Week 3 HW: Lab automation
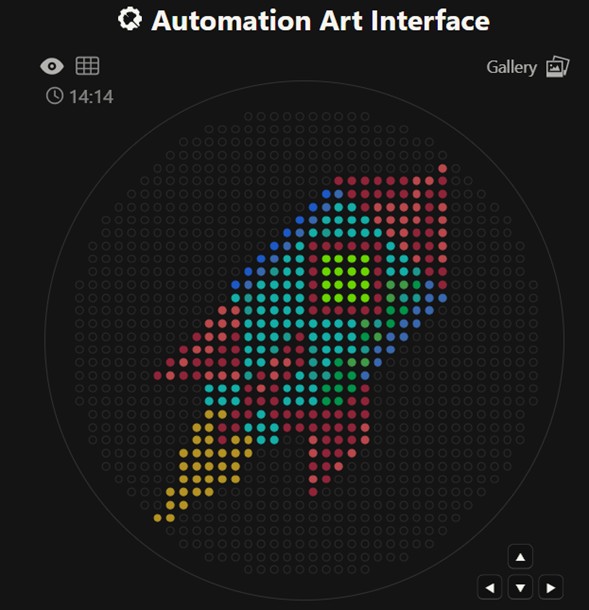
Generate an artistic design using the GUI at opentrons-art.rcdonovan.com.
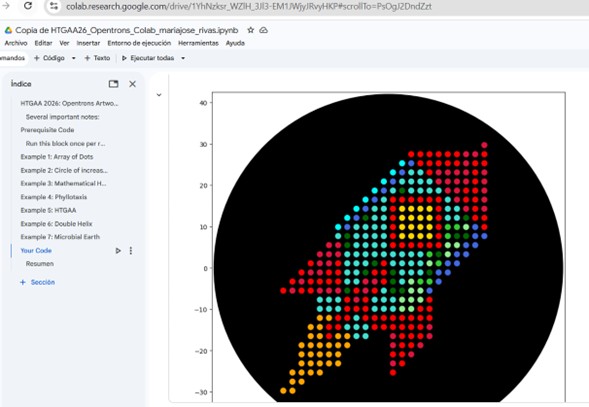
First, upload your drawing file. The platform allows you to edit the image with basic functions such as brightness, intensity, filters, etc. Next, modify the file by manually selecting points and colors to improve certain aspects. Generate the file and save it to the gallery with the name “rocket”. The link contains the coordinates, and you can also download the file with the .py extension. https://opentrons-art.rcdonovan.com/?id=w3dr10lr9t6cek6
Colab
Following the instructions in the week 3 assignment, I made a copy of the collab and the first thing was to understand the function of each component in the first block:
Robot Simulator/Mock
a. pipetteSim: Simulates the P20 pipette (max. 20 µL). Tracks aspirated/dispensed volumes, positions, colors, and generates a visualization using marplotlib. Validates errors such as: dispensing outside the dish, dispensing more than the aspirated volume, using the pipette without a tip, etc.
b. WellMock: Simulates a well in the labware.
c.LabwarMock: Simulates laboratory containers (racks, plates, etc.).
d. ModuleMock: Simulates modules such as the temperature sensor and thermocycler.
e. OpentronsMock: Simulates the robot’s main protocol, orchestrating the labware load, modules, and pipettes.
The visualization displays a black Petri dish (84 mm in diameter) with the dispensed points as a colored scatter plot.
Key Concepts for My Code:
a.Pick_up_tip(): Take a pipette tip (one per color to avoid cross-contamination).
b. Aspirate(vol, well): Draw liquid (colored bacteria) from a well. Maximum 20 µL per aspiration.
c. Dispense_and_detach(pipette, vol, location): Dispense a drop by moving up 5 mm, down, dispensing, and then back up (to avoid dragging/staining the drop).
d. Drop_tip: Release the tip after completing a color.
The coordinates are in mm relative to the center of the plate and must be within a radius of approximately 40 mm.
Settings: In the downloaded .py file from GUI at opentrons-art.rcdonovan, which contains the names of the proteins that express the color, add PROTEIN_COLOR_MAP to translate the protein names to matplotlib colors.
Also add set_offest() to labwareMock (no-op).
Each list contains (x, y) coordinates in millimeters relative to the center of the petri dish. Each coordinate is a point where the robot will deposit a drop of the corresponding color. These points were previously generated by page “x” and together form the figure. The list name corresponds to a fluorescent protein that produces a specific color when expressed:
| Protein | Color | Aspirate(uL) | Dispense (uL) | Points |
|---|---|---|---|---|
| mRFP1 | Red | 75.75 | 75.0 | 100 |
| mTurquoise2 | Turquoise | 58.8 | 57.75 | 77 |
| mScarlet_l | Carmesi | 35.25 | 34.5 | 46 |
| mKO2 | Orange | 24.75 | 24.0 | 32 |
| Azurite | Blue | 15.75 | 15.75 | 21 |
| Vanus | Gold | 12.0 | 12.0 | 16 |
| mJuniper | Dark green | 12.0 | 12.0 | 16 |
| mWasabi | Ligth green | 9.75 | 9.75 | 13 |
| Electra2 | Cyan | 6.0 | 6.0 | 8 |
| mClover3 | Lime | 5.25 | 5.25 | 7 |
Modifications to make the simulator work: The code is designed to run on the real robot, so some adjustments were necessary: 1.
Use AI assistance for this coding — Google Gemini or human assistance
I have limited experience programming in Python, so I relied on ChatGPT in developer mode, as well as Gemini integrated into Colab. I also asked for help from my colleague, MSc Rafael Pérez Aguirre, an expert in Artificial Intelligence, who helped me install the libraries correctly and properly apply the information that the AI had previously provided. I can’t say which tool was better, because both agents and my partner provided me with support.
I had problems with my code, so I asked a friend Rafael Peréz Aguirre, and he helped me. He also developed a library for the HTGAA community that, if you install it, lets you directly use the .py file that the operons-art website allows you to download. If you had problems with your code this week and haven’t finished your assignment yet, this will be very useful. I’m sharing the repository with you: https://github.com/Mozta/opentrons-bioart-sim
My Colab results: https://colab.research.google.com/drive/1YhNzksr_WZlH_3Jl3-EM1JWjyJRvyHKP#scrollTo=FF2b5c1P2_sH
Submit your Python file
Published Paper Utilizing Opentrons for Novel Biological Applications
One published paper that utilizes Opentrons automation for a novel biological application is: “An Open-Source Robotic Platform for Automated Biological Protocols” published in Micromachines (2024).
Overview of the Study
The paper describes the use of the Opentrons OT-2 liquid handling robot as a low-cost, open-source automation platform to perform complex biological protocols with high reproducibility. The researchers adapted the system to execute multistep workflows including liquid transfers, reagent mixing, incubation steps, and assay preparation. https://www.mdpi.com/2072-666X/15/6/708
Novel Biological Application
- Democratizing laboratory automation by using an affordable, open-source robotic system.
- Enabling reproducible and programmable biological workflows.
- Reducing human variability in sensitive biological assays.
Instead of relying on expensive industrial automation systems, the researchers demonstrate that Opentrons can be customized for:
- High-throughput screening
- Biological assay preparation
- Sample processing
- Reproducible experimental workflows
This represents a shift toward accessible automation in academic and small research laboratories.
Why This Is Significant? Traditional laboratory automation systems are often prohibitively expensive and closed-source. This study demonstrates that Opentrons can:
- Be customized through Python scripting
- Be modified with 3D-printed parts
- Enable scalable experimental designs
- Increase reproducibility in biological research
This makes it particularly relevant for emerging fields such as biofabrication, biosensor screening, and biomaterial experimentation, where iterative and programmable liquid handling is essential. a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
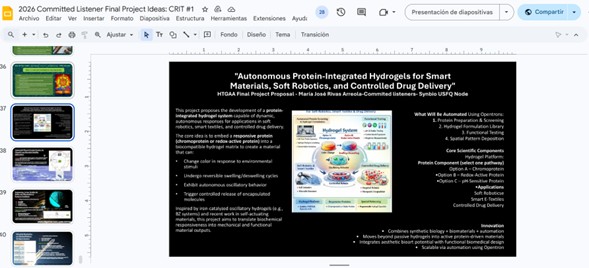
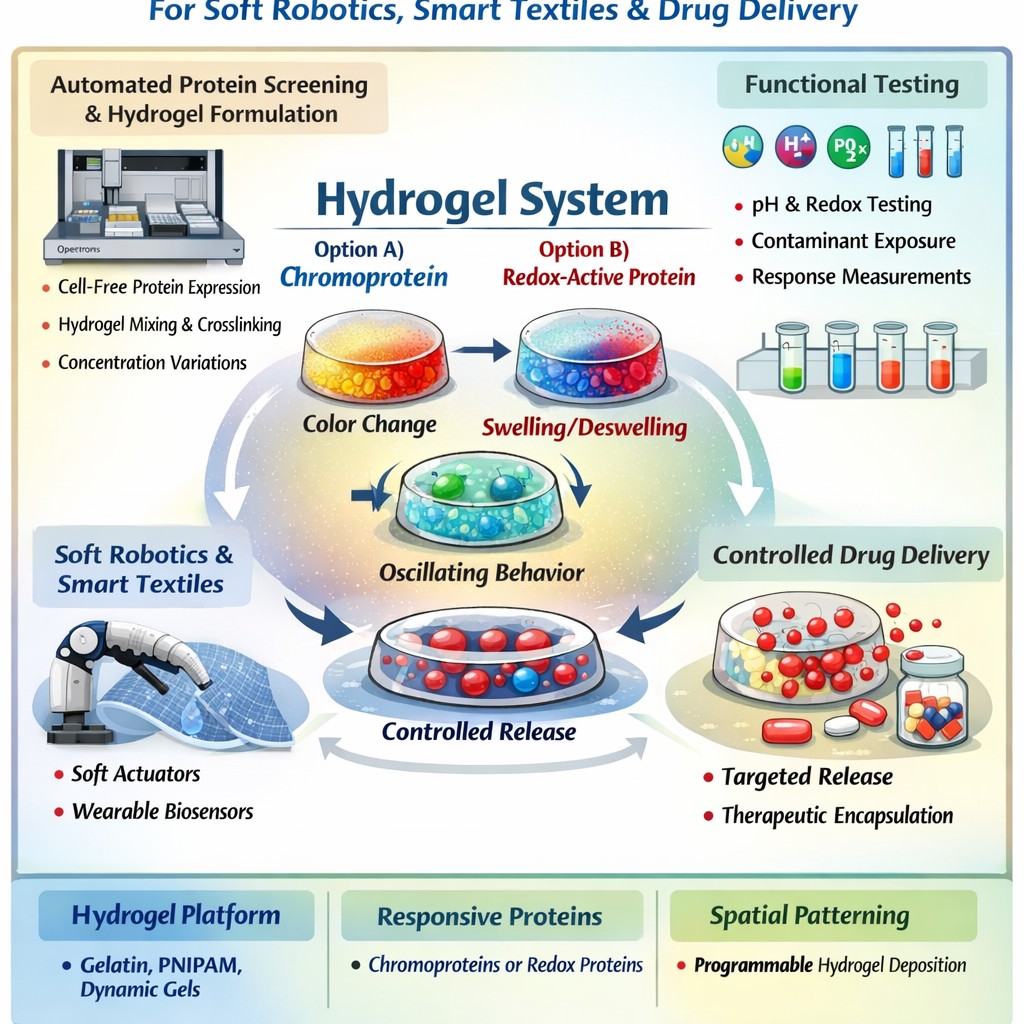
HTGAA Final Project Concept: Autonomous Protein-Integrated Hydrogels for Smart Materials, Soft Robotics, and Controlled Drug Delivery
Project Overview
This project proposes the development of a protein-integrated hydrogel system capable of dynamic, autonomous responses for applications in soft robotics, smart textiles, and controlled drug delivery.
The core idea is to embed a responsive protein (chromoprotein or redox-active protein) into a biocompatible hydrogel matrix to create a material that can:
- Change color in response to environmental stimuli
- Undergo reversible swelling/deswelling cycles
- Exhibit autonomous oscillatory behavior
- Trigger controlled release of encapsulated molecules
Inspired by iron-catalyzed oscillatory hydrogels (e.g., BZ systems) and recent work in self-actuating materials, this project aims to translate biochemical responsiveness into mechanical and functional material outputs.
What Will Be Automated Using Opentrons, the following processes will be automated:
- Protein Preparation & Screening
- Preparation of chromoprotein/redox-protein constructs
- Automated reaction setup for cell-free expression (if applicable)
- Screening of protein concentration gradients
- Hydrogel Formulation Library
- Automated mixing of hydrogel precursors (e.g., gelatin, GelMA, PNIPAM, alginate)
- Variation of crosslinking density
- Variation of protein incorporation ratios
- Functional Testing Automated preparation of:
- pH gradients
- Redox environments
- Contaminant exposure conditions
- High-throughput colorimetric or volumetric response screening
- Kinetic measurements of response time
- Spatial Pattern Deposition
- Programmed deposition of hydrogel-protein mixtures into defined geometries
- Pattern testing inspired by oscillatory gel geometry effects
Core Scientific Components
- Hydrogel Platform, Biocompatible, tunable matrix:
- Gelatin-based hydrogel
- PNIPAM-based thermo-responsive gel
- Dynamic covalent network (Schiff base / reversible crosslinking)
- Protein Component (select one pathway) Option A – Chromoprotein
- Visible color output
- Applications in bioart and smart textiles Option B – Redox-Active Protein
- Coupled redox changes drive volume change
- Applications in soft robotics
- Option C – pH-Sensitive Protein
- Detects infection-like acidic microenvironments
- Triggers drug release
Applications Soft Robotics Hydrogel undergoes periodic or stimulus-induced deformation a) Acts as a soft actuator b) Potential for autonomous motion or mechanical response Smart E-Textiles Protein-integrated fibers respond to: a)Sweat (pH) b)Environmental contamination c)Temperature changes Visible color shift or mechanical change enables wearable biosensing. a) Controlled Drug Delivery b) Hydrogel encapsulates therapeutic molecules. c) Protein-triggered structural change → controlled release under specific environmental conditions.
Innovation
- Combines synthetic biology + biomaterials + automation
- Moves beyond passive hydrogels into active protein-driven materials
- Integrates aesthetic bioart potential with functional biomedical design
- Scalable via automation using Opentron
Using the copilot tool to generate an image and loading information Hardware modifications: and Artistic dimension:, the result is: