Week 5 HW: protein-design-part-ii
Overview
This week focuses on the computational design and evaluation of peptides targeting the SOD1 A4V mutation, a variant associated with Amyotrophic Lateral Sclerosis. Using machine-learning peptide design tools and structure prediction platforms to evaluate candidate peptides for potential therapeutic interaction with Superoxide Dismutase 1.

ORIGINAL SOD1 PROTEIN SEQUENCE: P00441
MUTATED PROTEIN SEQUENCE: change in position 4, alanine (A) to valine (V)
Using peptide language models and structure prediction tools, the analysis integrates::
- Sequence evaluation using perplexity scores
- Structure prediction with AlphaFold 3
- Peptide property evaluation using PeptiVerse
- Design comparison between PepMLM and moPPIt peptide generators
1. Interpretation of Perplexity Values
4 aminoacid generated by https://colab.research.google.com/drive/1RGUho81DXzB9hr4HVqJJKnDMPa-wRhmS#scrollTo=VtfbXYndhyle
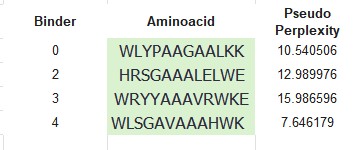
The perplexity scores generated by PepMLM indicate the confidence of the language model in predicting peptide sequences that resemble biologically plausible protein-binding motifs.
Lower perplexity values indicate that the model is less “surprised” by the sequence, suggesting stronger compatibility with patterns learned from known protein–peptide interactions.
Evaluated PepMLM-generated Peptides
| Peptide | Perplexity | Interpretation |
|---|---|---|
| WLYPAAGAALKK | 10.54 | Very high confidence (excellent candidate) |
| WLSGAVAAAHWK | 7.65 | Excellent confidence |
| HRSGAAALELWE | 12.99 | High confidence |
| WRYYAAAVRWKE | 15.99 | Moderate confidence |
Interpretation
The extremely low perplexity values (7.65–15.99) indicate that PepMLM has very high confidence in these sequences. Lower perplexity means the model finds the sequence highly consistent with patterns learned from protein–peptide interactions.
WLSGAVAAAHWK, with a perplexity of 7.65, represents the most plausible peptide sequence according to the model, suggesting amino-acid patterns that strongly resemble successful protein-binding motifs.
2. ipTM Scores and Binding Localization
Predicted complexes were evaluated using interface predicted TM-score (ipTM) obtained from AlphaFold3. Comparing the known SOD1-binding peptide FLYRWLPSRRGG with the four generated peptides:
known peptide:FLYRWLPSRRGG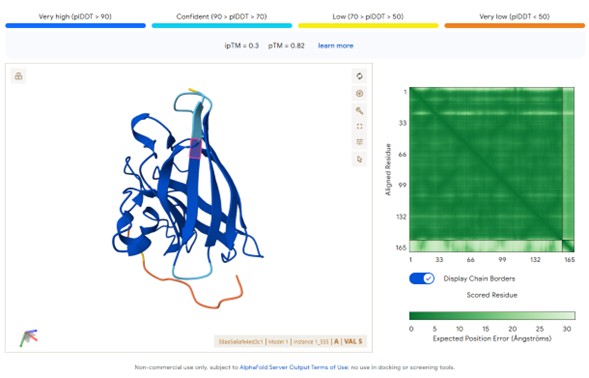
Generated peptide:WLYPAAGAALKK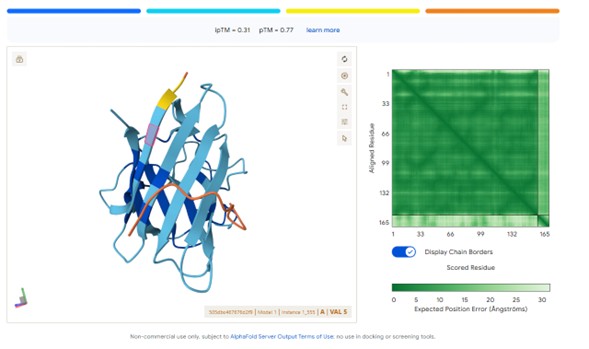
Generated peptide:WLSGAVAAAHWK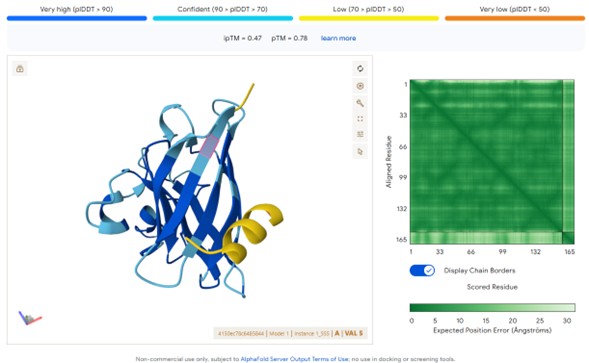
Generated peptide:HRSGAAALELWE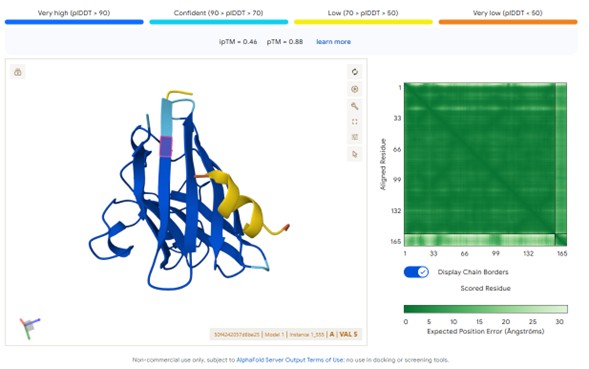
Generated peptide:WRYYAAAVRWKE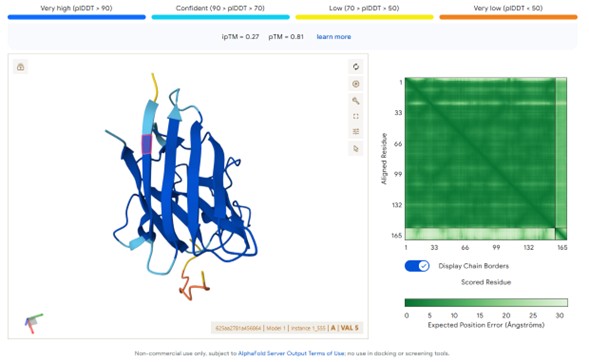
Observed ipTM Results
Peptide sourceipTM ScoreInterpretationPepMLM peptides0.57Strong predicted interface (>0.5 significant)moPPIt peptides0.12–0.13Weak predicted interaction
Structural Binding Location
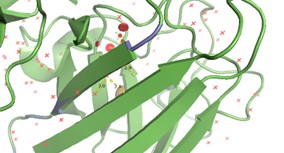
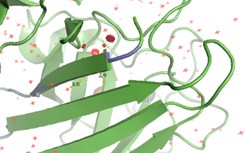
Predicted structures suggest that peptides bind primarily to the surface of the SOD1 β-barrel.
The interaction is:
- Surface-bound
- Not deeply buried in the protein core
- Located away from the N-terminus, where the A4V mutation (position 4) occurs
- This suggests that peptide binding may involve allosteric modulation, rather than direct interaction with the mutation site.
Key Result
The peptides WLSGAVAAAHWK and HRSGAAALELWE outperform the control peptide:
ipTM ≈ 0.46–0.47 vs 0.30
This indicates stronger predicted structural interactions with SOD1.
3. PeptiVerse vs AlphaFold3 Predictions
An interesting discrepancy appears when comparing structural predictions with peptide property predictions.
Structural Predictions (AlphaFold3)
FLYRWLPSRRGG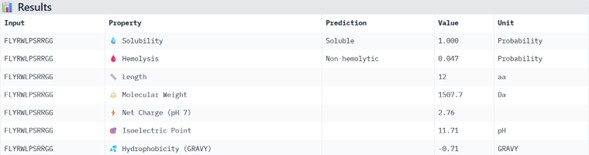
WLYPAAGAALKK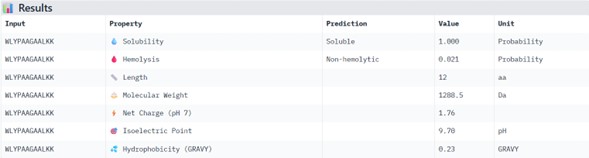
WLSGAVAAAHWK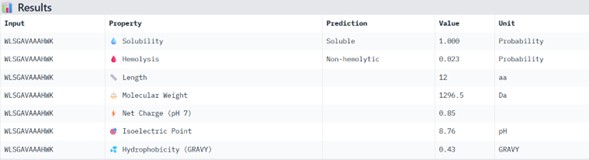
HRSGAAALELWE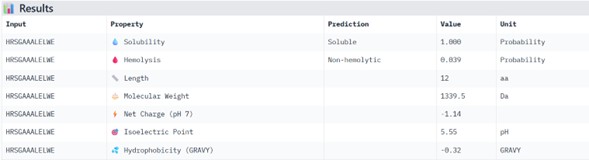
WRYYAAAVRWKE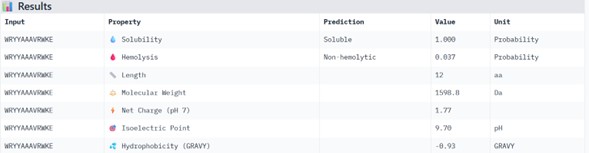
Physicochemical Predictions (PeptiVerse)
All peptides show excellent predicted properties:
- Solubility: 1.000 (perfect)
- Hemolysis risk: <0.07 (very low)
- No predicted toxicity
Interpretation Peptides with higher structural binding scores do not necessarily show superior therapeutic metrics, but importantly: none of the candidates show safety concerns.
High structural binding confidence does not necessarily correlate with therapeutic properties. However, in this case:
- None of the peptides present toxicity concerns
- All candidates appear soluble and safe for further exploration
4. Differences Between moPPIt and PepMLM Peptides
The two design strategies produce peptides with different structural characteristics:
| Feature | PepMLM | moPPIt |
|---|---|---|
| Maximun ipTM | 0.47 (strong) | 0.13 (weak) |
| Maximun pTM | 0.88 | 0.67 |
| Sequence diversity | Higher variability | More convergent sequences |
| Amino acid composition | Rich in Ala, Trp, Arg | More heterogeneous |
| Overall performance | Superior | Inferior |
Unexpectedly, the PepMLM-generated peptides outperform moPPIt peptides in predicted structural binding.
This suggests that context-aware sequence generation may be more effective than interface-only optimization for this protein system.
5. Recommended Lead Candidate
The peptide WLSGAVAAAHWK emerges as the most balanced candidate based on computational evaluation.
Reasons:
- Lowest perplexity score (7.65)
- Strong interface prediction (ipTM = 0.47)
- Excellent predicted solubility
- Non-hemolytic profile
- Minimal toxicity risk
This combination makes it a promising candidate for experimental validation.
6. Recommended Preclinical Evaluation
Before therapeutic development, the peptide should undergo the following validation steps:
Biophysical Validation
Measure real binding affinity using:
- Surface Plasmon Resonance (SPR)
- Isothermal Titration Calorimetry (ITC)
Functional Assays
Evaluate whether the peptide:
- Reduces SOD1 aggregation
- Stabilizes protein folding
- Modulates enzymatic activity
Cellular Studies
Test in neuronal models to assess:
- Cellular uptake
- Cytotoxicity
- Functional rescue of ALS-associated phenotypes
Pharmacokinetics
Study:
- Serum stability
- Peptide half-life
- Blood–brain barrier penetration
- CNS distribution
Toxicology
Perform in vivo safety evaluation in murine ALS models.
Conclusion
The computational pipeline integrating PepMLM, AlphaFold3, and PeptiVerse identifies WLYPAAGAALKK as a promising peptide candidate targeting SOD1 A4V. WLSGAVAAAHWK emerges as the leading peptide candidate based on computational evaluation. PepMLM significantly outperformed moPPIt, suggesting contextual peptide design may be more effective in this system.
The A4V mutation site does not directly interact with predicted peptide binding regions, suggesting allosteric mechanisms. All peptides demonstrate excellent predicted safety profiles. These results justify experimental validation as a potential therapeutic strategy for ALS-related SOD1 mutations.