Week 9 HW: Cell Free Systems
Part A: General and Lecturer-Specific Questions
1. Explain the main advantages of cell-free protein synthesis over traditional in vivo methods, specifically in terms of flexibility and control over experimental variables. Name at least two cases where cell-free expression is more beneficial than cell production.
Cell-free protein synthesis (CFPS) has several advantages compared to in vivo methods because it is an open system and we can control everything better.
First, it gives more flexibility, because we can add or remove components directly, like DNA, enzymes or cofactors. In living cells this is more difficult because everything depends on cell conditions.
Second, we have better control of experimental variables, like temperature, ion concentration or energy levels. Also, there are no problems with cell viability.
Another important advantage is that it is faster, because we don’t need to grow cells. The reaction starts immediately.
Also, CFPS allows modification of the translation system, so we can incorporate non-natural amino acids and expand the chemical diversity of proteins beyond the 20 standard amino acids.
Cases where CFPS is more useful:
Production of toxic proteins:
Some proteins can kill the cell, so they cannot be produced in vivo, but in cell-free systems it is possible.Rapid testing of proteins or genetic systems:
It is very useful in synthetic biology to test quickly without doing cloning and cell culture.
2. Describe the main components of a cell-free expression system and explain the role of each component.
A cell-free expression system can be understood as the core “cytoplasm” of a synthetic cell, where all the biochemical reactions needed for protein production take place. It is a fully controllable system because we can choose and adjust every component inside it, unlike in living cells.
The main component is the cell extract (lysate), which contains the molecular machinery like ribosomes, enzymes, and especially tRNAs. These tRNAs are very important because they decode the genetic information into proteins, and in cell-free systems they can even be modified to change how the genetic code is read.
Another key component is the DNA or genome, which provides the instructions to produce proteins. In synthetic systems, this genome can be minimized and fully designed depending on what proteins we want to express.
The system also includes small molecules, such as amino acids, nucleotides, and energy sources. These are essential because they act as building blocks and provide the energy needed for transcription and translation. One advantage is that we can control exactly which molecules are present, allowing us to modify the internal chemistry of the system.
Finally, although not always considered part of the core reaction, in synthetic cells this system is usually placed inside a liposome (membrane). The membrane allows compartmentalization and, together with membrane channels, enables communication with the environment, such as importing nutrients or exporting products.
In conclusion, a cell-free system is composed of the expression machinery, genetic material, and small molecules, all of which can be precisely controlled to direct protein production.
3. Why is energy provision regeneration critical in cell-free systems? Describe a method you could use to ensure continuous ATP supply in your cell-free experiment.
Energy regeneration is very important in cell-free systems because protein synthesis needs a lot of energy, mainly in the form of ATP and GTP.
Since there are no living cells, the system cannot produce energy by itself. If ATP is consumed and not regenerated, the reaction will stop very quickly and protein production will be very low.
Also, processes like transcription and translation are very energy demanding, so without a continuous energy supply, the system is not efficient.
One common method to ensure continuous ATP supply is to use an energy regeneration system, for example with phosphoenolpyruvate (PEP). In this case, PEP is used together with enzymes like pyruvate kinase to regenerate ATP from ADP.
Another option is using creatine phosphate + creatine kinase, which also helps to recycle ATP. These systems allow the reaction to continue for longer time and increase protein yield.
4. Compare prokaryotic versus eukaryotic cell-free expression systems. Choose a protein to produce in each system and explain why.
Cell-free expression systems can be divided into prokaryotic and eukaryotic, and they present important differences in terms of efficiency and protein complexity.
Prokaryotic systems, such as those based on E. coli, are usually faster, cheaper, and produce high protein yields. However, they have limitations because they cannot perform most post-translational modifications, and sometimes proteins do not fold correctly if they are complex.
In contrast, eukaryotic cell-free systems, like wheat germ, insect, or mammalian extracts, are more suitable for producing complex proteins. These systems allow post-translational modifications such as glycosylation and support proper protein folding. The disadvantage is that they are slower, more expensive, and sometimes give lower yields compared to prokaryotic systems.
An example of a protein that can be efficiently produced in a prokaryotic system is the green fluorescent protein (GFP). This protein is relatively simple, does not require complex modifications, and folds correctly in bacterial systems, so using a prokaryotic extract is more practical and efficient.
On the other hand, a good example for a eukaryotic system would be a human antibody. Antibodies require correct folding and post-translational modifications, especially glycosylation, to be functional. These processes cannot be properly carried out in prokaryotic systems, so a eukaryotic cell-free system is a better choice.
In conclusion, prokaryotic systems are ideal for simple and fast protein production, while eukaryotic systems are necessary when producing complex proteins that require modifications and proper folding.
5. How would you design a cell-free experiment to optimize the expression of a membrane protein? Discuss the challenges and how you would address them in your setup.
To optimize the expression of a membrane protein in a cell-free system, I would focus on designing an appropriate membrane environment. I would use liposomes composed of phospholipids and cholesterol, since cholesterol improves membrane fluidity and stability, which helps proper insertion of the protein.
The main challenge is that membrane proteins are hydrophobic and can aggregate if they do not interact correctly with the membrane. To address this, I would optimize the lipid composition, adjusting the ratio of phospholipids and cholesterol to create a membrane that favors correct insertion and folding.
I would also control membrane properties such as fluidity and permeability and possibly include membrane channels to allow exchange of molecules and maintain proper conditions inside the system.
In summary, optimizing the membrane composition and properties is key to avoid aggregation and ensure correct folding and functionality of the membrane protein.
6. Imagine you observe a low yield of your target protein in a cell-free system. Describe three possible reasons for this and suggest a troubleshooting strategy for each.
If a low yield of the target protein is observed in a cell-free system, there are several possible reasons related to expression efficiency, protein degradation, and folding.
First, one possible reason is inefficient transcription or translation. This can happen if the promoter is weak, the ribosome binding site is not optimal, or the codon usage is not well adapted. To solve this, the DNA can be optimized by using a strong promoter (for example T7), improving the ribosome binding site, and doing codon optimization to increase expression.
Second, protein degradation can also reduce the yield. This is usually caused by proteases present in the extract, which degrade the protein after it is produced. To solve this, we can use more purified extracts or add protease inhibitors to protect the protein.
Third, low yield can be due to incorrect protein folding or aggregation. If the protein does not fold correctly, it can become inactive or form aggregates. To improve this, we can add chaperones and adjust conditions like temperature or redox environment to help proper folding.
In conclusion, low protein yield can be caused by problems in expression, degradation, or folding, and these can be improved by optimizing the DNA and reaction conditions.
Homework question from Kate Adamala
Design an example of a useful synthetic minimal cell as follows:
1. Pick a function and describe it.
a. What would your synthetic cell do? What is the input and what is the output? The synthetic cell will detect early biofilm formation on oxygen delivery tubing and generate a visible signal to indicate contamination. The input will be quorum sensing molecules (AHL) produced by bacteria during early biofilm formation. These molecules diffuse into the synthetic cell and bind to the LuxR protein, activating gene expression. The output will be a visible signal, such as fluorescence (e.g., GFP), indicating the presence of bacterial contamination.
b. Could this function be realized by cell-free Tx/Tl alone, without encapsulation? No. Without encapsulation, the cell-free system would not be localized on the surface of the oxygen tubing, and the components could diffuse away or degrade more easily. The membrane provides compartmentalization, allowing the system to remain stable and concentrated at the site where biofilm formation occurs. This localization is important to ensure reliable detection and a clear signal in response to bacterial contamination.
c. Could this function be realized by genetically modified natural cell? Yes, this function could be achieved using genetically modified bacteria that detect quorum sensing molecules and produce a reporter signal. However, using living cells in medical devices such as oxygen tubing raises safety and regulatory concerns, including the risk of contamination or uncontrolled growth. In contrast, synthetic cells provide a safer and more controllable alternative, as they are non-living systems and can be designed to operate only under specific conditions.
d. Describe the desired outcome of your synthetic cell operation. The desired outcome is the early detection of biofilm formation on oxygen delivery tubing through a clear and visible signal. This would allow timely intervention, such as replacing or cleaning the tubing, before significant bacterial growth occurs. By providing a localized and real-time indication of contamination, the system aims to improve patient safety and reduce the risk of respiratory infections associated with biofilm formation.
2. Design all components that would need to be part of your synthetic cell.
a. What would be the membrane made of? The membrane would be composed of phospholipids (such as POPC) and cholesterol. This composition provides stability, fluidity, and biocompatibility, allowing the synthetic cell to maintain its structure while permitting diffusion of small molecules like quorum sensing signals.
b. What would you encapsulate inside? Enzymes, small molecules. The synthetic cell would encapsulate a cell-free transcription-translation (Tx/Tl) system, including ribosomes, enzymes, tRNAs, and energy components. It would also contain DNA encoding a quorum sensing detection circuit, including the LuxR protein and a reporter gene such as GFP under the control of a LuxR-activated promoter. Additionally, small molecules such as amino acids, nucleotides, and energy sources would be included to support protein expression.
c. Which organism your Tx/Tl system will come from? Is bacterial OK, or do you need a mammalian system for some reason? (hint: for example, if you want to use small molecule modulated promotors, like Tet-ON, you need mammalian) A bacterial cell-free system derived from E. coli would be used. This is suitable because the quorum sensing system (LuxR and AHL) and the reporter gene (GFP) are naturally compatible with bacterial expression systems. A mammalian system is not required, as no complex post-translational modifications or mammalian-specific regulatory elements are needed. Additionally, the system is encapsulated and immobilized within the material of the oxygen tubing, preventing direct exposure to the oxygen flow and ensuring safe operation in a medical environment.
d. How will your synthetic cell communicate with the environment? (hint: are substrates permeable? or do you need to express the membrane channel?) The synthetic cell will communicate with the environment through passive diffusion of small molecules. Quorum sensing molecules (AHL) are small and can diffuse across the lipid membrane into the synthetic cell. Inside, they activate the genetic circuit leading to reporter protein expression. The output signal, such as GFP fluorescence, does not need to exit the cell, as it can be detected externally. Therefore, no additional membrane channels are required.
3. Experimental details
a. List all lipids and genes. (bonus: find the specific genes; for example, instead of just saying “small molecule membrane channel” pick the actual gene.) Lipids: POPC (phosphatidylcholine) and cholesterol.
Genes: luxR gene encoding the quorum sensing regulator, and a reporter gene (gfp) under the control of a LuxR-activated promoter (Plux). These components enable detection of quorum sensing molecules and production of a visible fluorescence signal.
b. How will you measure the function of your system? The function of the system will be measured by detecting fluorescence from the reporter protein (GFP). Increased fluorescence intensity indicates the presence of quorum sensing molecules and activation of the synthetic cell. Fluorescence can be measured visually or using a fluorescence reader to quantify signal intensity.
Homework question from Peter Nguyen
Freeze-dried cell-free systems can be incorporated into all kinds of materials as biological sensors or as inducible enzymes to modify the material itself or the surrounding environment. Choose one application field — Architecture, Textiles/Fashion, or Robotics — and propose an application using cell-free systems that are functionally integrated into the material. Answer each of these key questions for your proposal pitch:
1. Write a one-sentence summary pitch sentence describing your concept.
I propose oxygen delivery tubing embedded with freeze-dried cell-free systems that detect early biofilm formation, trigger a visible color change, and produce enzymes to prevent microbial colonization.
2. How will the idea work, in more detail? Write 3-4 sentences or more.
The oxygen tubing would incorporate embedded freeze-dried cell-free systems immobilized within a protective matrix along the inner surface of the material. These systems would be programmed with DNA circuits that detect biofilm-associated signals, such as quorum-sensing molecules or bacterial metabolites. Upon exposure to moisture, the system activates and produces a visible color change through reporter proteins, providing an early warning of contamination. At the same time, it expresses enzymes that degrade the extracellular polymeric substances (EPS) matrix, disrupting biofilm formation. The cell-free components remain physically encapsulated within the material to prevent direct exposure to the patient while enabling localized detection and response.
3. What societal challenge or market need will this address?
Biofilm formation in oxygen delivery systems, particularly those involving humidification, poses a risk of respiratory infections and patient complications. Current prevention methods rely on strict sterilization protocols and disposable components, which increase costs and resource use. This project addresses the need for safer, self-monitoring medical devices that can detect and prevent contamination in real time, improving patient safety while reducing maintenance and waste.
4. How do you envision addressing the limitation of cell-free reactions (e.g., activation with water, stability, one-time use)?
To address the limitations of cell-free systems, the components would be encapsulated within stable, biocompatible matrices to improve durability and prevent direct exposure to the patient. Activation would be controlled by moisture present in the oxygen delivery system, ensuring the system functions only when needed. To overcome the limitation of one-time use, the tubing could incorporate replaceable or modular inner coatings that allow periodic renewal of the active components. Advances in stabilization of freeze-dried systems would further extend shelf life and maintain functionality under clinical conditions.
Homework question from Ally Huang
Freeze-dried cell-free reactions have great potential in space, where resources are constrained. As described in my talk, the Genes in Space competition challenges students to consider how biotechnology, including cell-free reactions, can be used to solve biological problems encountered in space. While the competition is limited to only high school students, your assignment will be to develop your own mock Genes in Space proposal to practice thinking about biotech applications in space!
For this particular assignment, your proposal is required to incorporate the BioBits® cell-free protein expression system, but you may also use the other tools in the Genes in Space toolkit (the miniPCR® thermal cycler and the P51 Molecular Fluorescence Viewer). For more inspiration, check out https://www.genesinspace.org/ .
1. Provide background information that describes the space biology question or challenge you propose to address. Explain why this topic is significant for humanity, relevant for space exploration, and scientifically interesting. (Maximum 100 words)
Spaceflight exposes biological systems to unique conditions, including increased radiation and microgravity, which can affect DNA stability. Telomeres are repetitive DNA sequences that protect chromosome integrity and are known to be sensitive to environmental stress and DNA damage. Changes in telomere integrity are associated with aging, genomic instability, and disease. Understanding how spaceflight conditions affect telomeric DNA is important for astronaut health during long missions. This topic is scientifically interesting because it helps us study DNA damage mechanisms in extreme environments and supports the development of safer strategies for human space exploration.
2. Name the molecular or genetic target that you propose to study. Examples of molecular targets include individual genes and proteins, DNA and RNA sequences, or broader -omics approaches. (Maximum 30 words)
Synthetic telomeric DNA sequences (TTAGGG repeats) used as a model to study telomere-associated DNA damage under spaceflight conditions.
3. Describe how your molecular or genetic target relates to the space biology question or challenge your proposal addresses. (Maximum 100 words)
The molecular target, synthetic telomeric DNA sequences, represents the repetitive regions found at the ends of chromosomes that are particularly sensitive to damage. By analyzing the integrity of these sequences after exposure to spaceflight conditions, we can assess how radiation and microgravity affect telomere stability. Since telomeres play a key role in protecting genomic DNA, damage to these regions can lead to genomic instability and cellular dysfunction. Therefore, studying telomeric DNA provides a relevant model to understand how space conditions impact DNA integrity and potential risks for astronaut health.
4. Clearly state your hypothesis or research goal and explain the reasoning behind it. (Maximum 150 words)
Spaceflight conditions, including radiation and microgravity, increase telomere-associated DNA damage compared to Earth conditions. Telomeres are repetitive DNA sequences that are particularly sensitive to environmental stress and DNA damage. In space, higher radiation levels and altered physical conditions can lead to strand breaks and base modifications, which may affect DNA integrity. Since telomeres play an important role in protecting chromosomes, damage in these regions could contribute to genomic instability and long-term health risks for astronauts.
This experiment aims to evaluate whether exposure to spaceflight conditions reduces the integrity of telomeric DNA. By comparing samples processed in space and on Earth, differences in PCR amplification and reporter expression can indicate levels of damage. This approach allows us to study DNA stability in extreme environments using a simple and controlled system.
5. Outline your experimental plan - identify the sample(s) you will test in your experiment, including any necessary controls, the type of data or measurements that will be collected, etc. (Maximum 100 words)
Identical synthetic telomeric DNA samples will be prepared and divided into Earth and spaceflight conditions. In space, samples will be processed using the miniPCR® thermal cycler to amplify telomeric sequences. PCR products will then be used as templates in the BioBits® cell-free system to express GFP. Fluorescence will be measured using the P51 Molecular Fluorescence Viewer. Earth samples will follow the same protocol as controls. Data collected will include PCR amplification efficiency and fluorescence intensity. Lower amplification and fluorescence in space samples will indicate increased telomere-associated DNA damage.
Homework Part B: Individual Final Project
Final Project Instructions
We’d like students to start exploring their final project in depth this week! Of your three Aims, for this week you should have at least Aim 1 decided and written down.
1. Put your chosen final project slide in the appropriate slide deck following the instructions on slide 1:
- MIT/Harvard/Wellesley ONE FINAL PROJECT IDEA
- Committed Listener ONE FINAL PROJECT IDEA (DONE) 2. and submit this Final Project selection form if you have not already. (DONE)
3.Begin planning how you will write your final project documentation based on these guidelines.
4. Prepare your first DNA order and put it in the “Twist (MIT)” or “Twist (Nodes)” tab of the 2026 HTGAA Ordering: DNA, Reagents, Consumables spreadsheet, as appropriate. (DONE, See below).
Final DNA Insert Design Description for twist order
The original PETase (HMW2-based) sequence from Assignment 2 was replaced with the csgA gene from Escherichia coli, aligning the design with the final project’s focus on biofilm formation. The csgA gene encodes the main structural component of curli fibers, which are essential for surface adhesion and biofilm development.
csgA Sequence
Length: 456 bp
Description: Coding sequence including start and stop codons
The insert was designed as a complete expression cassette, including a constitutive promoter (BBa_J23106), a ribosome binding site (RBS, GCCACC), an ATG start codon, the csgA coding sequence, a TAA stop codon, and a BBa_B0015 double terminator to ensure proper transcription and translation in E. coli.
For cloning and propagation, the backbone vector selected is pUC19, chosen for its high copy number, ampicillin resistance marker, and well-characterized multiple cloning site.
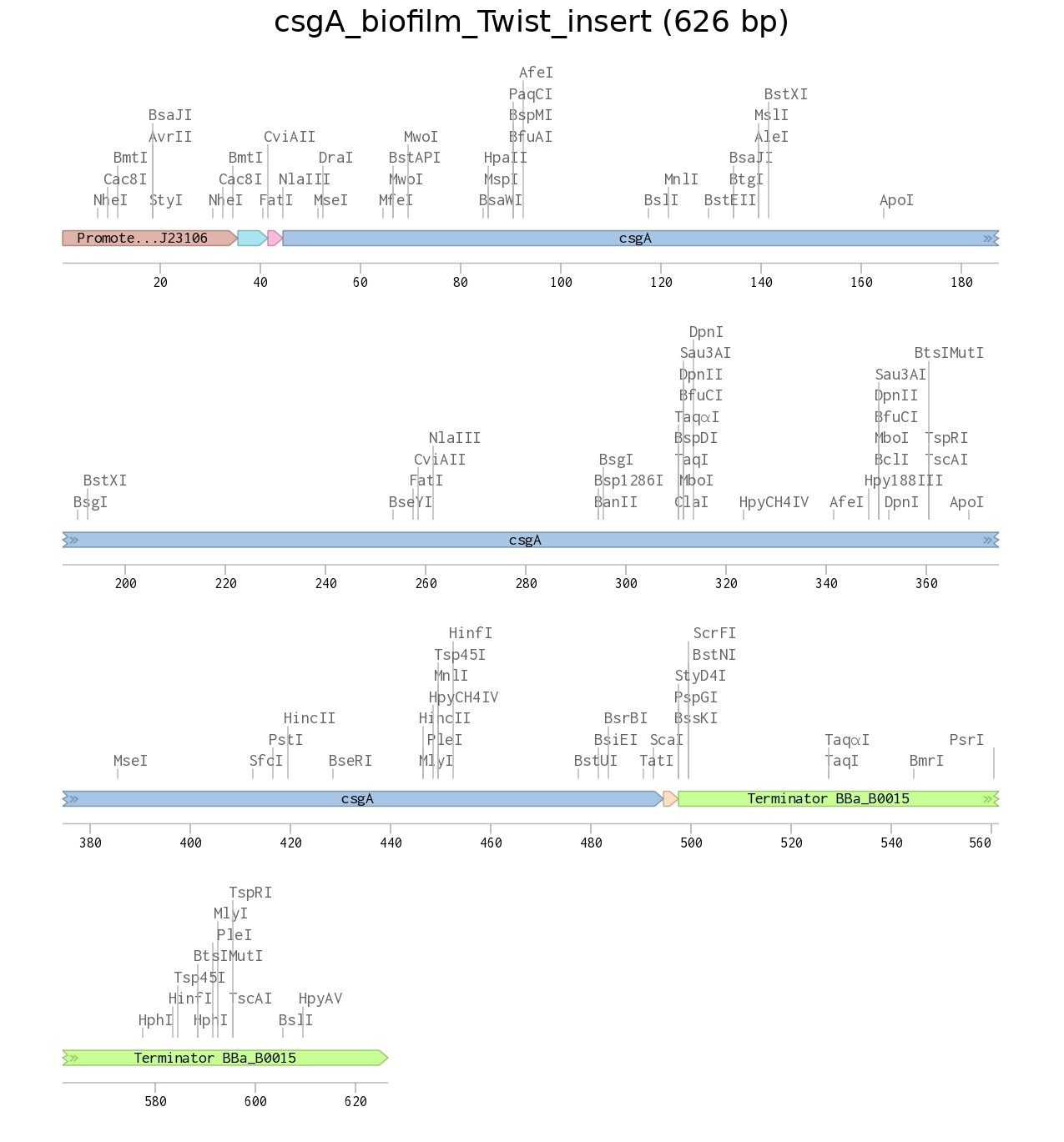 Figure 1. pUC19 backbone | 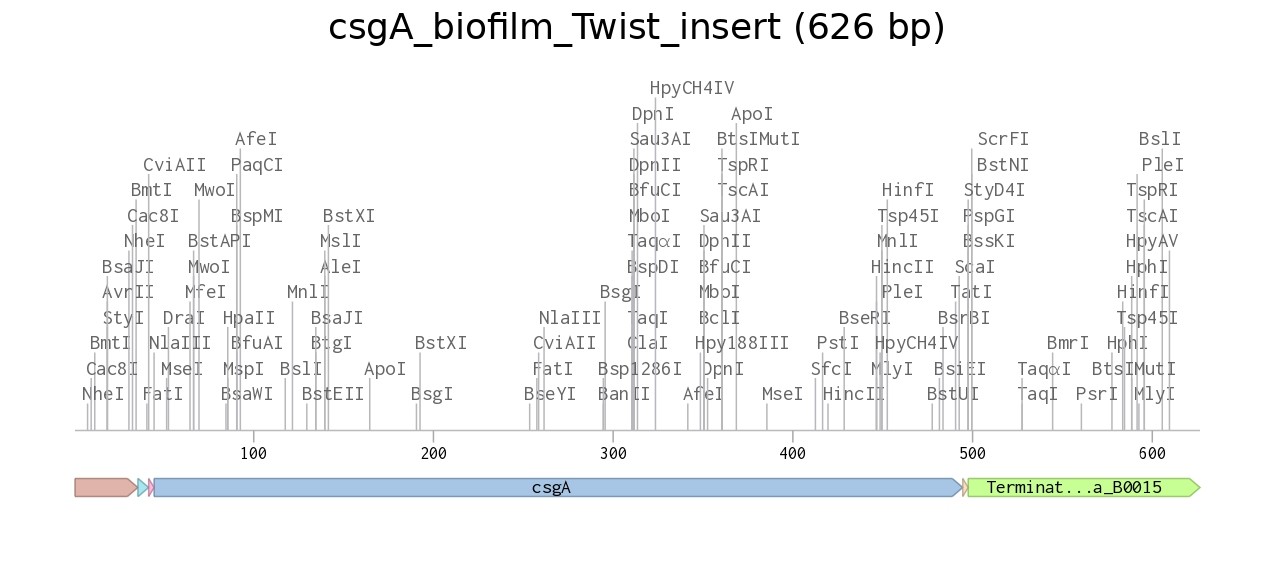 Figure 2. Recombinant plasmid |
The designed insert sequence can be accessed here: Benchling sequence link.