Documentation.
The Plastisphere Code:
Decoding How Microbes Stick to Synthetic Polymers
Section 1: Abstract
Microplastic (MP) pollution is one of the most important environmental problems today. Recent studies have shown that bacteria can colonize plastic surfaces forming biofilms known as the “plastisphere”, which behave very differently from surrounding microbial communities (Di Pippo et al., 2020; Zhai et al., 2023). The impact of this colonization is not only ecological: plastisphere biofilms can act as vectors for pathogens and antibiotic-resistance genes, with Pseudomonas and Bacillus often identified as dominant hosts of resistance genes in these environments (Li et al., 2025). In addition to bacteria, more than 200 fungal species have also been reported to colonize and degrade synthetic plastics, showing that microbial adhesion to polymers is a broad and still poorly understood phenomenon (Ekanayaka et al., 2022). At the molecular level, bacterial attachment to synthetic polymers is largely mediated by surface proteins such as curli amyloids (CsgA) and other adhesins, but the structural and physicochemical features that determine why specific proteins bind plastic surfaces are still not well understood (Sano et al., 2023).
The general objective of this project is to use computational tools to identify and characterize microbial adhesion proteins that interact with synthetic polymers. The working hypothesis is that physicochemical and structural traits — surface hydrophobicity, charge distribution, and conserved adhesion motifs — can predict whether a protein binds plastic surfaces. Two specific sub-hypotheses are tested: (H1) three structurally distinct amyloid adhesins (CsgA, FapC, TasA) will show convergent binding modes against the same polymer despite their different folds; and (H2) all three will display stronger interactions with polystyrene than with polyethylene due to π-π stacking with aromatic residues. These features can then guide the rational design of bio-adhesive systems.
The project is organized into three specific aims. Aim 1 (in silico) computationally identifies and characterizes a panel of three bacterial amyloid adhesion proteins — CsgA from Escherichia coli, FapC from Pseudomonas fluorescens and TasA from Bacillus subtilis — that interact with polyethylene, combining structure retrieval from the AlphaFold Protein Structure Database (Jumper et al., 2021), surface analysis in PyMOL, and protein–polymer complex prediction with the AlphaFold 3 web server (Abramson et al., 2024); in parallel, three E. coli expression cassettes are designed in silico and validated for Twist Bioscience synthesis. Aim 2 (development) uses the residue-level adhesion map from Aim 1 to design anti-adhesion peptides with PepMLM (Chen et al., 2025), moPPIt and PeptiVerse, with structural validation in AlphaFold 3 prior to in vitro testing on PE and polystyrene surfaces, using E. coli strains transformed with the expression cassettes designed in Aim 1 to produce each adhesin in a controlled biofilm-formation assay.Aim 3 (visionary) extends the framework to clinically relevant biofilm proteins such as Bap from Staphylococcus aureus and to programmable applications including anti-adhesion peptides for medical devices, non-toxic antifouling coatings, enhanced microbial degradation of microplastics by enzymes such as PETase (Austin et al., 2018), engineered living materials and diagnostic platforms.
Section 2 — Project Aims
Aim 1 — Experimental Aim (this project)
The first aim of this project is to identify and computationally characterize a panel of three bacterial amyloid adhesion proteins — CsgA from Escherichia coli, FapC from Pseudomonas fluorescens, and TasA from Bacillus subtilis — that interact with synthetic polymer surfaces. The mature chain of each protein (signal peptide removed) is retrieved from the AlphaFold Protein Structure Database, and solvent-exposed hydrophobic residues are identified in PyMOL using per-residue SASA filtering and visual inspection of the surface. Protein–polymer complexes are then predicted with the AlphaFold 3 web server (alphafoldserver.com), using palmitate (PLM, a 16-carbon hydrocarbon ligand) as a proxy for polyethylene (PE), since the public server does not accept arbitrary SMILES strings. The top-ranked complex is analyzed in PyMOL to identify protein residues within 5 Å of the ligand, producing a residue-level adhesion map per protein. In parallel, three E. coli expression cassettes (one per adhesin) are designed in silico using the IDT Codon Optimization Tool and assembled with standard iGEM BioBrick parts (BBa_J23106 promoter, BBa_B0034 ribosome binding site, BBa_B0015 terminator), validated against Twist Bioscience synthesis specifications. Two sub-hypotheses are tested with this aim: (H1) the three structurally distinct amyloid adhesins will show convergent binding modes against the same polymer despite having different folds; and (H2) all three proteins will display stronger interactions (higher ipTM scores) with polystyrene than with polyethylene, due to π–π stacking between aromatic residues (Phe, Tyr, Trp) and the phenyl groups of polystyrene.
Aim 2 — Development Aim (next step beyond this course)
Building on the residue-level adhesion map produced in Aim 1, the next step is the rational design of anti-adhesion peptides able to block bacterial colonization of synthetic polymers, using PepMLM to generate candidate binders from each adhesin sequence, moPPIt to optimize peptide binding toward the specific adhesion-critical residues identified in Aim 1, and PeptiVerse to evaluate therapeutic-relevant properties (binding affinity, solubility, net charge, hemolysis probability), with structural validation of the predicted peptide–adhesin complexes through the AlphaFold 3 web server. The most promising lead peptides will be advanced to in vitro biofilm-formation assays on PE and polystyrene surfaces, using E. coli strains transformed with the expression cassettes designed in Aim 1 to produce each adhesin and quantifying biofilm formation with crystal violet staining in the presence and absence of the candidate peptides.
Aim 3 — Visionary Aim (long-term vision)
The long-term vision of this project is to enable rational, residue-level control of microbial adhesion to synthetic polymers, transforming biofilm formation on plastics from an unavoidable phenomenon into an engineerable interface between living and non-living matter. As a natural extension, the same framework would be applied to additional clinically relevant biofilm proteins, such as Bap from Staphylococcus aureus, to address hospital-acquired biofilm infections on indwelling medical devices. Programmable applications of this approach include anti-adhesion peptides for catheters, implants and food-contact surfaces, non-toxic antifouling coatings, enhanced microbial degradation of microplastics through bioremediation, engineered living materials with controlled cell–surface attachment, and diagnostic platforms based on selective pathogen capture.
Section 3. BACKGROUND
3.1. Literature Context
Two recent peer-reviewed studies illustrate both the relevance and the current limitations of microbial adhesion research on synthetic polymers. Li et al. (2025) characterized the mangrove plastisphere as a hotspot for high-risk antibiotic-resistance genes (ARGs) and pathogenic bacteria, showing that polyethylene (PE), polystyrene (PS) and polyvinyl chloride (PVC) enrich virulence and resistance determinants compared to surrounding sediments, with Pseudomonas and Bacillus identified as dominant hosts of these genes. Their findings demonstrate that microbial colonization of microplastics (MPs) is not a neutral ecological event but an active vector for antimicrobial resistance and pathogen dissemination, reinforcing the urgent need to understand which microbial proteins drive this colonization. From a molecular perspective, Sano et al. (2023) provided a high-resolution view of one such adhesion system: they identified CsgI (YccT) as a periplasmic inhibitor of curli fimbriae formation in Escherichia coli, demonstrating that the polymerization of CsgA — the main amyloid component of curli involved in solid-surface adhesion and biofilm formation — can be modulated through the EnvZ/OmpR two-component system and through direct inhibition of monomer assembly. Together, these two studies highlight a clear gap in the field: while the plastisphere is increasingly recognized as a public-health-relevant microbial niche, and while individual adhesion proteins such as CsgA are now structurally and functionally well described, no systematic computational framework currently links the structural and physicochemical features of microbial adhesion proteins to their ability to bind specific synthetic polymer surfaces, which is precisely the gap that the present project addresses.
3.2. Innovation
This project introduces three innovative contributions to the study of microbial adhesion on synthetic polymers:
It applies a comparative computational approach to three structurally different amyloid adhesins — CsgA, FapC and TasA — from distinct bacterial genera. This comparison aims to identify common physicochemical and structural features involved in adhesion to plastic surfaces. Although functional amyloids and plastisphere colonization have been studied separately, a systematic cross-genus comparison focused specifically on plastisphere-relevant amyloid adhesins remains limited.
The project repurposes recent generative machine-learning tools originally developed for therapeutic peptide design (PepMLM, moPPIt and PeptiVerse) for an environmental biotechnology context. In this way, the work explores how models created for drug discovery can be transferred to materials science applications and biofilm control strategies.
The project challenges the traditional antibiotic-based paradigm of microbial control by proposing inhibition of surface attachment at the residue level rather than bacterial killing. By acting before stable colonization occurs, this strategy is expected to reduce selective pressure on microbial viability and therefore lower the risk of resistance development, which represents a major limitation of current antimicrobial approaches.
Overall, the project expands the scope of synthetic biology by combining comparative structural microbiology, generative artificial intelligence and anti-adhesion design within a framework oriented toward environmental biofilm management.
3.3. Significance
Microplastic pollution and antimicrobial resistance are two of the biggest global problems today for public health and the environment, and the plastisphere sits at the intersection of both: plastic surfaces in ecosystems act as platforms where pathogenic bacteria and antibiotic-resistance genes accumulate (Li et al., 2025). This project addresses a critical gap in this field — the lack of a systematic, residue-level understanding of which bacterial proteins drive plastic colonization and how this colonization could be modulated without antibiotics. The proposed computational pipeline integrates structural prediction, protein–polymer docking and generative peptide design, advancing scientific capability across structural microbiology, environmental biotechnology and synthetic biology, three fields that currently work in isolation.
Beyond the immediate research context, anti-adhesion strategies derived from this framework could have several real-world impacts:
Clinical practice: reducing biofilm-driven hospital-acquired infections on catheters, implants and food-contact surfaces, a problem responsible for substantial mortality, morbidity and healthcare costs worldwide.
Environmental biotechnology: limiting the dissemination of antimicrobial resistance through plastic waste.
Bioremediation: the same framework could be reversed to enhance microbial adhesion to microplastics, accelerating their degradation by enzymes such as PETase (Austin et al., 2018).
If the aims of this project are achieved, the results would provide a starting point for understanding microbial adhesion to plastics at the molecular level, and could support future research on non-antibiotic strategies to control biofilm formation.
3.4. Bioethical Considerations
This project has several ethical implications that should be openly discussed. The framework can be used in two opposite ways: it can block microbial adhesion to plastics, or it can enhance it. This makes the project a clear example of dual-use research, because the same design principles that prevent biofilm formation could also be misused to create microbes with stronger colonization capacities. Four ethical principles are particularly relevant to this work:
- Non-maleficence: Anti-adhesion strategies block bacterial attachment instead of killing the bacteria, so they should not produce the selective pressure that drives antibiotic resistance.
- Beneficence: Reducing biofilm infections on medical devices could save lives, and improving microplastic bioremediation could protect the environment.
- Justice: Biofilm infections affect vulnerable groups more strongly, such as patients in intensive care, elderly people, or individuals with limited access to good healthcare. Any therapy derived from this work should be available in a fair way and should not increase existing health inequalities.
- Responsibility: The dual-use nature of the methodology requires careful evaluation of potential applications, with input from biosafety and bioethics experts.
To make sure the project is conducted in a responsible way, several measures are proposed below.
Proposed actions:
- The work will remain at the computational level during this course.
- All generated data, code and structural models will be shared in open repositories, so that other researchers can verify the results.
- Any future experimental validation will be done under standard biosafety conditions and using only non-pathogenic laboratory strains.
- Any potential release of engineered organisms into the environment would require a previous ecological risk assessment and regulatory approval.
Assumptions that may not be fully correct:
- Computational predictions may not fully describe how proteins behave in real biological conditions.
- Polymer surfaces in real environments are not flat and clean as in the models, but weathered, broken and covered by other biological material.
- Microbial communities are very adaptive and may find alternative ways to attach to plastics, even if one specific protein is blocked.
Possible unintended consequences:
- Bacteria could develop adhesion variants that resist the designed peptides.
- The peptides could have off-target effects on human or commensal proteins.
- Beneficial environmental biofilms could be disrupted if interventions are used without caution.
Alternative strategies that should not be replaced by this work:
- Better plastic waste management and waste reduction at the source.
- Antimicrobial stewardship programmes.
- Physical surface modifications, such as nanotopography.
- Phage-based biofilm control.
From a public-health perspective, this project is relevant because it addresses the overlap between hospital-acquired infections, antimicrobial resistance and environmental contamination by microplastics, which are three problems considered priority areas by major public-health and biomedical research organizations.
Section 4. EXPERIMENTAL DESIGN, TECHNIQUES, TOOLS, AND TECHNOLOGY
4.1. Plan
This project is primarily computational, with two complementary outputs: (i) an in silico characterization of three bacterial amyloid adhesion proteins interacting with synthetic polymers, and
(ii) an in silico DNA design of three E. coli expression cassettes that will support future in vivo validation. The work is organized into seven sub-aims. A schematic workflow figure (Figure 1) summarizes the full pipeline.
Sub-aim 1.1 - Protein selection and structural retrieval
- Retrieve protein sequences from UniProt:
- CsgA: P28307 (Escherichia coli K-12)
- FapC: P0DXF5 (Pseudomonas fluorescens)
- TasA: P54507 (Bacillus subtilis 168)
Figure 1. Top panel: the two synthetic polymer ligands used in this work, polyethylene (PE, right) modelled as a 16-carbon aliphatic chain, and polystyrene (PS, left) modelled as a 4-styrene oligomer with phenyl rings, both rendered as sticks with carbon atoms in grey and hydrogens implicit. Bottom panel: the three target bacterial amyloid adhesion proteins, CsgA, FapC and TasA , shown as cartoons of their AlphaFold-predicted structures, with β-strands highlighted in marine and α-helices in firebrick. CsgA and FapC display repetitive amyloid architectures, while TasA shows a more compact globular β-sheet fold.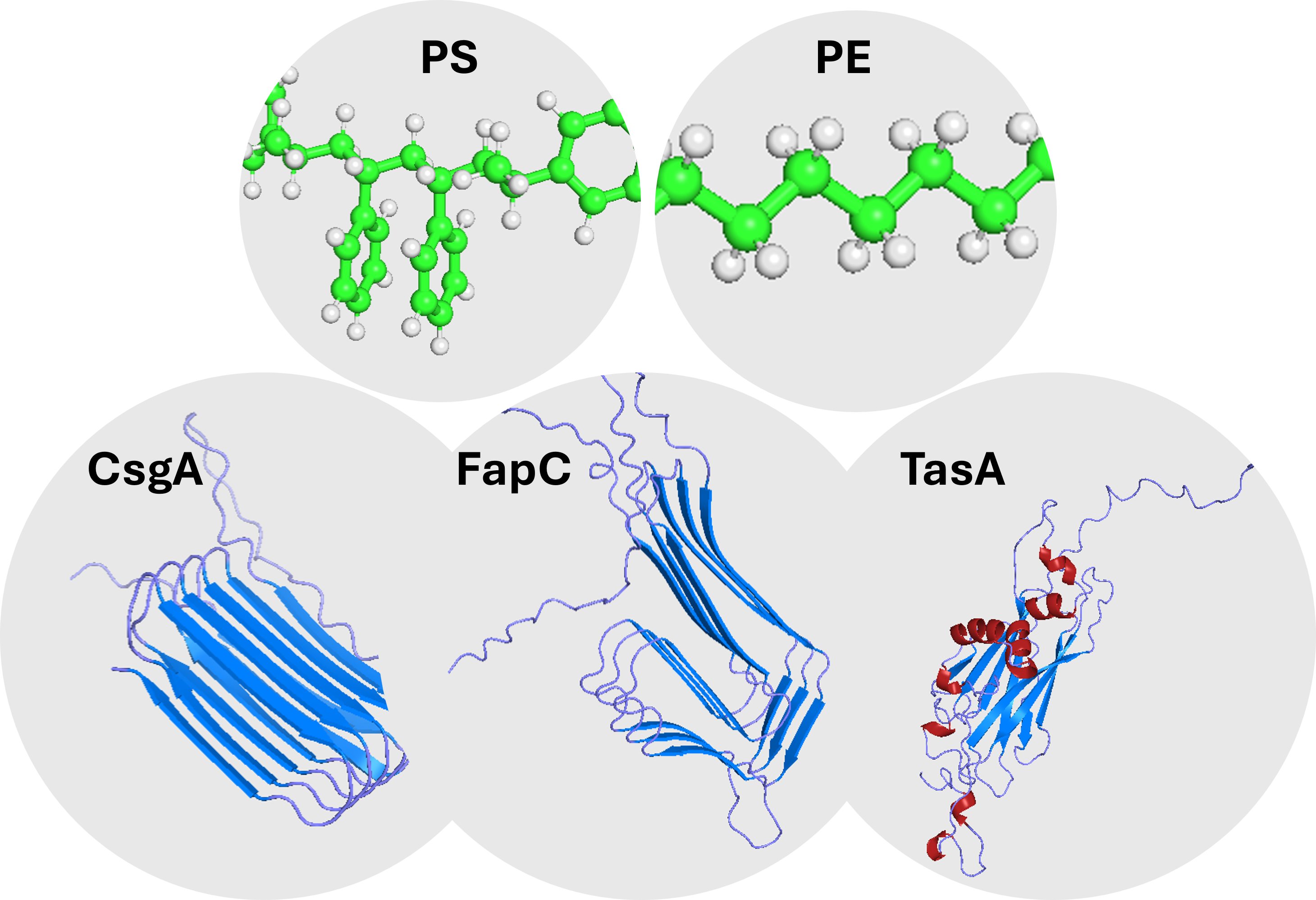
- Note signal peptide and mature chain boundaries for each protein.
- Download AlphaFold-predicted structures from the AlphaFold Protein Structure
- Database (.pdb format).
- Inspect per-residue confidence (pLDDT); regions with pLDDT < 70 are considered flexible/disordered (typical of amyloids).
Expected output: three annotated .pdb files with mature chain boundaries.
Sub-aim 1.2 — Surface analysis in PyMOL
Open each .pdb file in PyMOL.
- Calculate per-atom solvent accessible surface area (SASA) with “get_area, load_b=1”.
- Filter for hydrophobic side-chain residues (Ala, Val, Leu, Ile, Met, Phe, Trp, Tyr) with β-carbon SASA > 5 Ų.
- Curate the list to mature chain only and select 15 residues per protein, prioritizing aromatic residues (key for π-π stacking with PS) and the most exposed hydrophobic positions.
- Save annotated surface images.
Expected output: three lists of 15 candidate adhesion residues, ready as input for docking interpretation.
Sub-aim 1.3 — Protein–polymer docking with AlphaFold 3 and Boltz-1
Predict protein–polymer complexes with two independent tools:
- AlphaFold 3 web server (alphafoldserver.com): mature protein chain + palmitate (PLM, CCD code) as a 16-carbon proxy for polyethylene (PE), since the public server does not accept arbitrary SMILES strings.
- Boltz-1 (via ColabFold-style Colab notebook): mature protein chain + custom SMILES of polystyrene (PS) oligomer: CC(c1ccccc1)CC(c1ccccc1)CC(c1ccccc1)CC(c1ccccc1)C
Record ipTM and pTM confidence scores for the top-ranked model in each case.
Expected output: six predicted complexes (3 proteins × 2 polymer proxies) with ipTM/pTM scores.
Sub-aim 1.4 — Contact identification and visualization
For each top-ranked complex, open the .cif file in PyMOL and identify protein residues within 5 Å of any ligand atom using: “select ligand, not polymer, select contacts, byres (polymer within 5 of ligand), iterate contacts and name CA, print(resi, resn)”
Apply a standardized visualization preset to ensure that all complexes are rendered with consistent style for direct comparison: “bg_color white, hide everything, show cartoon, polymer, color slate, polymer show sticks, ligand color orange, ligand util.cnc ligand show spheres, contacts and name CA color magenta, contacts and name CA set sphere_scale, 1, contacts set sphere_transparency, 0.2, contacts orient
The protein is shown as a slate-blue cartoon, the ligand as orange sticks with heteroatoms color-coded (oxygen red, nitrogen blue), and the contact residues as semi-transparent magenta spheres at the Cα position. This combination provides a clear visual identification of which protein regions interact with the ligand. Capture the rendered viewport as a screenshot and save it as the corresponding figure for each complex.
Expected output: residue-level adhesion map per protein per polymer combination, with one comparable image per complex.
Sub-aim 1.5 — DNA cassette design (mandatory HTGAA component)
Reuse the CsgA expression cassette previously assembled by the author during HTGAA preparatory tasks as a structural template.
Design two new equivalent cassettes for FapC and TasA:
- Codon-optimize the protein sequence for E. coli K12 using the IDT
- Codon Optimization Tool.
- Assemble the cassette with the same architecture as the CsgA template:
- 5’ element: BBa_J23106 (Anderson constitutive promoter, 35 bp)
- BBa_B0034 ribosome binding site (12 bp Shine-Dalgarno)
- ATG start codon
- Codon-optimized full CDS (signal peptide + mature chain)
- TAA stop codon
- 3’ element: BBa_B0015 (rrnB T1 + T7 TΦ double terminator, 129 bp)
Figure 2. Common architecture of the three E. coli expression cassettes designed for CsgA, FapC and TasA.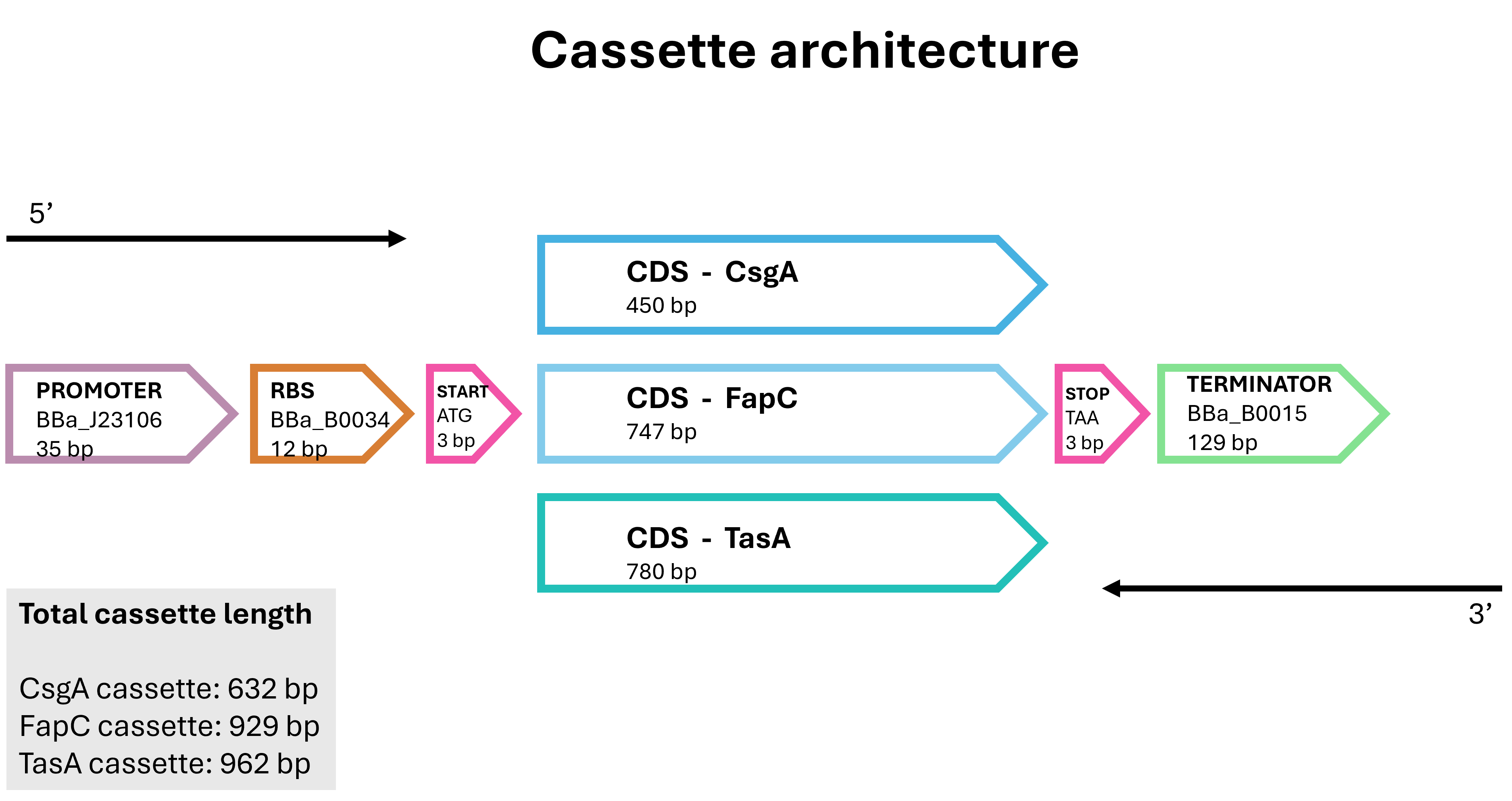
- Validate all cassettes against Twist Bioscience synthesis specifications: – GC content 40–60%. – No repeats longer than 8 bp. – No problematic secondary structures.
Expected output: three Twist-ready cassettes (CsgA already assembled; FapC and TasA designed in this work). These constructs are intended for future synthesis and transformation into E. coli strains, where they will be used in Sub-aim 1.7 (in vivo validation) to express each adhesin and quantify biofilm formation on PE and PS surfaces in the presence and absence of the anti-adhesion peptides designed in Aim 2.
Sub-aim 1.6 — Comparative analysis
- Compile docking results across all 6 protein × polymer combinations into a comparative table (ipTM, pTM, contact residues).
- Test sub-hypothesis H1: do the three proteins share patterns of contact residues against the same polymer?
- Test sub-hypothesis H2: are ipTM scores higher for PS than for PE due to π-π stacking?
- Generate comparative figures (heatmap of ipTM, structural alignment of contacts).
Expected output: integrated comparative analysis testing the two sub-hypotheses.
Sub-aim 1.7 — In vivo validation (future, beyond this course)
The in silico results will be validated experimentally in a future stage of the project, following these steps:
- Synthesis and cloning. Order the three Twist-ready cassettes; clone each into a low-copy plasmid backbone (e.g., pSEVA or pBR322) using Gibson assembly.
- Transformation. Transform each plasmid into E. coli (DH5α for maintenance; an E. coli ΔcsgA strain for CsgA to avoid endogenous interference).
- Constitutive expression. Since the J23106 promoter is constitutive, the corresponding adhesin is expressed without inducer.
- Biofilm assay on plastic surfaces. Inoculate cultures into microtiter plates of PE and PS, incubate 24–48 h at 28 °C without shaking; quantify biofilm formation by crystal violet staining (OD590).
- Anti-adhesion peptide test (linked to Aim 2). Repeat the biofilm assay with synthesized lead peptides; a successful peptide would reduce biofilm formation in a dose-dependent way.
Expected output: a quantitative wet-lab validation of the in silico predictions and an experimental platform for testing anti-adhesion peptides.
4.2. We discussed and practiced various techniques related to synthetic biology throughout the semester. Place a check next to the techniques relevant to your project.
 Figure 3. Techniques relevant to the project.
Figure 3. Techniques relevant to the project.
4.3. Expand upon two techniques you checked in the previous question by describing how you would utilize those techniques in your final project. (min. 4 sentences)
Chassis Selection. Chassis selection means choosing the bacterial strain that fits best for the expression of each adhesion protein. In this project, the choice depends on the biology of each protein. CsgA will be expressed in an E. coli ΔcsgA knockout strain, so that the natural curli system of E. coli does not produce extra adhesion and confuse the results (Barnhart & Chapman, 2006). FapC and TasA will be expressed in standard E. coli strains such as BL21 or DH5α, because both proteins come from other organisms (Pseudomonas fluorescens and Bacillus subtilis), and E. coli does not produce them naturally. This way, the biofilm formed in the assay can be linked specifically to the recombinant adhesin produced by each cassette.
Bacterial Culturing. Bacterial culturing will be used to test the in silico predictions in living cells, through a biofilm-formation assay on plastic surfaces. Each E. coli strain will be grown in M9 minimal medium and added to microtiter plates of polyethylene (PE) and polystyrene (PS), and incubated for 24 to 48 hours at 28 °C without shaking, so that a biofilm can form. The amount of biofilm will be measured by crystal violet staining: the biofilm is washed, stained, and the dye is then dissolved in ethanol and read at OD590 (O’Toole, 2011). An empty-vector strain will be used as negative control. The same protocol will be repeated adding the lead anti-adhesion peptides designed in Aim 2, to check whether they reduce biofilm formation in a dose-dependent way. This is the main functional test of the project.
4.4. Identify any How To Grow (Almost) Anything Industry Council companies which are associated with your final project (optional)
The following companies and platforms are directly associated with this project:
Boltz.bio: Boltz is a modern protein structure prediction tool that can complement AlphaFold2 and ColabFold. It will be used to refine or cross-check the predicted structures of CsgA, FapC and TasA, especially in low-confidence regions. Twist Biosciences: Twist will be used to synthesize the FapC and TasA expression cassettes designed in this project. All DNA designs are validated against Twist synthesis rules (GC content, repeats and secondary structures) before submission. Addgene: Addgene is the main public repository of plasmids. It will be used to source the backbone vector for the expression cassettes and to find reference plasmids that already contain related adhesion proteins. SecureDNA: SecureDNA performs biosecurity screening of synthetic DNA orders. Since this project has a dual-use dimension (anti-adhesion design could in principle be misused to enhance microbial colonization), SecureDNA-type screening of synthesis orders is consistent with the ethical safeguards described in the previous section.
Section 5. RESULTS AND QUANTITATIVE EXPECTATIONS.
5.1. Validated aspect
The validation aspect chosen is the in silico characterization of how three bacterial amyloid adhesion proteins (CsgA, FapC and TasA) interact with two synthetic polymer surfaces (polyethylene and polystyrene), combined with the in silico design of three E. coli expression cassettes that will support future in vivo testing. This validation directly tests the central hypothesis of Aim 1, that bacterial amyloid adhesins recognize plastic surfaces through specific structural and physicochemical features, and provides both computational evidence (docking scores and contact residues) and a ready-to-synthesize experimental platform.
5.2. Detailed protocol
The validation followed the protocol described below.
Protein sequence retrieval. The protein sequences of CsgA (UniProt P28307, Escherichia coli K-12), FapC (UniProt P0DXF5, Pseudomonas fluorescens) and TasA (UniProt P54507, Bacillus subtilis 168) were downloaded from UniProt in FASTA format. The signal peptide and mature chain regions were noted for each protein, and only the mature chain was used for the docking analyses, since this is the functional form of the protein outside the cell.
Protein structure retrieval. The AlphaFold-predicted 3D structures of the three proteins were downloaded from the AlphaFold Protein Structure Database in .pdb format. The mean per-residue confidence (pLDDT) was recorded for each structure (CsgA 81.12, FapC 79.94, TasA 80.12), all classified as in the Confident range (pLDDT 70–90).
Surface analysis in PyMOL. Each .pdb file was opened in PyMOL. Per-atom solvent accessible surface area (SASA) was calculated with the command
get_area, load_b=1. Hydrophobic side-chain residues (Ala, Val, Leu, Ile, Met, Phe, Trp, Tyr) with β-carbon SASA above 5 Ų were selected, curated to the mature chain only, and reduced to 15 representative residues per protein, prioritising aromatic residues and the most exposed hydrophobic positions.Protein–polymer docking with AlphaFold 3. Each protein–polymer complex was predicted with the AlphaFold 3 web server (alphafoldserver.com). Polyethylene (PE) was modelled using palmitate (CCD code PLM, a 16-carbon hydrocarbon), since the public AlphaFold 3 server does not accept arbitrary SMILES strings. The top-ranked model (rank 0) was used for downstream analysis, and the ipTM and pTM scores were recorded.
Cross-validation with Boltz-1. As an independent validation, the same protein–polymer complexes were also predicted using Boltz-1, run through the ColabFold-style notebook (https://colab.research.google.com/github/sokrypton/ColabFold/blob/main/Boltz1.ipynb). Boltz-1 accepts custom SMILES as ligand input, allowing the use of the real polyethylene oligomer (
CCCCCCCCCCCCCCCC) and a polystyrene oligomer (CC(c1ccccc1)CC(c1ccccc1)CC(c1ccccc1)CC(c1ccccc1)C). The ipTM, pTM and confidence scores were extracted from the JSON output for each prediction. A custom Python cell was added at the end of the notebook to scan the entire output directory and print the contents of all generated JSON files (including the confidence scores per model), since the default notebook output does not display these scores directly in the cell output.Contact residue identification. For each top-ranked complex, the .cif file was opened in PyMOL and the protein residues within 5 Å of any ligand atom were identified using the script:
select ligand, not polymer/select contacts, byres (polymer within 5 of ligand)/iterate contacts and name CA, print(resi, resn). The resulting list was recorded for each protein–polymer combination.Visualization of complexes. A standardised visualization preset was applied to render each complex with consistent style: protein as slate-coloured cartoon, ligand as orange sticks (with O atoms in red), and contact residues as semi-transparent magenta spheres at the Cα position. Screenshots were captured for the figures.
DNA cassette design. The protein sequences of FapC and TasA were optimised for expression in E. coli K12 using the IDT Codon Optimization Tool. Each cassette was assembled in silico with the same architecture as the previously built CsgA cassette: BBa_J23106 promoter (35 bp), BBa_B0034 ribosome binding site (12 bp), ATG start codon, codon-optimised full coding sequence (signal peptide + mature chain), TAA stop codon, and BBa_B0015 double terminator (129 bp).
Cassette validation. The total length and GC content of each cassette were calculated with the Sequence Manipulation Suite (bioinformatics.org/sms2/dna_stats.html), and verified to fall within Twist Bioscience synthesis specifications (gBlock fragment range 125–3000 bp, GC content 40–60%). GC content is the percentage of guanine (G) and cytosine (C) nucleotides in a DNA sequence and is a key parameter for commercial synthesis: values outside the 40–60% range tend to produce secondary structures or unstable regions that compromise the synthesis quality, so all three cassettes were verified to fall within this safe interval.
5.3. Techniques Used
Several synthetic biology techniques covered in the HTGAA course were used in this validation. First, Databases (UniProt, NCBI and the AlphaFold Protein Structure Database) were used to retrieve the protein sequences, signal peptide annotations and predicted 3D structures of the three target adhesins. Second, Protein Design was applied through the AlphaFold 3 web server and PyMOL to predict protein–polymer complexes, identify solvent-exposed hydrophobic residues, and characterize the contact residues that mediate interaction with the polymer. Third, Use of Boltz-1 was applied as an independent cross-validation tool, since this model accepts custom SMILES strings and therefore allows direct docking against the real polyethylene and polystyrene oligomers. Fourth, Models and Notebooks were used through Google Colab notebooks (ColabFold and Boltz-1 notebooks) to run the predictions on free GPU instances. Finally, DNA Construct Design and Designing a Twist Order were applied to assemble the three E. coli expression cassettes with iGEM standard BioBrick parts (BBa_J23106, BBa_B0034 and BBa_B0015) and to validate them against Twist Bioscience synthesis specifications.
5.4. Project Data
Three complementary outputs were obtained:
- Three Twist-ready DNA expression cassettes (one per adhesin), validated against synthesis specifications.
- Surface analysis of the three target adhesins, identifying their candidate adhesion residues.
- Nine protein–polymer docking predictions (3 proteins × 3 prediction conditions: AlphaFold 3 with palmitate as PE proxy, Boltz-1 with real PE, and Boltz-1 with real PS).
5.4.1. Expression Cassettes
Three E. coli expression cassettes were designed in silico, one for each target adhesin (CsgA, FapC and TasA). The CsgA cassette was already assembled in a HTGAA homework and was used as the structural template for the design of the FapC and TasA cassettes. All three cassettes share the same modular BioBrick architecture, ensuring direct comparability between the three constructs. The architecture and final maps of the three cassettes are shown in Figure 3, and their compliance with Twist Bioscience synthesis specifications is summarized in Table 2.
Figure 4. Designed E. coli expression cassettes for the three target adhesins (CsgA, FapC and TasA), generated and visualized in Benchling. Top: linear maps showing the modular BioBrick architecture (BBa_J23106 promoter, BBa_B0034 ribosome binding site, ATG start codon, codon-optimized full coding sequence, TAA stop codon, BBa_B0015 double terminator). Bottom: plasmid maps showing the same cassettes inserted into a low-copy expression backbone, ready for transformation into E. coli during the future experimental phase (Sub-aim 1.7).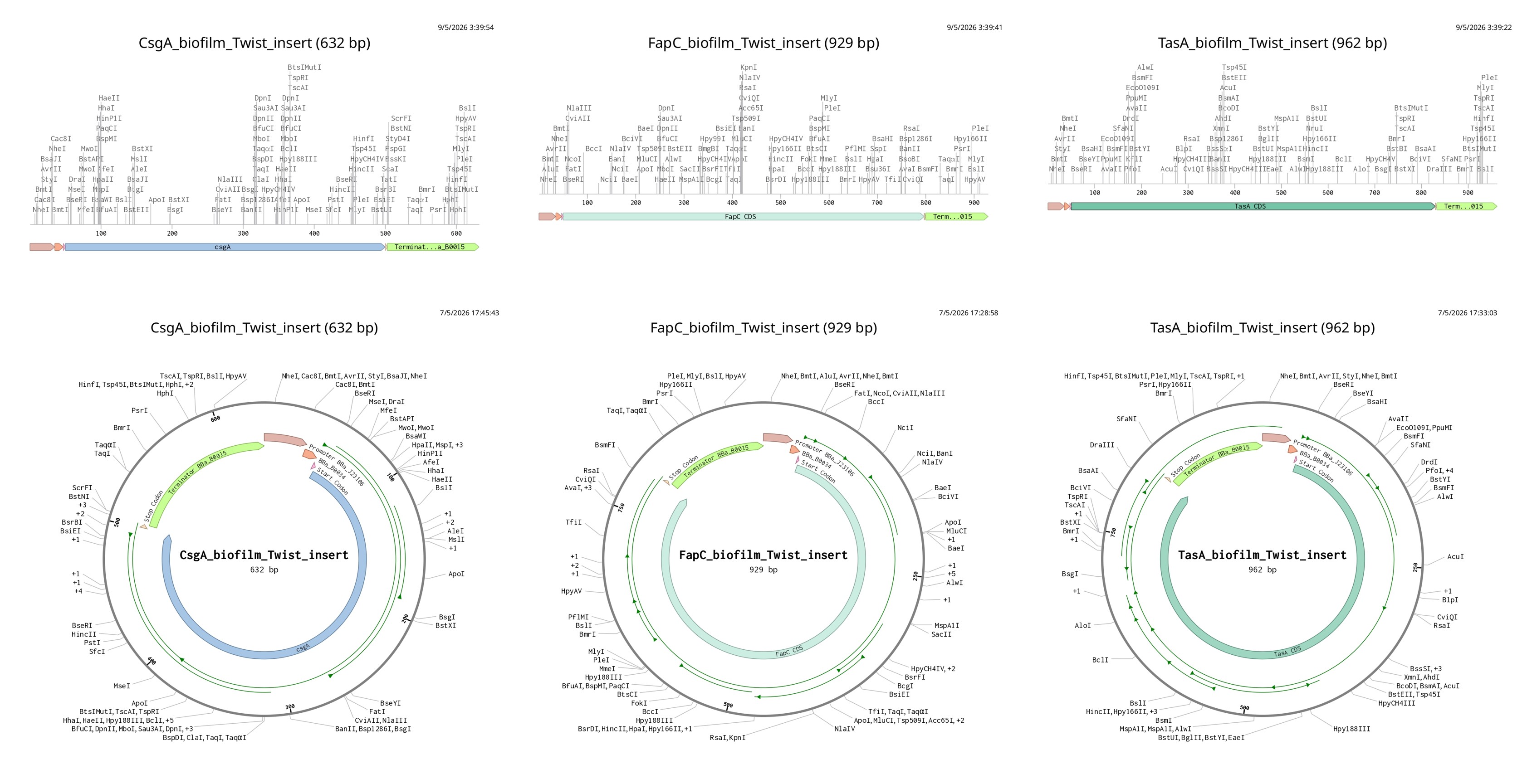
Validation of the three designed E. coli expression cassettes against Twist Bioscience synthesis specifications (Table 2). The total length and GC content of each cassette were calculated using the Sequence Manipulation Suite (bioinformatics.org/sms2/dna_stats.html). All three cassettes fall within the recommended GC content range (40–60%) and within the gBlock gene fragment length range (125–3000 bp), confirming that they are ready for commercial synthesis.
Table 2. Validation of the three E. coli expression cassettes against Twist Bioscience synthesis specification.
| Cassette | Length (bp) | GC content (%) | GC within 40–60% | Within gBlock 125–3000 bp |
|---|---|---|---|---|
| CsgA | 632 | 50.00 | ✓ | ✓ |
| FapC | 929 | 51.45 | ✓ | ✓ |
| TasA | 962 | 45.74 | ✓ | ✓ |
5.4.2. Hydrophobic Surface analysis
The three AlphaFold-predicted structures were analyzed in PyMOL to identify candidate adhesion residues. Per-atom solvent accessible surface area (SASA) was calculated and used to filter for solvent-exposed residues with two complementary properties: non-aromatic hydrophobic side chains (Ala, Val, Leu, Ile, Met), which mediate general hydrophobic interactions with both polyethylene and polystyrene and are therefore expected to be the main contributors to PE binding; and aromatic residues (Phe, Tyr, Trp), which combine hydrophobic character with π–π stacking interactions against the phenyl rings of polystyrene and are therefore expected to be especially relevant for PS recognition. The resulting list was curated to the mature chain only and reduced to 15 representative residues per protein. The selected residues are summarized in Table 3.
| Protein | Mature chain length (aa) | Selected hydrophobic surface residues (15 per protein) |
|---|---|---|
| CsgA | 131 | 26, 48, 50, 56, 58, 79, 97, 101, 106, 118, 124, 130, 142, 146, 151 |
| FapC | 226 | 30, 55, 64, 81, 82, 106, 112, 120, 129, 147, 183, 193, 197, 220, 250 |
| TasA | 234 | 29, 39, 70, 72, 89, 92, 109, 124, 139, 161, 191, 200, 217, 226, 233 |
The surface of each protein was rendered as a semi-transparent representation combined with the cartoon, coloured according to the chemical nature of each residue: slate for polar and uncharged residues, yellow for non-aromatic hydrophobic residues (Ala, Val, Leu, Ile, Met, Pro), orange for aromatic residues (Phe, Trp, Tyr), red for negatively charged residues (Asp, Glu), and marine blue for positively charged residues (Arg, Lys, His). The yellow and orange patches on the surface highlight the solvent-exposed hydrophobic regions identified by the SASA analysis, which are the most likely candidates to mediate adhesion to a hydrophobic plastic surface. The orange aromatic residues are particularly relevant for binding to polystyrene, since their planar rings can engage in π–π stacking with the phenyl groups of the polymer.
Figure 5. Surface analysis of the three target adhesion proteins (CsgA, FapC and TasA) rendered in PyMOL, showing solvent-exposed hydrophobic patches (yellow and orange) as candidate plastic-binding regions.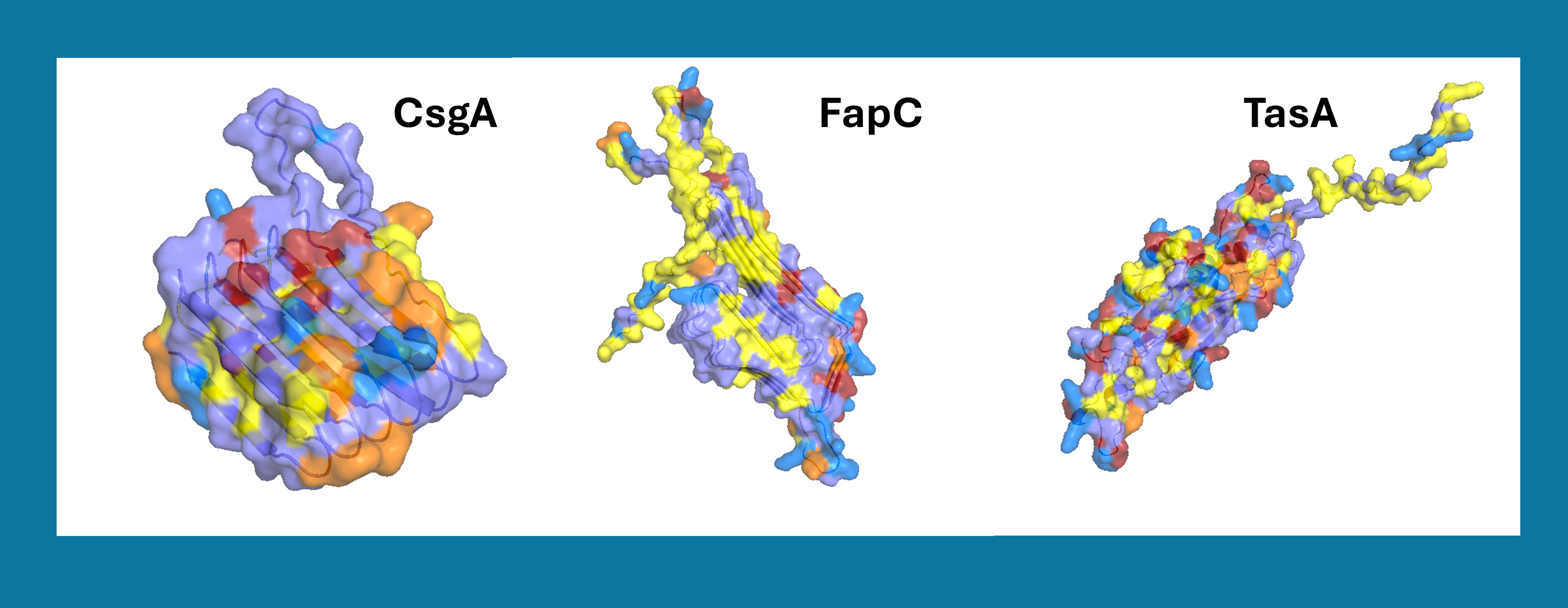
5.4.3. Protein–Polymer Docking Predictions
Once the candidate adhesion residues had been identified, each of the three target proteins was docked against polyethylene (PE) and polystyrene (PS) using two independent prediction tools: AlphaFold 3 with palmitate as PE proxy, and Boltz-1 with the real PE and PS oligomers as ligands. This combination yielded nine top-ranked complexes (3 proteins × 3 prediction conditions) that were evaluated through their confidence scores (ipTM and pTM) and through the identification of the protein residues in direct contact with the polymer. The confidence scores of the nine predictions are summarized in Table 4, and a representative view of each complex is shown in Figure 4.
The two polymers used in this study differ in their chemical structure and therefore in the type of interactions they can form with proteins. Polyethylene (PE) is a linear chain of repeating –CH₂–CH₂– units, and is purely hydrophobic with no functional groups capable of specific interactions beyond van der Waals forces. Polystyrene (PS) shares the same aliphatic backbone but each second carbon carries a phenyl ring (–C₆H₅), giving the polymer both hydrophobicity and the ability to form π–π stacking interactions with aromatic residues. Since the public AlphaFold 3 server does not accept arbitrary SMILES strings, polyethylene was modeled using palmitate (PLM, CCD code), a 16-carbon aliphatic chain (–CH₃(CH₂)₁₄COO⁻) included in the AlphaFold 3 ligand library. Palmitate reproduces the hydrocarbon backbone and the hydrophobic character of a short PE oligomer, although the carboxylate group at one end is absent in real PE; this small difference does not affect the bulk of the binding interaction.
Table 4. Confidence scores (ipTM / pTM) of the in silico protein–polymer docking predictions. Each cell shows the ipTM (interface predicted TM-score) and pTM (predicted TM-score) of the top-ranked model produced by each prediction tool.
| Protein | AF3 + PE (PLM proxy) | Boltz + PE | Boltz + PS |
|---|---|---|---|
| CsgA | 0.79 / 0.83 | 0.68 / 0.89 | 0.77 / 0.91 |
| FapC | 0.86 / 0.81 | 0.74 / 0.82 | 0.82 / 0.87 |
| TasA | 0.63 / 0.67 | 0.86 / 0.89 | 0.82 / 0.89 |
The ipTM (interface predicted TM-score) measures the model’s confidence in the relative position between the protein and the ligand, and is the most relevant metric for evaluating a docking prediction. The pTM (predicted TM-score) measures the model’s confidence in the overall fold of the complex, including both the protein structure and the ligand. Both scores range from 0 to 1, with values above 0.6 generally considered as confident, and values above 0.8 considered as high confidence.
The nine top-ranked complexes obtained from the two prediction tools are shown in Figure 4, arranged in a 3 × 3 grid (Figure 5). The upper row contains the three AlphaFold 3 predictions of each protein bound to palmitate (PE proxy). The middle row contains the three Boltz-1 predictions with the real polyethylene oligomer (CCCCCCCCCCCCCCCC), and the lower row contains the three Boltz-1 predictions with the polystyrene oligomer (CC(c1ccccc1)CC(c1ccccc1)CC(c1ccccc1)CC(c1ccccc1)C). All complexes are rendered with the same visualization preset (slate cartoon for the protein, orange sticks for the ligand, and semi-transparent magenta spheres at the Cα position of each contact residue) to allow direct visual comparison between proteins and prediction conditions.
Figure 6. Top-ranked predicted complexes between the three adhesion proteins (CsgA, FapC, TasA) and the two polymer ligands, organized in a 3 × 3 grid. Top row: AlphaFold 3 predictions with palmitate (PLM) as polyethylene proxy. Middle row: Boltz-1 predictions with the real polyethylene oligomer. Bottom row: Boltz-1 predictions with the polystyrene oligomer. Protein is shown as slate cartoon, ligand as orange sticks (oxygen atoms in red), and the protein residues within 5 Å of the ligand as semi-transparent magenta spheres at the Cα position.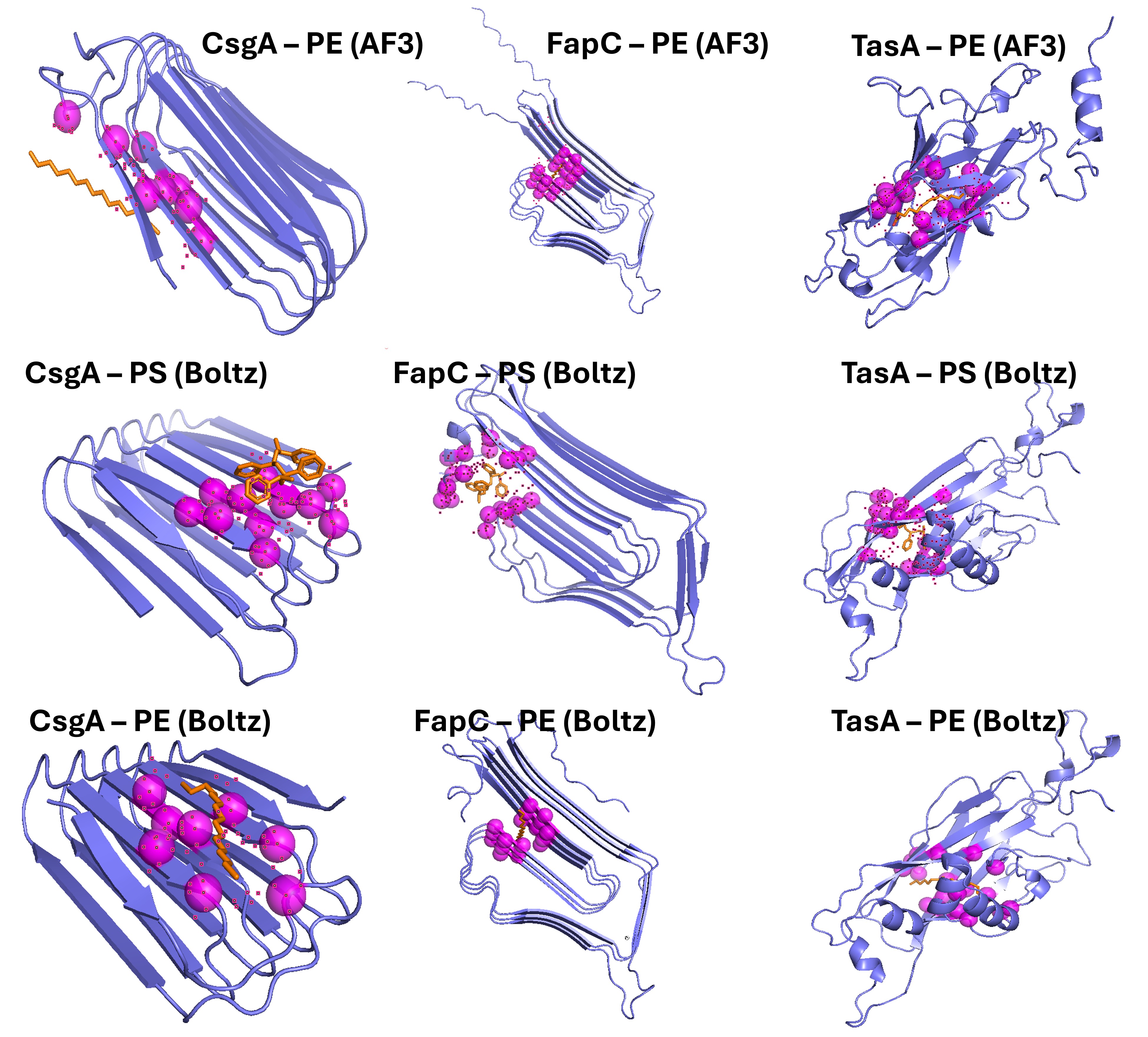
For each top-ranked complex, the protein residues within 5 Å of the ligand were identified in PyMOL and recorded as the predicted contact residues. The complete list, with aromatic residues highlighted as candidate π–π stacking partners with polystyrene, is summarized in Table 5.
| Complex | Number of contacts | Aromatic residues in contact |
|---|---|---|
| CsgA + PE (AF3) | 11 | F77, F122 |
| CsgA + PE (Boltz) | 8 | W86 |
| CsgA + PS (Boltz) | 11 | W86 |
| FapC + PE (AF3) | 17 | none |
| FapC + PE (Boltz) | 17 | none |
| FapC + PS (Boltz) | 21 | W6, F226 |
| TasA + PE (AF3) | 21 | F12, F43, F199 |
| TasA + PE (Boltz) | 19 | F12, F43, F199 |
| TasA + PS (Boltz) | 21 | F12, F43, F199, F201 |
Table 5. Contact residues identified in PyMOL within 5 Å of the ligand for each top-ranked docking complex, with aromatic residues highlighted as candidate π–π stacking partners with polystyrene.
Key observations from the docking data:
- All nine predictions show ipTM > 0.6, indicating that the three adhesins are predicted to interact with both PE and PS with confidence.
- FapC shows the highest scores overall, consistent with its repetitive amyloid architecture and abundance of solvent-exposed hydrophobic residues.
- For CsgA and FapC, ipTM is higher with PS than with PE in Boltz, which is consistent with sub-hypothesis H2 (stronger interaction with PS due to π–π stacking with aromatic residues).
- For TasA, ipTM is slightly higher with PE than with PS in Boltz, indicating that H2 is not universal: TasA seems to prefer PE, possibly because its globular β-sheet domain offers a different binding interface compared to the linear amyloid repeats of CsgA and FapC.
- Aromatic residues (Phe, Trp, Tyr) appear in the contact lists of PS dockings (W6, F226 in FapC; F12, F43, F199, F201 in TasA; W86 in CsgA), supporting the role of π–π stacking in PS recognition.
5.5. Troubleshooting
Several challenges and limitations were identified during the in silico validation of this project.
Challenges and limitations:
The AlphaFold 3 public server does not accept arbitrary SMILES strings, so palmitate (PLM) had to be used as a proxy for polyethylene. PLM is a 16-carbon hydrocarbon that reproduces the hydrophobic character of PE but adds a small carboxylate group at one end that does not exist in real PE. This may slightly bias the binding pose toward the carboxylate region.
The Boltz-1 Colab notebook had software incompatibilities between TensorFlow and Protobuf libraries that required manual reinstallation of specific package versions (protobuf 6.31.1) before the predictions could be executed. This is a known issue of community-maintained Colab notebooks and required several runtime restart cycles.
Both AlphaFold 3 and Boltz-1 use stochastic sampling, so different runs of the same input can produce slightly different binding poses, as observed in two independent CsgA + PE Boltz predictions. This variability needs to be considered when interpreting individual contact residues.
The polymer ligands used (16-carbon PE oligomer and 4-styrene PS oligomer) are short representations of the much larger real plastic surfaces. The predicted contacts approximate the initial binding event but cannot fully reproduce the behavior of bacterial adhesion to a full plastic slab in the environment.
Alternative strategies for future work:
Full-slab molecular dynamics simulations using GROMACS combined with CHARMM-GUI Polymer Builder would model the protein on top of a realistic polymer surface, providing a more accurate description of the adhesion process at the cost of significantly higher computational resources.
Restraint-based docking with HADDOCK 2.4, using custom polymer parameters, would allow the introduction of experimental knowledge about the binding interface (e.g., specific residues identified by mutagenesis).
Experimental wet-lab validation through the biofilm-formation assays described in Sub-aim 1.7, using the three E. coli strains transformed with the cassettes designed in this work, would provide a direct functional readout of the in silico predictions.
SECTION 6. AI Use Disclosure / Statement
This document was written with the assistance of a large language model (Claude, Anthropic). AI was used to support English drafting, to structure the text, and to format references. All scientific ideas, the project design, the choice of tools, and the interpretation of results are the author’s own, based on the work carried out during the HTGAA course.
SECTION 7. ADDITIONAL INFORMATION
7.1. References
Abramson, J., Adler, J., Dunger, J., et al. (2024). Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature, 630(8016), 493–500. https://doi.org/10.1038/s41586-024-07487-w
Austin, H. P., Allen, M. D., Donohoe, B. S., et al. (2018). Characterization and engineering of a plastic-degrading aromatic polyesterase. Proceedings of the National Academy of Sciences, 115(19), E4350–E4357. https://doi.org/10.1073/pnas.1718804115
Barnhart, M. M., & Chapman, M. R. (2006). Curli biogenesis and function. Annual Review of Microbiology, 60, 131–147. https://doi.org/10.1146/annurev.micro.60.080805.142106
Chen, L. T., Quinn, Z., Dumas, M., et al. (2025). Target sequence-conditioned design of peptide binders using masked language modeling. Nature Biotechnology. Advance online publication. https://doi.org/10.1038/s41587-025-02761-2
Di Pippo, F., Venezia, C., Sighicelli, M., et al. (2020). Microplastic-associated biofilms in lentic Italian ecosystems. Water Research, 187, 116429. https://doi.org/10.1016/j.watres.2020.116429
Ekanayaka, A. H., Tibpromma, S., Dai, D., et al. (2022). A review of the fungi that degrade plastic. Journal of Fungi, 8(8), 772. https://doi.org/10.3390/jof8080772
Jumper, J., Evans, R., Pritzel, A., et al. (2021). Highly accurate protein structure prediction with AlphaFold. Nature, 596(7873), 583–589. https://doi.org/10.1038/s41586-021-03819-3
Li, H.-Q., Wang, W.-L., Shen, Y.-J., & Su, J.-Q. (2025). Mangrove plastisphere as a hotspot for high-risk antibiotic resistance genes and pathogens. Environmental Research, 274, 121282. https://doi.org/10.1016/j.envres.2025.121282
O’Toole, G. A. (2011). Microtiter dish biofilm formation assay. Journal of Visualized Experiments, (47), 2437. https://doi.org/10.3791/2437
Sano, K., Kobayashi, H., Chuta, H., et al. (2023). CsgI (YccT) is a novel inhibitor of curli fimbriae formation in Escherichia coli preventing CsgA polymerization and curli gene expression. International Journal of Molecular Sciences, 24(5), 4357. https://doi.org/10.3390/ijms24054357
Zhai, X., Zhang, X.-H., & Yu, M. (2023). Microbial colonization and degradation of marine microplastics in the plastisphere: A review. Frontiers in Microbiology, 14, 1127308. https://doi.org/10.3389/fmicb.2023.1127308
7.2. Supplies and Budget
The following table summarizes the planned reagents, consumables and equipment for the future experimental phase (Sub-aim 1.7), including the Twist Bioscience DNA synthesis of the three cassettes designed in this work.
| Item | Description | Estimated cost (USD) | Supplier |
|---|---|---|---|
| Twist DNA synthesis | Three gene fragments (CsgA 632 bp, FapC 929 bp, TasA 962 bp) | ~250 | Twist Bioscience |
| Plasmid backbone | Low-copy expression vector (e.g., pSEVA221) | ~75 | Addgene |
| Gibson Assembly Master Mix | NEBuilder HiFi DNA Assembly, 50 reactions | ~290 | New England Biolabs |
| E. coli strains | DH5α (cloning), BL21 (expression), ΔcsgA knockout | ~180 | Addgene / CGSC |
| Plasmid Miniprep Kit | QIAprep Spin Miniprep Kit, 250 preps | ~330 | Thermo Fisher |
| LB broth and M9 minimal medium | Bacterial growth media for cloning and biofilm assay | ~80 | Millipore Sigma |
| 96-well microtiter plates | Polyethylene and polystyrene flat-bottom plates for biofilm assay | ~150 | Thermo Fisher |
| Crystal violet 0.1% | Biofilm staining reagent | ~40 | Millipore Sigma |
| Microplate reader (OD590) | Equipment usage for quantification of biofilm staining | institutional | e.g., Thermo Multiskan |
| Estimated total | Excluding equipment usage | ~1,400 | — |
Table 6. Estimated reagents, consumables and equipment for the future experimental phase of the project (Sub-aim 1.7), with approximate costs and trustworthy suppliers.