Week 11 HW: Bioproduction & Cloud Labs
Cloud laboratories are making science accessible, affordable, and reproducible. Our aim this semester is to showcase how they can enable human creativity at scale, and how they provide a platform for collaboration and community.
How To Grow (Almost) Anything is about synthetic biology, bioengineering, robotics, automation, art, and AI. But it is also about friendship, shared purpose, and the freedom to build beyond what we know and to be inspired by what can be. To that end, the goal with this cloud lab unit and homework assignment is to inspire collaboration and creativity while designing a scientifically rigorous cell-free fluorescent protein optimization experiment together.
Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork
1. Contribute at least one pixel to this global artwork experiment before the editing ends on Sunday 4/19 at 11:59 PM EST.
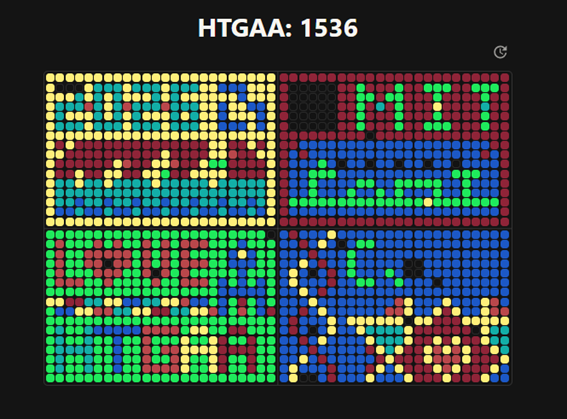
Figure 1. Screen shoot from Collective Artwork webpage.
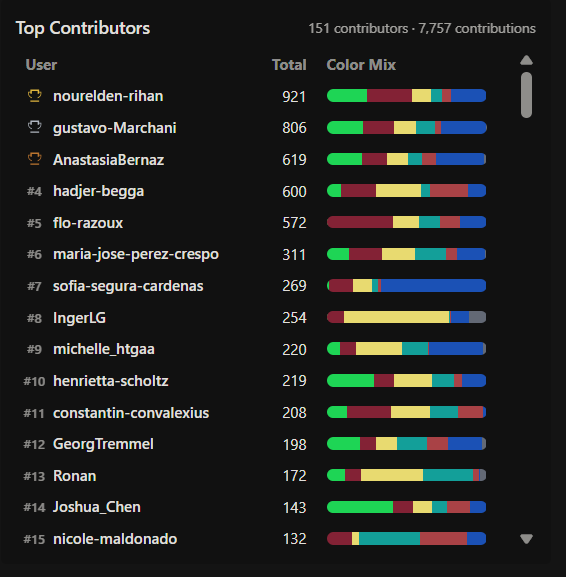
Figure 2. Screen shoot from Collective Artwork webpage.
2. Make a note on your HTGAA webpages including:what you contributed to the community bioart project, what you liked about the project, and what about this collaborative art experiment could be made better for next year.
Participation in the Bioart Project At the beginning of the experiment, I tried to contribute by adding several pixels to the design. However, I noticed that my contributions were disappearing very quickly, and nothing I did remained in the final artwork. I tried at different times to modify some pixels, but they were constantly reverted, as if someone — or maybe a machine — was restoring the design the whole time. In the end, I just tried to leave at least one pixel as a minimal contribution, but it was not possible. Every change I made was undone almost immediately. Only a few pixeles remained (in red circle) from a total of 311 inputs.
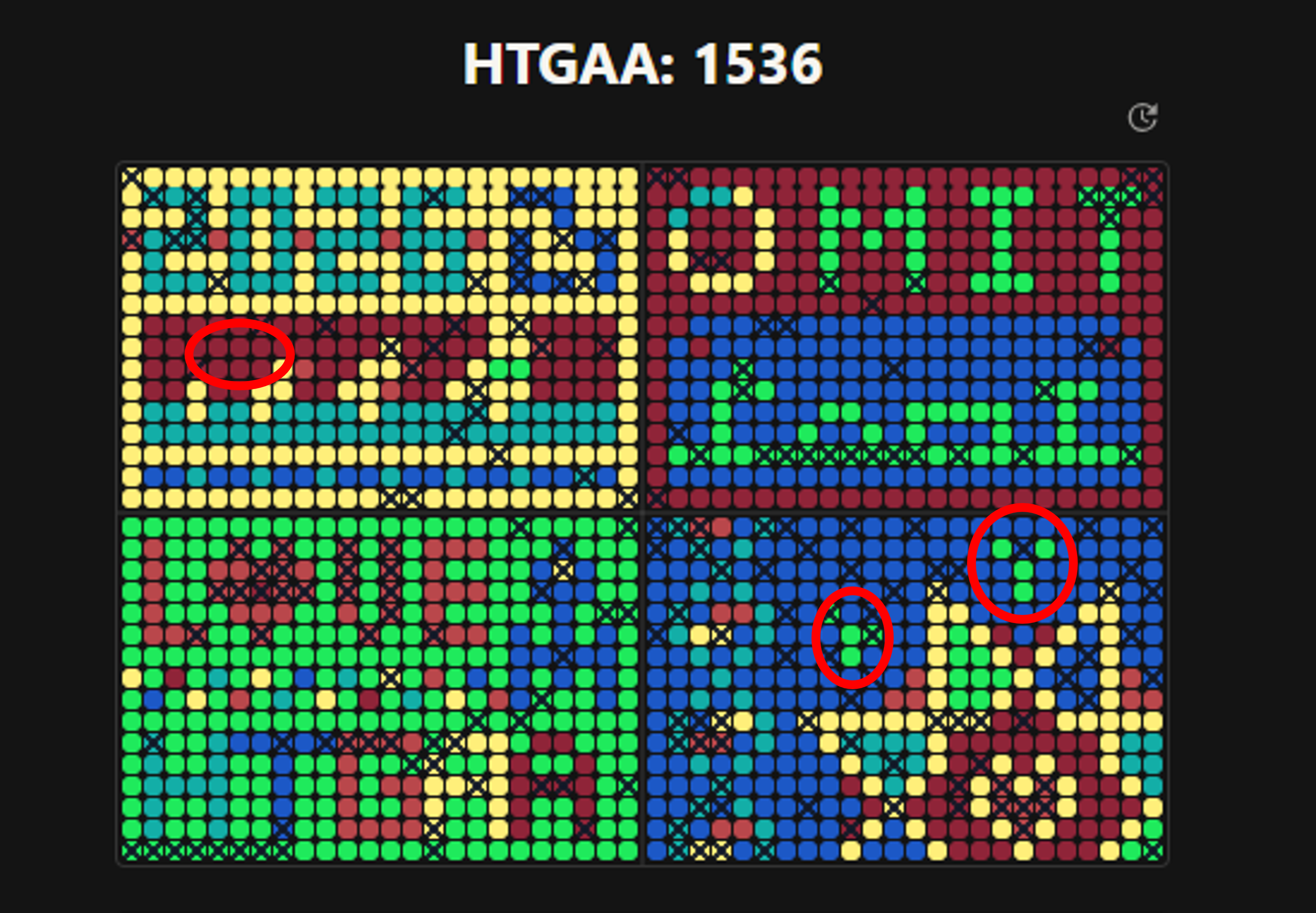
Figure 3. Screen shoot from final Collective Artwork webpage. My contribution in red circles.
Reflection on the Experience Because of this, I did not feel that the exercise was truly cooperative. Even though the objective was to build a collective artwork, in practice the design became unchangeable at some point during the experiment. This limited the real participation of many users. Also, in my case, the large number of pixels I added was not because I wanted to contribute more than others, but because I was trying to understand what was happening with the pixels I placed. Even so, none of my pixels remained in the final result, so my contribution is not reflected. This is not what collaboration should be — all participants should be able to see at least a small part of their work in the final design.
Suggestions for Improvement To improve this kind of experience in the future, some rules or systems could be implemented, for example:
- Set a limit on the number of pixels each person can modify or input.
- Avoid constant overwriting of other participants’ contributions.
- Encourage building a collective design while respecting individual inputs. If the time between infpus of each participant increases it could prevent an individual to create a particular design.
- Ensure that every user can leave at least one visible contribution in the final artwork.
In conclusion, the goal should be that everyone can contribute in a meaningful way and that these contributions remain visible, reinforcing the collaborative nature of the experiment.
Part B: Cell-Free Protein Synthesis | Cell-Free Reagents
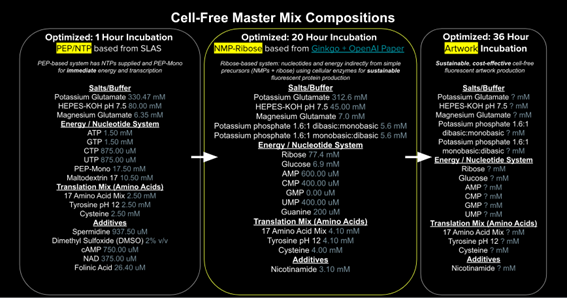
Figure 4. Cell-Free master mix compositions.
1. Referencing the cell-free protein synthesis reaction composition (the middle box outlined in yellow on the image above, also listed below), provide a 1-2 sentence description of what each component’s role is in the cell-free reaction.
E. coli Lysate
- BL21 (DE3) Star Lysate (includes T7 RNA Polymerase) - Crude cell extract from E. coli BL21(DE3) Star that provides the core machinery for protein synthesis — ribosomes, tRNAs, translation factors, and metabolic enzymes. It also contains T7 RNA Polymerase, which transcribes DNA templates under T7 promoters into mRNA, enabling both transcription and translation in the cell-free reaction.
Salts/Buffer
- Potassium Glutamate — Provides K⁺ ions required for ribosome assembly and translation fidelity, while glutamate acts as a biocompatible counter-ion that mimics the intracellular environment better than chloride.
- HEPES-KOH pH 7.5 — A biological buffer that maintains the reaction at physiological pH (~7.5), keeping enzymes active and preventing pH shifts as metabolic reactions consume or release protons.
- Magnesium Glutamate — Supplies Mg²⁺, an essential cofactor for RNA polymerase, ribosomes, and most nucleotide-dependent enzymes; it also stabilizes RNA structure and tRNA–ribosome interactions.
- Potassium Phosphate Monobasic (KH₂PO₄) — Provides inorganic phosphate (Pᵢ) needed for energy regeneration, and together with the dibasic form establishes a secondary phosphate buffer that stabilizes pH.
- Potassium Phosphate Dibasic (K₂HPO₄) — The basic counterpart of the phosphate pair; contributes K⁺ and Pᵢ, and works with the monobasic form to maintain pH and support phosphorylation reactions in energy metabolism.
Energy / Nucleotide System
- Ribose — A 5-carbon sugar that serves as the scaffold for ribonucleotide biosynthesis; cellular enzymes phosphorylate it to form PRPP, which combines with nitrogenous bases to generate NTPs for transcription.
- Glucose — The main energy source of the reaction; it feeds into glycolysis to regenerate ATP (and other NTPs indirectly), sustaining the energy supply throughout the 20-hour incubation.
- AMP (Adenosine Monophosphate) — A low-cost precursor of ATP; adenylate kinase and NDP kinase in the lysate phosphorylate it stepwise (AMP → ADP → ATP) to provide the ATP used by T7 RNA polymerase and the ribosomes.
- CMP (Cytidine Monophosphate) — Precursor of CTP; lysate kinases convert CMP → CDP → CTP, which the RNA polymerase incorporates as cytidine residues in the nascent mRNA. GMP (Guanosine Monophosphate) — Normally the precursor of GTP (essential for translation elongation factors and transcription); in this formulation it is set to 0 μM and replaced by free guanine, which is recycled into GMP via the salvage pathway.
- UMP (Uridine Monophosphate) — Precursor of UTP; phosphorylated by cellular kinases (UMP → UDP → UTP) to provide the uridine residues incorporated into mRNA by RNA polymerase.
- Guanine — A free nitrogenous base that enters the salvage pathway: the enzyme HGPRT combines guanine with PRPP (derived from ribose + ATP) to generate GMP directly, which is then phosphorylated to GTP — providing an inexpensive, sustainable route to GTP without adding GMP.
Translation Mix (Amino Acids)
- 17 Amino Acid Mix — Balanced stock containing 17 of the 20 standard amino acids, providing the building blocks the ribosome incorporates into the nascent protein; tyrosine and cysteine are supplied separately, and glutamate is already present in the buffer salts.
- Tyrosine — Added separately (prepared at pH 12) because of its very low solubility at neutral pH, which would cause precipitation in the main mix; supplies the tyrosine residues needed by the ribosome during translation.
- Cysteine — Added separately and fresh because free cysteine oxidizes rapidly, forming disulfide-linked cystine that cannot be used for translation; provides cysteine residues critical for disulfide bonds and active sites in many proteins.
Additives
- Nicotinamide — Precursor of NAD⁺/NADH, the essential redox cofactor that sustains glycolysis (particularly the GAPDH step) and other energy-regenerating reactions, keeping the NAD⁺ pool replenished throughout the 20-hour reaction.
Backfill
- Nuclease Free Water — Ultrapure water treated to remove RNases and DNases, used to bring the reaction to its final volume; it must be nuclease-free to prevent degradation of the DNA template and newly synthesized mRNA, which would halt transcription and translation.
Table 1. Role of each reagent in the 20-hour cell-free master mix.
| Category | Reagent | What it does |
|---|---|---|
| Biological core | E. coli BL21(DE3) Star Lysate | Provides ribosomes, tRNAs, translation factors, metabolic enzymes, and T7 RNA polymerase — the full machinery for transcription + translation |
| Salts / Buffer | Potassium Glutamate | Supplies K⁺ for ribosome assembly and translation fidelity; glutamate mimics the intracellular environment |
| HEPES-KOH pH 7.5 | Biological buffer that keeps the reaction at physiological pH (~7.5) so enzymes stay active | |
| Magnesium Glutamate | Supplies Mg²⁺, essential cofactor for RNA polymerase, ribosomes, and nucleotide-dependent enzymes; stabilizes RNA | |
| Potassium Phosphate Monobasic (KH₂PO₄) | Provides inorganic phosphate (Pᵢ) for energy regeneration; part of secondary phosphate buffer | |
| Potassium Phosphate Dibasic (K₂HPO₄) | Basic half of the phosphate pair; together with monobasic maintains pH and supports phosphorylation | |
| Energy / Nucleotides | Ribose | Sugar backbone for nucleotide biosynthesis; converted to PRPP, which combines with bases to form NTPs |
| Glucose | Main energy source; feeds glycolysis to regenerate ATP throughout the reaction | |
| AMP | Precursor of ATP; lysate kinases phosphorylate it: AMP → ADP → ATP | |
| CMP | Precursor of CTP; supplies cytidine residues for mRNA synthesis | |
| GMP | Would be the precursor of GTP, but is set to 0 in this mix — replaced by guanine + salvage pathway | |
| UMP | Precursor of UTP; supplies uridine residues for mRNA synthesis | |
| Guanine | Free base converted to GMP via HGPRT + PRPP (salvage pathway), then phosphorylated to GTP | |
| Translation | 17 Amino Acid Mix | 17 of the 20 amino acids, the building blocks for protein synthesis (Tyr, Cys, and Glu added separately) |
| Tyrosine | Added separately because of low solubility at neutral pH (prepared at pH 12) | |
| Cysteine | Added separately and fresh because it oxidizes to cystine; essential for disulfide bonds and active sites | |
| Additives | Nicotinamide | Precursor of NAD⁺/NADH, required for glycolysis (GAPDH step) and energy regeneration |
| Backfill | Nuclease Free Water | Ultrapure water free of RNases/DNases used to bring the reaction to final volume without degrading nucleic acids |
2. Describe the main differences between the 1-hour optimized PEP-NTP master mix and the 20-hour NMP-Ribose-Glucose master mix shown in the Google Slide above. (2-3 sentences) The 1-hour PEP-NTP mix already contains the full nucleotides (ATP, GTP, CTP, UTP) and uses PEP as a direct energy source, so the reaction starts fast but ends fast too, and it is expensive. The 20-hour NMP-Ribose-Glucose mix is different because it only provides cheap precursors — NMPs (AMP, CMP, UMP), free guanine, ribose and glucose — and the enzymes in the E. coli lysate are the ones that build the NTPs and regenerate ATP through glycolysis. So the main idea is that the 1-hour mix works fast but is short and expensive, while the 20-hour mix is slower but lasts much longer and is cheaper, which is better for producing fluorescent proteins at a large scale.
3. Bonus question: How can transcription occur if GMP is not included but Guanine is? Transcription can still happen because the E. coli lysate has the enzymes of the purine salvage pathway, so it can make GTP from guanine without needing GMP. First, the ribose in the mix is converted into PRPP, and then the enzyme HGPRT joins guanine + PRPP to make GMP directly inside the reaction. After that, two kinases (guanylate kinase and NDP kinase) add more phosphates to transform GMP → GDP → GTP. This GTP is then used by the T7 RNA polymerase to add guanine to the mRNA during transcription. Adding guanine instead of GMP makes the reaction much cheaper, which is important when many reactions are running at the same time.
Part C: Planning the Global Experiment | Cell-Free Master Mix Design
1. Given the 6 fluorescent proteins we used for our collaborative painting, identify and explain at least one biophysical or functional property of each protein that affects expression or readout in cell-free systems. (Hint: options include maturation time, acid sensitivity, folding, oxygen dependence, etc) (1-2 sentences each). The amino acid sequences are shown in the HTGAA Cell-Free Benchling folder.
- sfGFP (superfolder GFP): sfGFP folds and matures very fast (~14 min) and is very robust, producing a strong and reliable green signal even in non-ideal cell-free conditions. It still needs oxygen for chromophore maturation, but because of its fast and efficient folding, it is one of the most consistent readouts in cell-free systems.
- mRFP1: mRFP1 has a slow maturation time (~60–90 min) and a strong oxygen dependence, so in a sealed cell-free reaction with limited O₂, the red signal appears late and is relatively dim compared to faster red FPs.
- mKO2 (monomeric Kusabira Orange 2): mKO2 is acid-sensitive (pKa ~5–6), so when the cell-free reaction accumulates organic acids from glycolysis and the pH drops, the orange fluorescence decreases even if the protein is fully expressed.
- mTurquoise2: mTurquoise2 has a very high quantum yield (~0.93) and low oxygen sensitivity, which makes it one of the brightest and most reliable cyan reporters in oxygen-limited cell-free reactions.
- mScarlet-I: mScarlet-I has a faster maturation time (~36 min) than most red FPs and is very bright, although it still requires oxygen for chromophore formation — so it gives a good red readout that is slower than sfGFP but faster than mRFP1.
- Electra2: Electra2 is a blue fluorescent protein (BFP) with brightness comparable to mTagBFP2, and like other GFP-family proteins it needs oxygen to mature its chromophore, so its signal can be limited in sealed cell-free reactions with depleted O₂. Its blue emission (~450 nm) also requires UV/violet excitation, which can increase autofluorescence from the lysate.
Table 2. Properties of the 6 proteins used in the collaborative painting.
| # | Protein | Color / Emission | Key property for cell-free | Reference (DOI) |
|---|---|---|---|---|
| 1 | sfGFP | Green ~510 nm | Fast and robust folding (~14 min) — the most reliable readout in cell-free | Pédelacq et al., Nat Biotechnol 2006 — 10.1038/nbt1172 |
| 2 | mRFP1 | Red ~607 nm | Slow maturation (~60–90 min) + strong O₂ dependence — signal appears late and dim | Campbell et al., PNAS 2002 — 10.1073/pnas.082243699 |
| 3 | mKO2 | Orange ~565 nm | Acid-sensitive (pKa ~5–6) — loses signal if glycolysis drops the pH | Sakaue-Sawano et al., Cell 2008 — 10.1016/j.cell.2007.12.033 |
| 4 | mTurquoise2 | Cyan ~475 nm | Record quantum yield (~0.93) — the brightest and most reliable CFP | Goedhart et al., Nat Commun 2012 — 10.1038/ncomms1738 |
| 5 | mScarlet-I | Red ~593 nm | Improved maturation (~36 min) vs original mScarlet — bright and fast | Bindels et al., Nat Methods 2017 — 10.1038/nmeth.4074 |
| 6 | Electra2 | Blue ~450 nm | Bright BFP (~mTagBFP2 level), O₂-dependent, requires UV/violet excitation | Papadaki et al., Sci Rep 2022 — 10.1038/s41598-022-13214-0 |
2. Create a hypothesis for how adjusting one or more reagents in the cell-free mastermix could improve a specific biophysical or functional property you identified above, in order to maximize fluorescence over a 36-hour incubation. Clearly state the protein, the reagent(s), and the expected effect.
Question 1 — Hypothesis (adjusted to 2 µL custom supplement at t = 0)
Protein: mTurquoise2.
Reagent(s) to adjust: In the 2 µL custom supplement slot, I will add a 10× concentrated stock that contains the 17-amino-acid mix, tyrosine, cysteine, AMP, CMP, UMP, guanine, glucose, and nicotinamide. This will almost double the starting concentration of amino acids and nucleotide precursors compared to the standard 20-hour master mix.
Expected effect: mTurquoise2 is already very bright (quantum yield ~0.93) and not very sensitive to oxygen, so the main limitation during a 36-hour reaction is that the amino acids and NMPs get used up after ~18–20 hours. When that happens, transcription and translation stop, and the fluorescence signal does not grow anymore. If I start the reaction with more of these substrates, protein synthesis should stay active for longer, and the final fluorescence at 36 hours should be higher than in wells that only receive water in the 2 µL custom slot.
Reasoning: Because brightness and folding are not the problem for mTurquoise2, the best way to improve the signal is to keep the reaction running longer. More amino acids help translation, and more NMPs and guanine help the lysate enzymes regenerate NTPs through the salvage and kinase pathways. I am not adding more Mg²⁺ or phosphate because those are already at a good level, and adding more could actually slow the enzymes down. Good controls would be a water-only 2 µL addition (to see the normal plateau) and, if possible, amino-acids-only and NMPs-only supplements, to see which type of substrate is really limiting the reaction.
3. The second phase of this lab will be to define the precise reagent concentrations for your cell-free experiment. You will be assigned artwork wells with specific fluorescent proteins and receive an email with instructions this week (by April 24). You can begin composing master mix compositions here.
Hypothesis:
The fluorescence at 36 h is limited mainly by substrate exhaustion. Adding more amino acids and more nucleotide/energy precursors at t = 0 will keep the TX-TL reaction productive longer and give more final fluorescence.
I customize 8 wells: 4 contiguous mRFP1 wells and 4 scattered sfGFP wells. The same 4 supplement conditions go to both proteins. I chose this pair because they are very different in biophysics, so the same supplement should affect them in a different way, and that is informative by itself.
sfGFP: folds and matures very fast (~14 min) and the folding is robust even when conditions are not perfect (Pédelacq et al., Nat Biotechnol 2006). Maturation needs O₂ but folding is so efficient that O₂ is normally not the bottleneck. → Expression-limited: more time of synthesis = more fluorescence.
mRFP1: maturation is slow (t½ ≈ 60–90 min) and has strong O₂ dependence (Campbell et al., PNAS 2002). Low quantum yield (~0.25), passes through a green intermediate, so if the reaction stops early, much of the protein stays immature. → Expression-limited AND maturation-limited.
A supplement that only helps the synthesis side should help sfGFP a lot but help mRFP1 less.
Why these reagents?
The 20-h base master mix (NMP + ribose, Ginkgo + OpenAI paper) regenerates NTPs from simple precursors. At 36 h I think the limiting things are: (1) free amino acids, especially aromatics and Cys; (2) the NMP pool (AMP, CMP, UMP) and guanine, used to regenerate NTPs through salvage and kinase pathways; (3) NAD⁺/NADH turnover (nicotinamide as precursor), to keep glycolysis active past 20 h.
I do not change salts (K-glutamate, Mg-glutamate), buffer (HEPES, K-phosphate), lysate volume or DNA template. Changing ionic strength, pH or Mg²⁺ would test many things at the same time.
Experimental design — 2×2 factorial
The same 4 conditions go to 4 sfGFP wells and 4 mRFP1 wells.
| Well | Condition | 2 µL custom supplement |
|---|---|---|
| W1 | Control | Only water |
| W2 | +AA | +25% boost of 17-AA mix, Tyr, Cys |
| W3 | +Energy | ~2× AMP, CMP, UMP, guanine; +50% nicotinamide; +2× glucose; small ribose top-up |
| W4 | +AA + Energy | W2 and W3 together |
Target final concentrations (20 µL reaction)
| Reagent | W1 | W2 | W3 | W4 |
|---|---|---|---|---|
| 17 AA mix | 4.063 mM | 5.06 | 4.063 | 5.06 |
| Tyrosine pH 12 | 4.063 mM | 5.06 | 4.063 | 5.06 |
| Cysteine | 4.000 mM | 5.00 | 4.000 | 5.00 |
| AMP | 0.625 mM | 0.625 | 1.25 | 1.25 |
| CMP | 0.375 mM | 0.375 | 0.75 | 0.75 |
| UMP | 0.375 mM | 0.375 | 0.75 | 0.75 |
| Guanine | 0.156 mM | 0.156 | 0.31 | 0.31 |
| Nicotinamide | 3.125 mM | 3.125 | 4.625 | 4.625 |
| Glucose | 1.250 g/L | 1.250 | 2.50 | 2.50 |
| Ribose | 11.625 g/L | 11.625 | 12.625 | 12.625 |
All other reagents (salts, buffer, phosphates) stay at base.
Reaction (same for every well) 6 µL lysate (E. coli BL21 Star DE3) + 10 µL 2× Master Mix + 2 µL DNA template (20 ng/µL) + 2 µL custom supplement (W1–W4) = 20 µL total.
The only variable across the 8 wells is the supplement and the FP. Any difference should come from the supplement.
Predictions
sfGFP: clear order W4 > W2 ≈ W3 > W1, effects of AA and energy more or less additive. Every extra hour of active translation gives proportionally more fluorescence.
mRFP1: same order but smaller magnitude and possible saturation: W4 ≥ W2 ≥ W3 > W1. If the W4-over-W1 effect in mRFP1 is much smaller than in sfGFP, the limit is chromophore maturation, not substrates, and the 36-h design should focus on O₂ (well geometry, lysate dilution) instead of more substrate.
Well selection and submission on the rcdonovan.com platform
After deciding the experimental design, I went to my personal link in the Donovan platform and assigned the 8 wells. I picked the wells in the following way and the four conditions W1 (control), W2 (+AA), W3 (+Energy) and W4 (+AA+Energy) were assigned across the wells of each protein, so each protein has all four supplement conditions represented.
I contributed with 8 wells, four per proteine, two per condition.
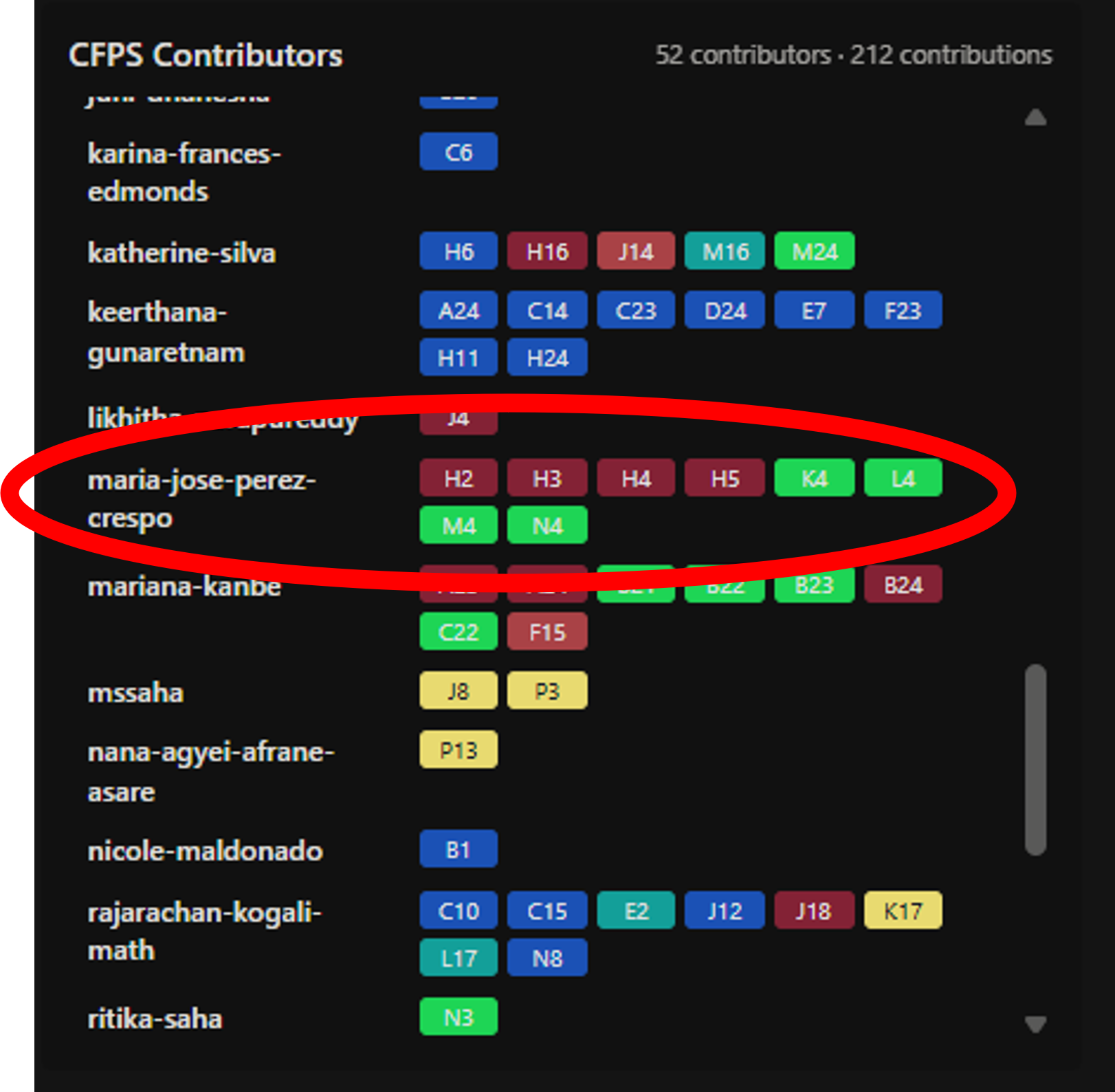
Figure 5: My contribution on the rcdonovan.com platform — 8 wells assigned to my HTGAA username (2026a-maria-jose-perez-crespo), with the four 2×2 supplement conditions distributed between mRFP1 (H2–H5) and sfGFP (K4–N4).
- mRFP1 (red, 4 wells): I chose 4 contiguous wells in a horizontal row at Q1: H2, H3, H4, H5. I chose 4 contiguous wells in a horizontal row at Q1: H2, H3, H4, H5. I selected them contiguous so I can easily see and visualize the effect of the supplement modifications across the four conditions (W1–W4) directly on the artwork.
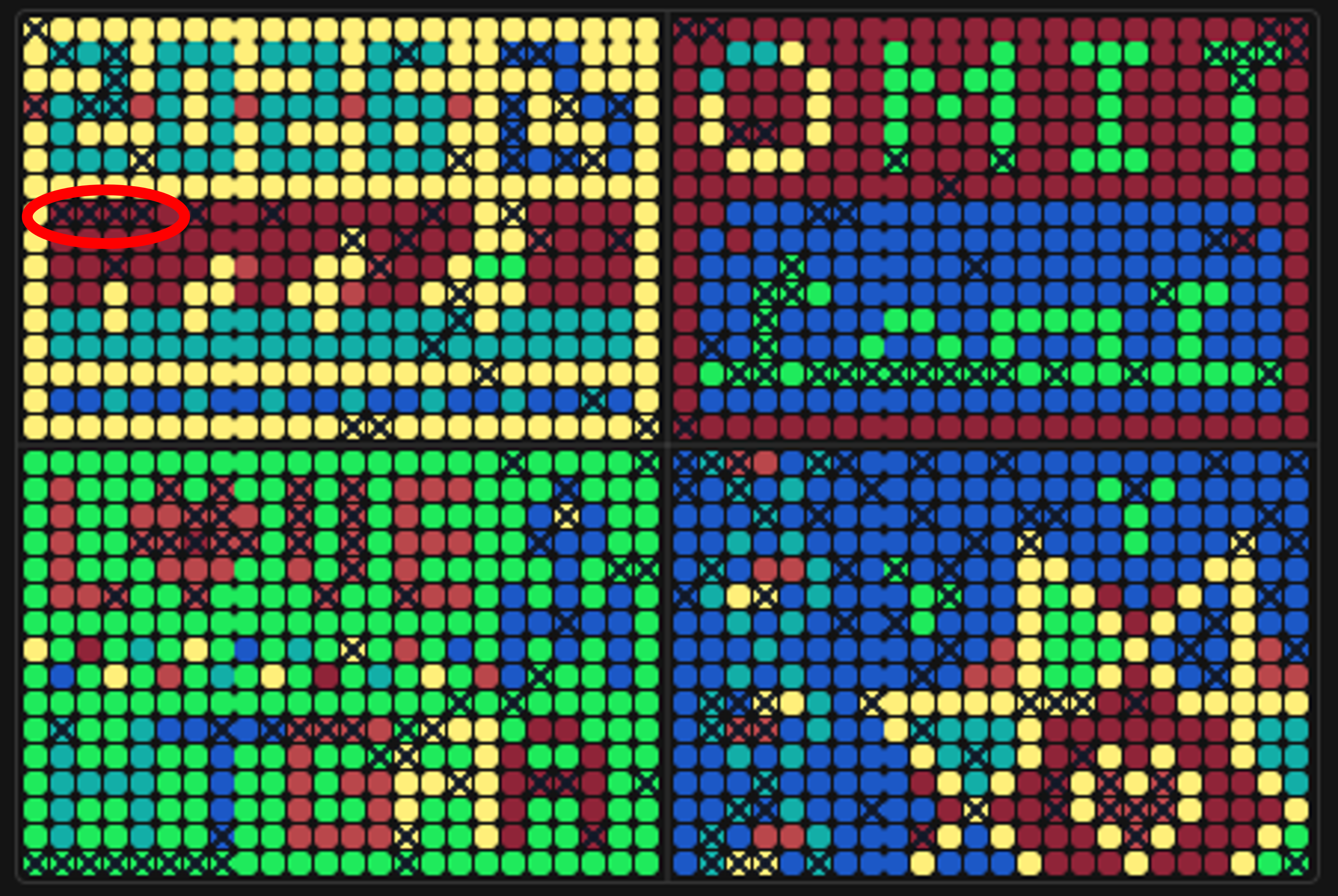
Figure 6. Position of the mRFP1 wells (H2–H5) in quadrant Q1 of the artwork canvas.
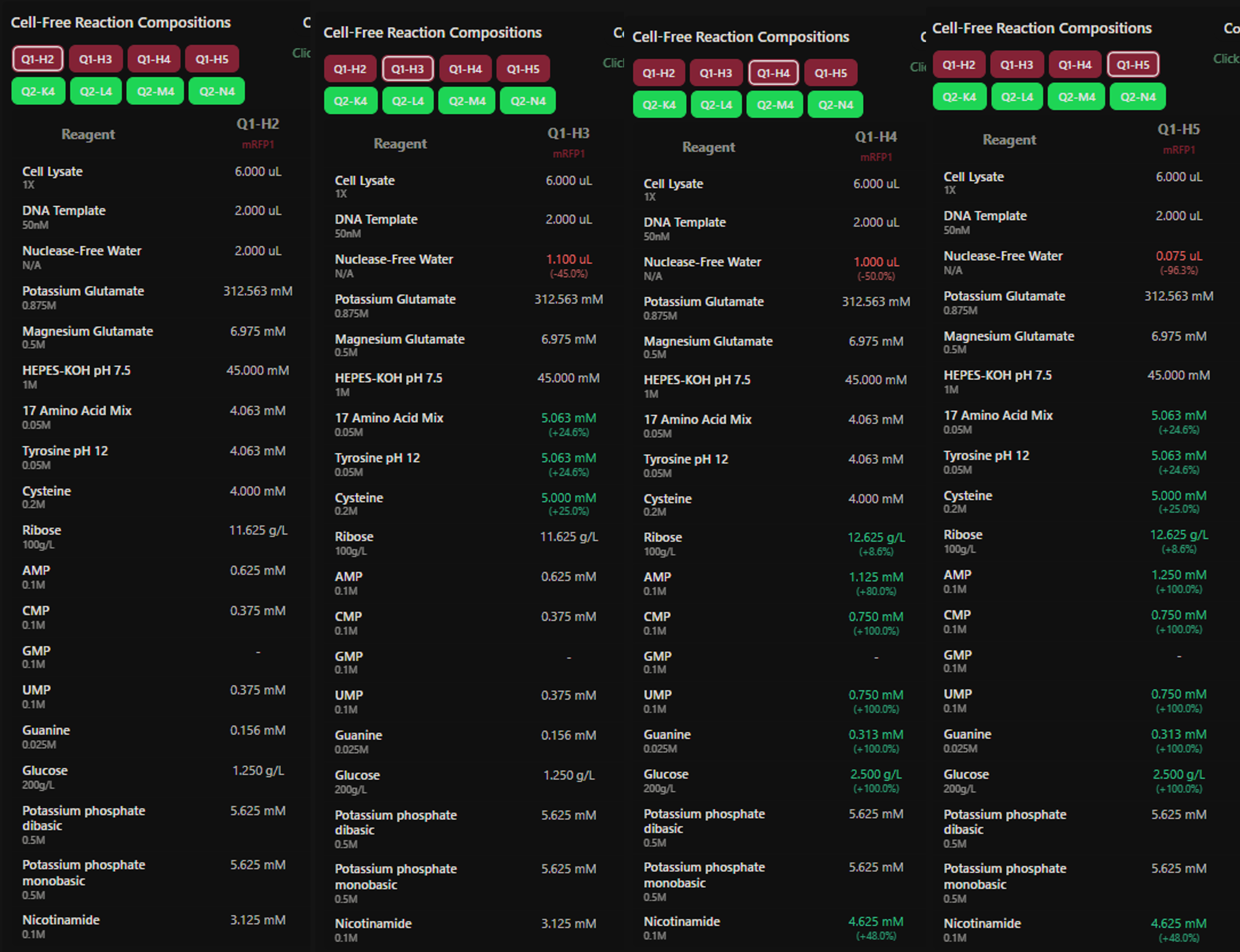
Figure 7: Custom supplement mix assigned to the mRFP1 wells (H2–H5), one condition per well (W1 control, W2 +AA, W3 +Energy, W4 +AA+Energy).
- sfGFP (green, 4 wells): the green wells assigned to me were scattered across the artwork, so I picked 4 of them in a vertical line at Q2: K4, L4, M4, N4 to easyly visualize the final effect.
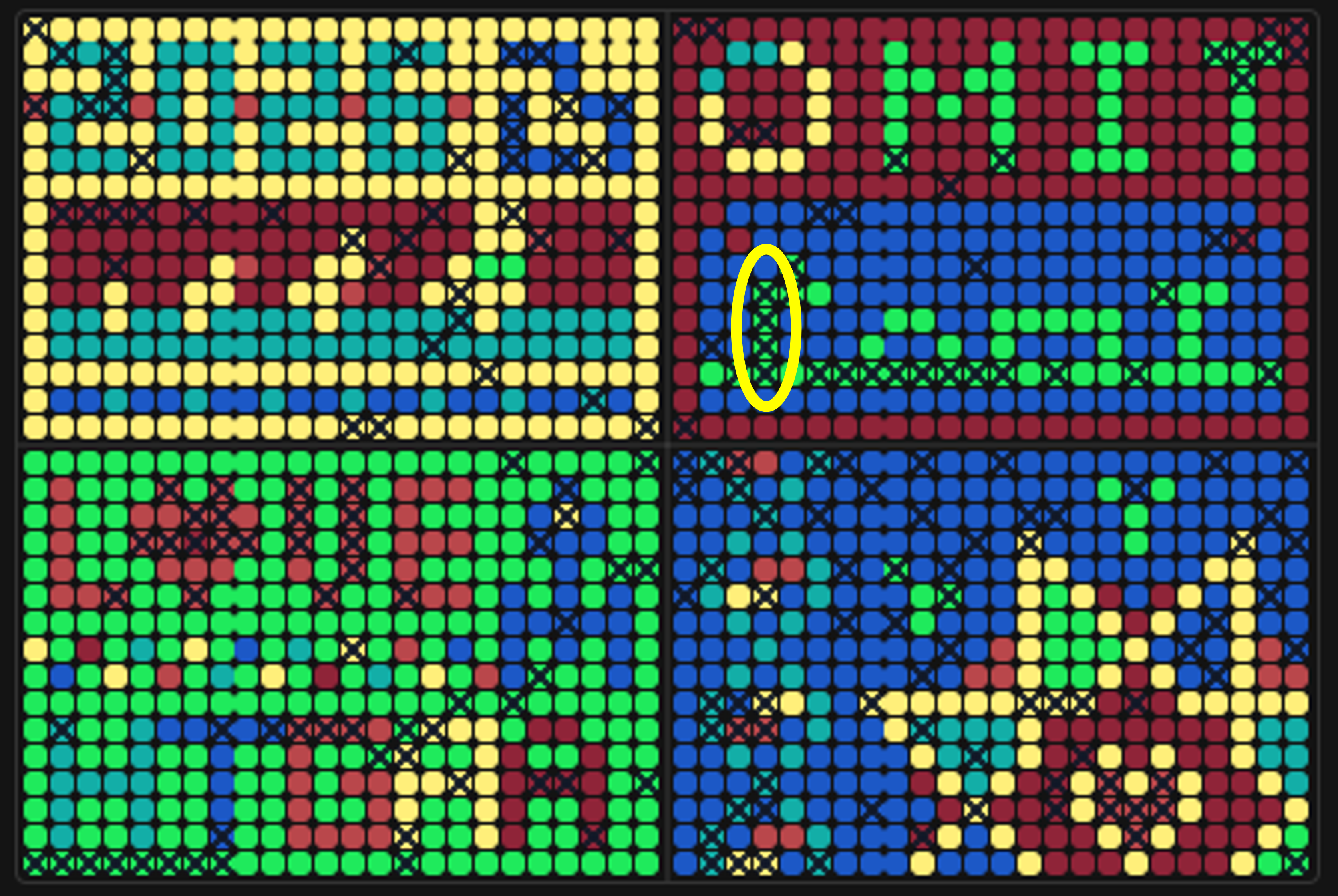
Figure 8. Position of the sfGFP wells (K4-N4) in quadrant Q2 of the artwork canvas.
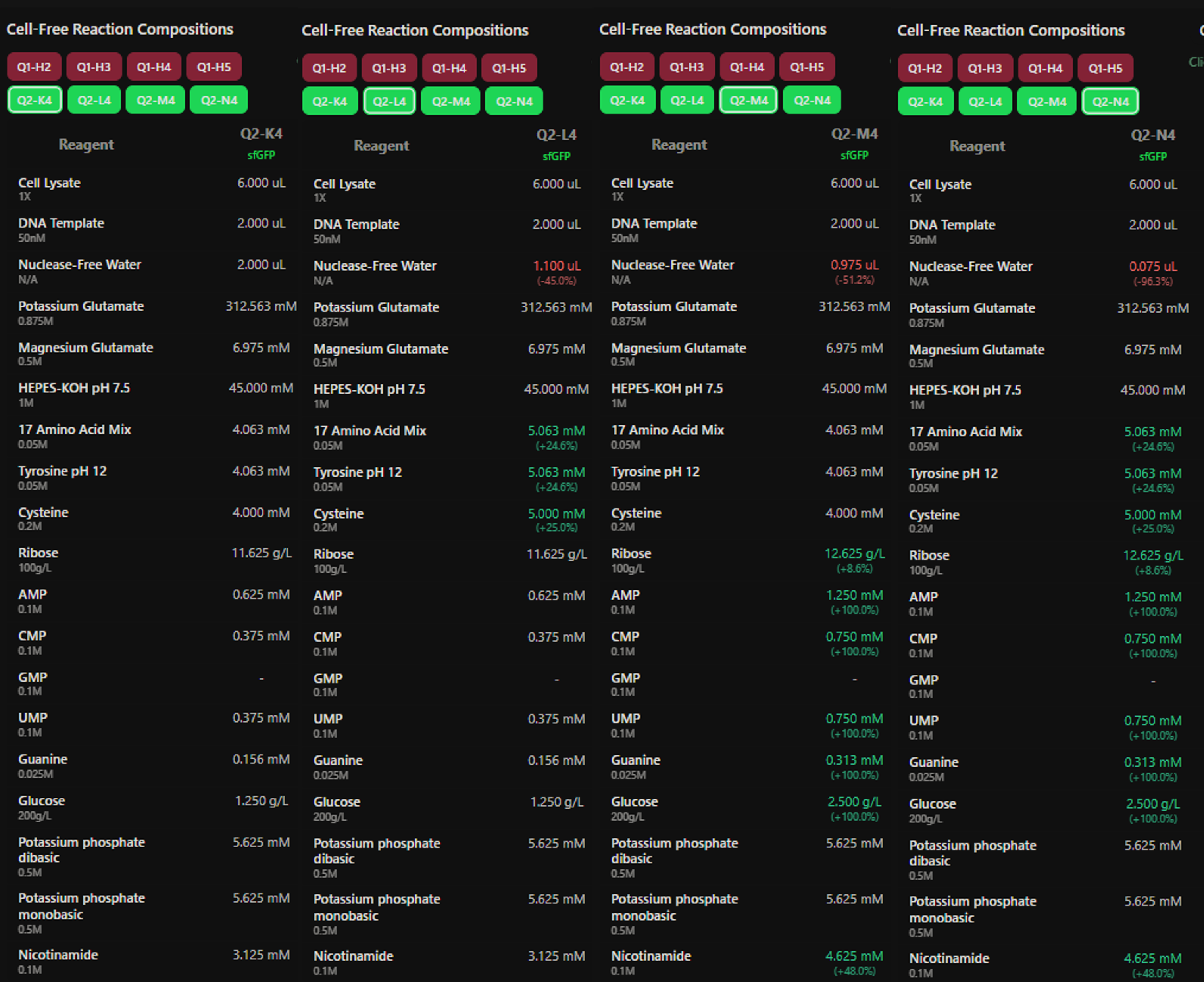
Figure 9: Custom supplement mix assigned to the sfGFP wells (K4-N4), one condition per well (W1 control, W2 +AA, W3 +Energy, W4 +AA+Energy).
PENDING PENDING
- The final phase of this lab will be analyzing the fluorescence data we collect to determine whether we can draw any conclusions about favorable reagent compositions for our fluorescent proteins. This will be due a week after the data is returned (date TBD!). The reaction composition for each well will be as follows:
6 μL of Lysate 10 μL of 2X Optimized Master Mix from above 2 μL of assigned fluorescent protein DNA template 2 μL of your custom reagent supplements Total: 20 μL reaction
Plan of analysis
For each protein:
- Background correction: subtract the mean of empty/dropout wells in the same plate region.
- Main effects and interaction:
- AA effect = mean(W2, W4) − mean(W1, W3)
- Energy effect = mean(W3, W4) − mean(W1, W2)
- Interaction = (W4 − W3) − (W2 − W1); ≈ 0 means additive, > 0 means synergy.
- Compare effect size between the two proteins: big effect in sfGFP and small in mRFP1 would support that mRFP1 is limited after the synthesis (folding, maturation, O₂) and not by substrates.
- If kinetic data is returned, fit a simple synthesis-then-plateau model per well, to separate “how long the reaction was productive” from “how much protein accumulated per unit time”.
Why this is useful for the global experiment?
Pairing a synthesis-limited reporter (sfGFP) with a maturation-limited reporter (mRFP1) under the same factorial perturbation separates effects on TX-TL throughput from effects on post-translational maturation — exactly the distinction the global 36-h master mix optimization has to make.
References
- Pédelacq et al. Nat Biotechnol (2006) — sfGFP. DOI: 10.1038/nbt1172
- Campbell et al. PNAS (2002) — mRFP1. DOI: 10.1073/pnas.082243699
- Ginkgo + OpenAI cell-free protein synthesis paper — base of the 20-h NMP-ribose master mix.
Part D: Build-A-Cloud-Lab | (optional) Bonus Assignment
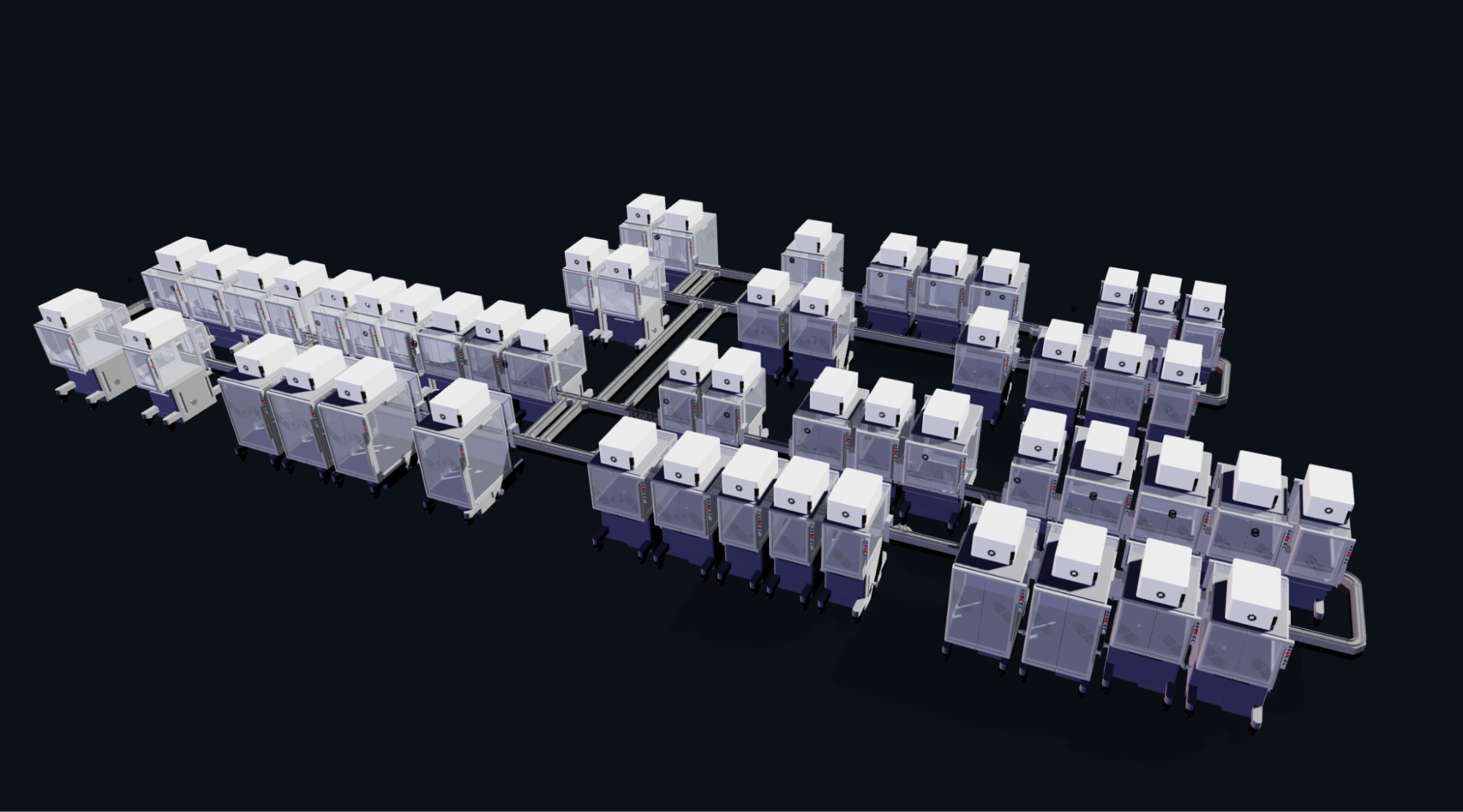

Figure 3. Cell-Free master mix compositions.
Use this simulation tool to create an interesting looking cloud lab out of the Ginkgo Reconfigurable Automation Carts. This is just a minimal implementation so far, but I would love to see some fun designs!