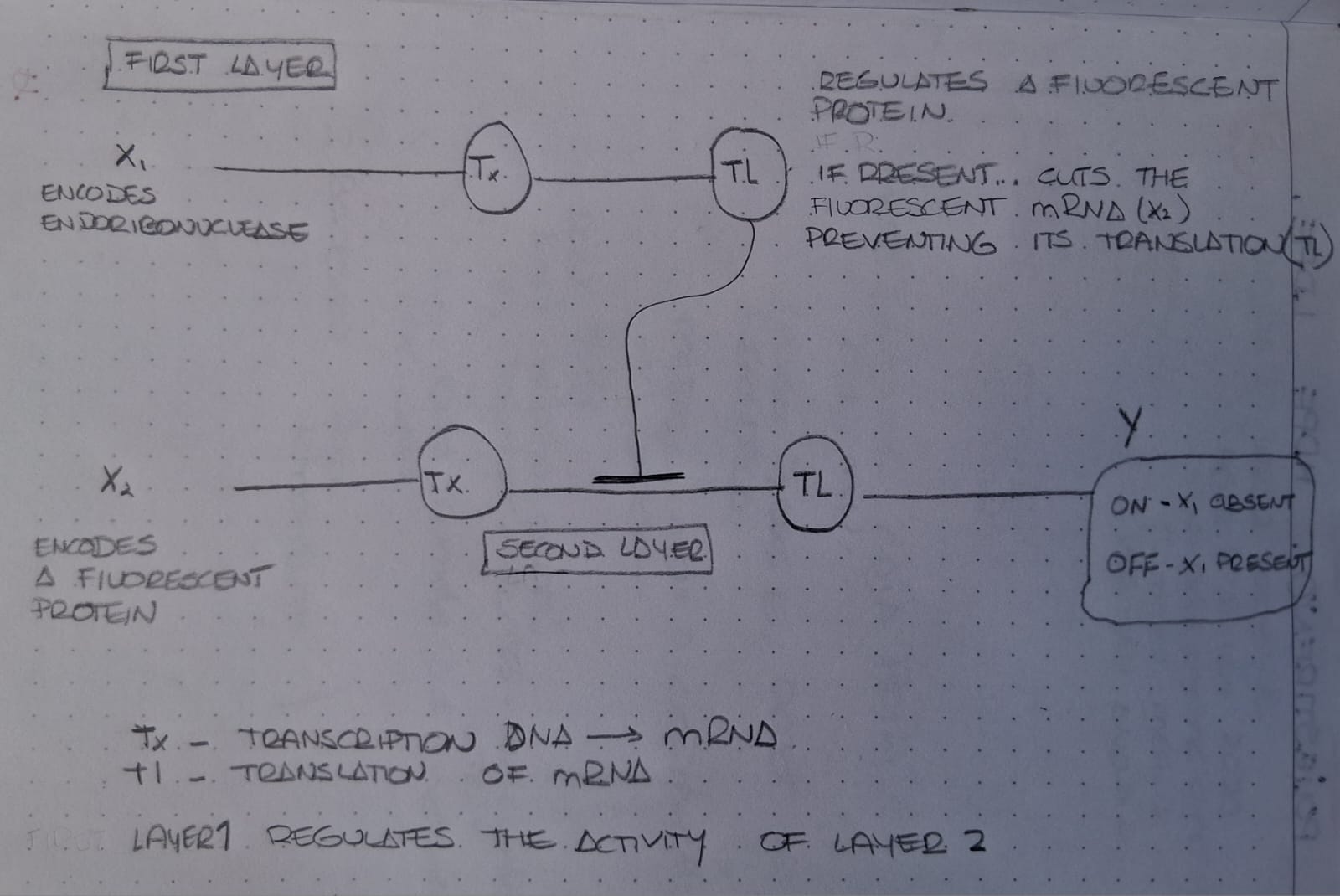

The intracellular multilayer perceptron shown in the figure operates through a sequential regulatory process involving two layers of genetic computation. In the first layer, input X1 encodes an endoribonuclease. After transcription (Tx) and translation (Tl), this gene produces the regulatory protein that functions as an intermediate computational node. At the same time, input X2 encodes a fluorescent protein whose messenger RNA is generated through transcription.
The key feature of the multilayer system is that the product of layer 1 regulates the activity of layer 2. Once synthesized, the endoribonuclease recognizes and cuts the fluorescent mRNA produced from X2, preventing its translation. If X1 is absent, no regulatory enzyme is produced, allowing the fluorescent mRNA to be translated and fluorescence to be observed. Conversely, when X1 is present, the endoribonuclease suppresses translation, turning the fluorescent output OFF.
Thus, the cell performs computation in two consecutive stages: the first layer generates a regulatory signal, and the second layer determines the final observable output. This sequential control mimics the behavior of a two-layer perceptron, where an intermediate node processes input information before producing the final response.
Part 2: Fungal Materials
1. What are some examples of existing fungal materials and what are they used for? What are their advantages and disadvantages over traditional counterparts?
Fungal materials, primarily made from mycelium (the root network of fungi), are emerging as sustainable alternatives to conventional materials.
- They are used in biodegradable packaging as substitutes for plastic foam, in leather-like textiles for fashion products, in construction materials such as insulation panels and acoustic elements, and in furniture and interior design components.
- Their main advantages include renewability, biodegradability, low energy production, and the ability to grow using agricultural waste.
- They are lightweight and provide good thermal and sound insulation.
However, compared to traditional materials, fungal materials can:
- Be less durable
- Be more sensitive to moisture
- Be slower to manufacture
- Sometimes be more expensive
Large-scale production and consistent performance are still developing challenges.
2. What might you want to genetically engineer fungi to do and why? What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
- Fungi could be genetically engineered to produce useful materials and solve environmental problems.
- For example, they could be modified to grow stronger biomaterials, produce medicines, create biodegradable plastics, or help break down waste and pollution.
- Scientists may also engineer fungi to make self-repairing building materials or more resistant textiles.
Advantages of using fungi compared to bacteria:
- Fungi grow as networks, which makes them good for creating large materials.
- They can release enzymes and proteins outside their cells, making production easier.
- Fungi can make more complex molecules that bacteria often cannot produce.
Limitations:
- Fungi usually grow more slowly
- Fungi are harder to modify genetically than bacteria
Overall:
Fungi are very useful for synthetic biology when complex products or materials are needed.
Assignment Part 3: First DNA Twist Order
Review the Individual Final Project documentation guidelines.
Descargar informeSubmit this Google Form with your draft Aim 1, final project summary, HTGAA industry council selections, and shared folder for DNA designs. DUE MARCH 20 FOR MIT/HARVARD/WELLESLEY STUDENTS
3. Review Part 3: DNA Design Challenge of the week 2 homework.
- Design at least 1 insert sequence and place it into the Benchling/Kernel/Other folder you shared in the Google Form above.
- Document the backbone vector it will be synthesized in on your website.
Work flow will be as follow:
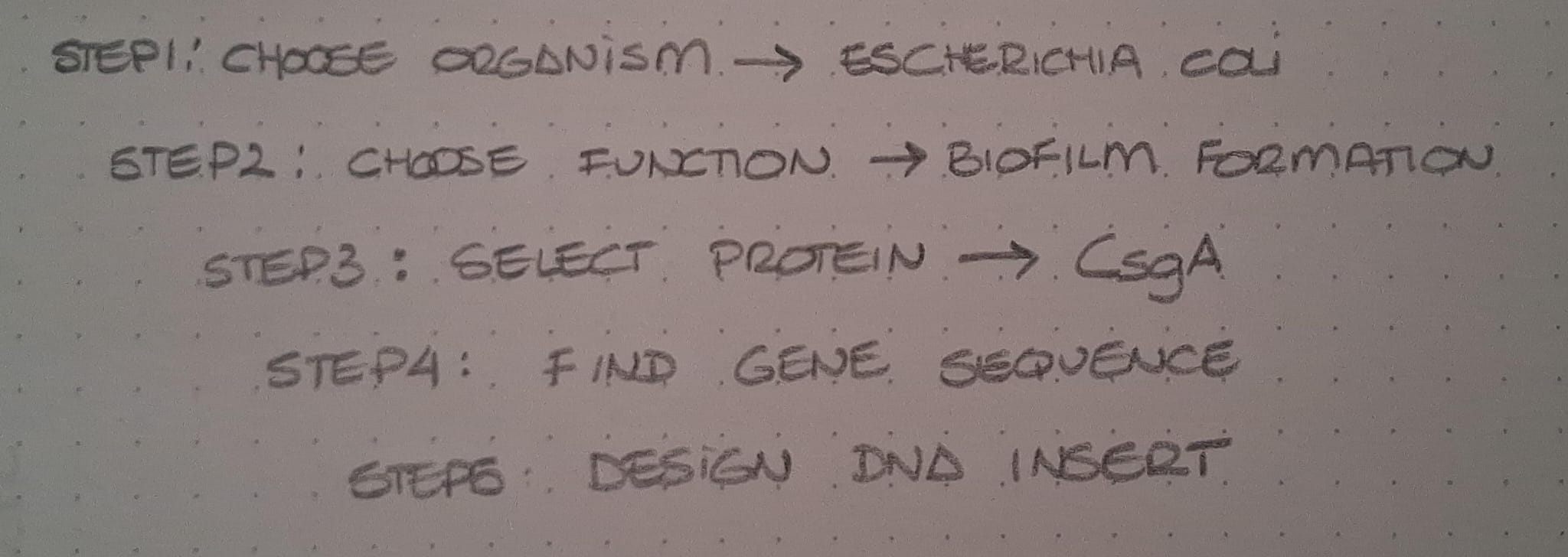

Step 1: Choose the Model Organism
For this project, we selected Escherichia coli as the model organism. It is well-studied, easy to manipulate in the lab, and has well-characterized biofilm-forming proteins. This choice allows us to design a DNA insert that promotes biofilm formation on plastic surfaces in a predictable and explainable way.
Step 2: Define the Biological Function
In this step, we selected biofilm formation as the target function, specifically focusing on initial adhesion to plastic surfaces. This function is essential because it allows bacteria to attach to the material and establish a stable biofilm, which is necessary for further activity on the surface.
Step 3: Select the Target Protein
In this step, we selected the CsgA protein from Escherichia coli. CsgA is the main structural component of curli fibers, which are extracellular protein structures involved in surface adhesion and biofilm formation. This makes it a suitable candidate for promoting attachment to plastic surfaces.
Step 4: Select the Reference Sequence
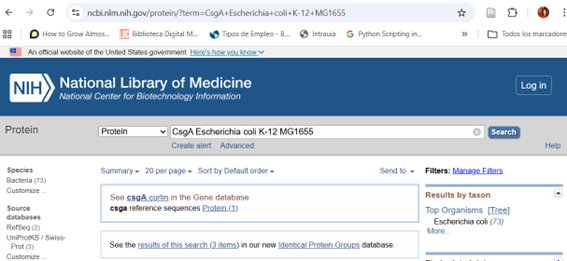

Selection of the Coding Region
Although the complete genome of Escherichia coli MG1655 is available, only the specific coding sequence (CDS) of the csgA gene was selected for this design. The full genome contains thousands of genes and non-coding regions that are not relevant to the intended function.
By isolating only the csgA CDS, we ensure that the insert contains the minimal genetic information required to produce the target protein (CsgA), which is responsible for curli fiber formation and surface adhesion. This approach simplifies the design, avoids unnecessary genetic material, and allows precise control of gene expression when the sequence is inserted into a plasmid vector.

Step 5: Design of the DNA Insert
The selected DNA insert corresponds to the coding sequence (CDS) of the csgA gene from Escherichia coli K-12 MG1655. This sequence encodes the major curlin subunit, which is the main structural component of curli fibers involved in surface adhesion and biofilm formation.
The insert was designed to include only the CDS, ensuring that it contains the minimal genetic information required to produce the target protein. This allows efficient expression when placed under the control of an appropriate promoter in a plasmid vector.
This design supports the overall objective of promoting biofilm formation on plastic surfaces, as the expression of CsgA enhances bacterial adhesion and the establishment of a stable biofilm.
Once the coding sequence was obtained in FASTA format, it was uploaded into Benchling as a linear DNA sequence for visualization and further design.
 Figure 1. pUC19 backbone | 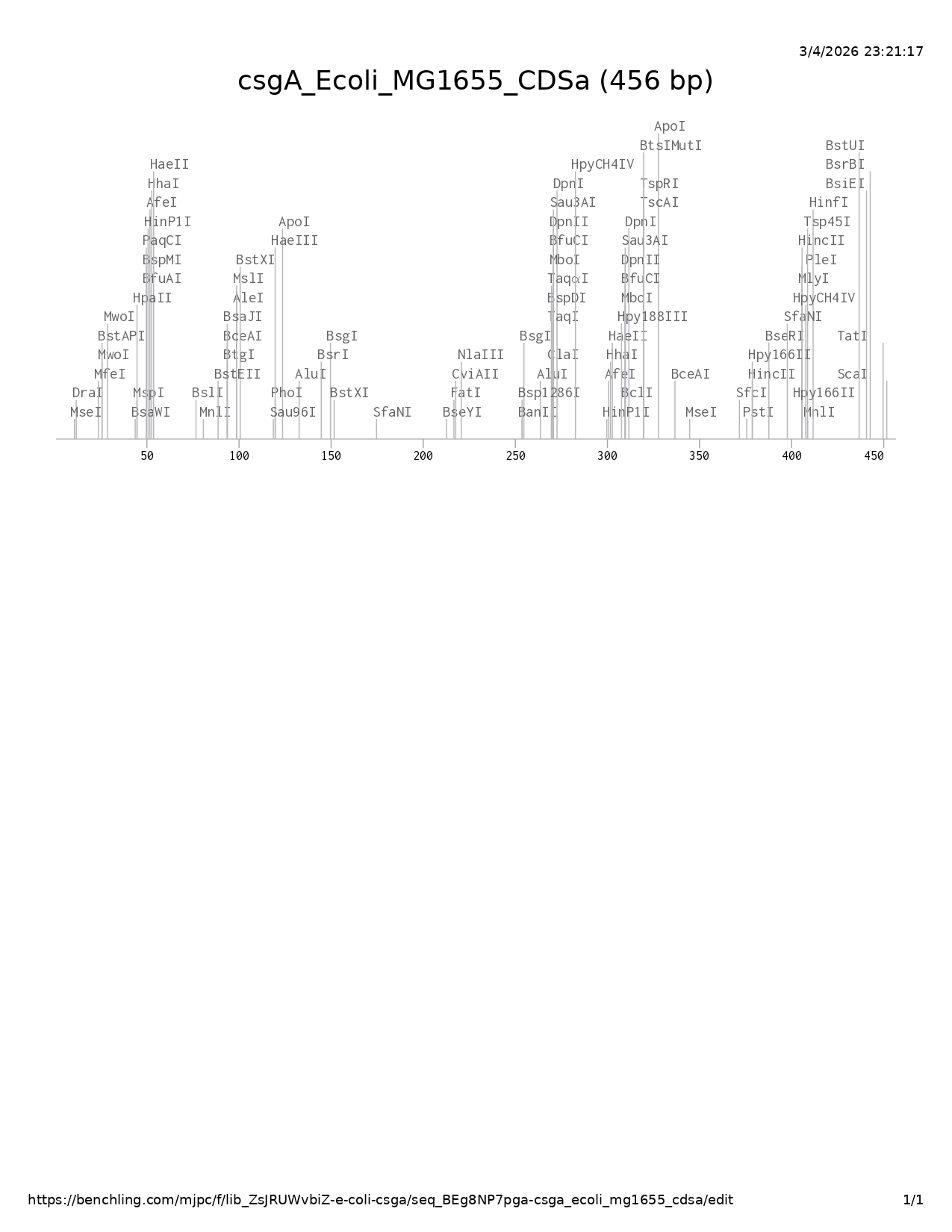 Figure 2. Recombinant plasmid |
Step 6: Selection of the Backbone Vector
Step 6: Selection of the Backbone Vector and Insert Integration
The pUC19 plasmid was selected as the backbone vector due to its widespread use and reliability for cloning in Escherichia coli. It contains an origin of replication, an ampicillin resistance gene, and a multiple cloning site (MCS) for DNA insertion.
The csgA coding sequence was inserted into the MCS of pUC19, specifically between the SalI and PstI restriction sites.
The resulting recombinant plasmid is expected to replicate in E. coli and enable the expression of the CsgA protein, promoting bacterial adhesion and biofilm formation on plastic surfaces.
 Figure 1. pUC19 backbone |  Figure 2. Recombinant plasmid |