Week 6 HW: genetic circuits part i
1. What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
The Phusion High-Fidelity PCR Master Mix contains several key components required for PCR:
- Phusion DNA polymerase – the enzyme that synthesizes new DNA strands by adding nucleotides complementary to the template DNA.
- dNTPs (deoxynucleotide triphosphates) – the building blocks used by the polymerase to create the new DNA strand.
- Reaction buffer – maintains the optimal pH and salt conditions needed for the enzyme to function properly.
- MgCl₂ (magnesium ions) – acts as a cofactor required for DNA polymerase activity and helps stabilize primer-template interactions.
- Some mixes may also include additives (e.g., GC buffer or DMSO) that help amplify difficult templates such as GC-rich DNA.
2. What are some factors that determine primer annealing temperature during PCR?
Several factors influence the primer annealing temperature:
- Primer melting temperature (Tm) – annealing temperature is usually set a few degrees below the Tm.
- Primer length – longer primers generally have higher melting temperatures.
- GC content – primers with higher GC content bind more strongly and therefore have higher Tm values
- Salt and buffer conditions – ion concentration (such as Mg²⁺) can affect primer binding stability.
- Primer complementarity or secondary structures – hairpins or dimers can affect efficient annealing.
3. Compare PCR and restriction enzyme digests for creating linear DNA fragments.
PCR Protocol:
- Uses primers, DNA polymerase, dNTPs, and thermal cycling.
- Amplifies a specific DNA region defined by the primers.
Advantages:
- Can amplify specific DNA fragments from very small amounts of template.
- Allows modification of DNA sequences (e.g., adding overlaps for Gibson assembly).
- Does not require restriction sites in the sequence.
Disadvantages:
- Can introduce mutations if low-fidelity polymerases are used.
- Requires primer design.
Restriction enzyme digest Protocol:
- Uses restriction enzymes that recognize specific DNA sequences and cut at those sites.
- Performed by incubating DNA with the enzyme and buffer at a specific temperature.
Advantages:
- Produces precise cuts at known recognition sites.
- Does not rely on amplification.
Disadvantages:
- Only works if the required restriction sites exist in the DNA.
- Less flexible for modifying sequences.
When to use each:
- PCR is preferable when you need to amplify a fragment, modify sequences, or add overlaps for cloning.
- Restriction digests are preferable when the correct restriction sites already exist and you want a simple, precise cut without amplification.
4. How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning? To ensure compatibility with Gibson cloning:
- Design PCR primers that add overlapping sequences (usually 20–40 bp) between the fragments and the vector.
- Make sure the overlap sequences are identical between adjacent fragments.
- Verify that the reading frame and orientation of the insert are correct.
- Confirm fragment sequences using DNA sequencing or in-silico design tools before assembly.
These overlaps allow the Gibson Assembly enzymes to join the fragments seamlessly.
5. How does the plasmid DNA enter the E. coli cells during transformation? During transformation, plasmid DNA enters competent E. coli cells that have been chemically treated or electrocompetent.
In chemical transformation (most common in teaching labs):
- Cells are treated with calcium chloride to make the membrane permeable.
- The plasmid DNA binds to the cell surface.
- A heat shock step briefly increases membrane permeability.
- DNA passes through the membrane into the cell.
- Cells recover and replicate the plasmid.
In electroporation, an electric pulse creates temporary pores in the membrane that allow DNA to enter.
6. Describe another assembly method: Golden Gate Assembly
a) Golden Gate Assembly
Is a molecular cloning method that allows for the simultaneous, seamless, and ordered assembly of multiple DNA fragments into a destination vector in a single “one-pot” reaction. It uses Type IIS restriction enzymes (e.g., BsaI) and T4 DNA ligase to cut and paste DNA, leaving no scar sequences at the junctions.
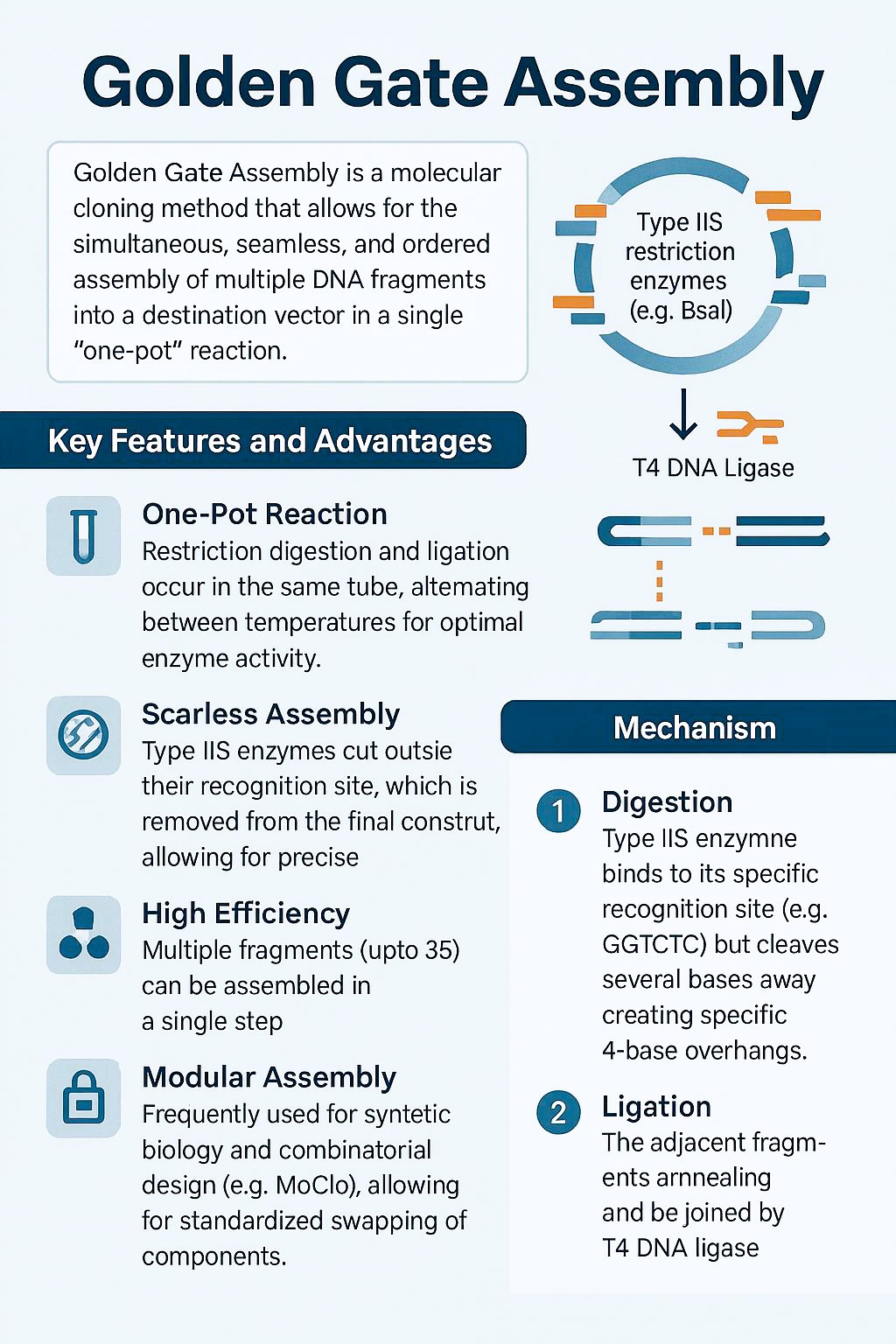 Image designed in copilot
Image designed in copilot
b) Golden Gate
Open Benchling account
Import Backbone sequence: M77789.2, Cloning vector pUC19, complete sequence:
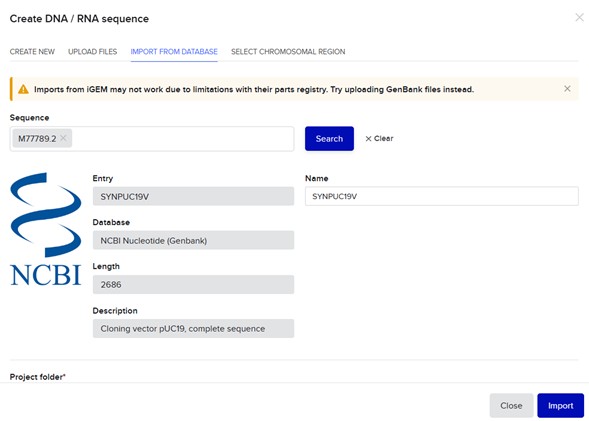
Import insert sequence: AH002844.2, Homo sapiens insulin (INS) gene, complete cds:
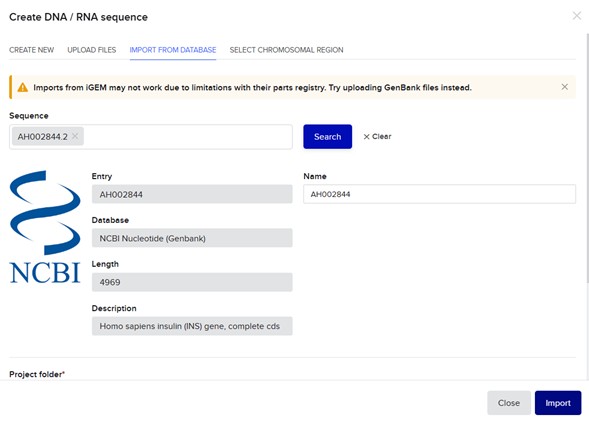
Select backbone + select insert + Pick assembly:
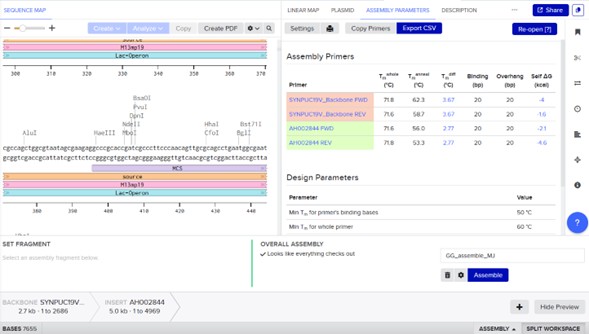
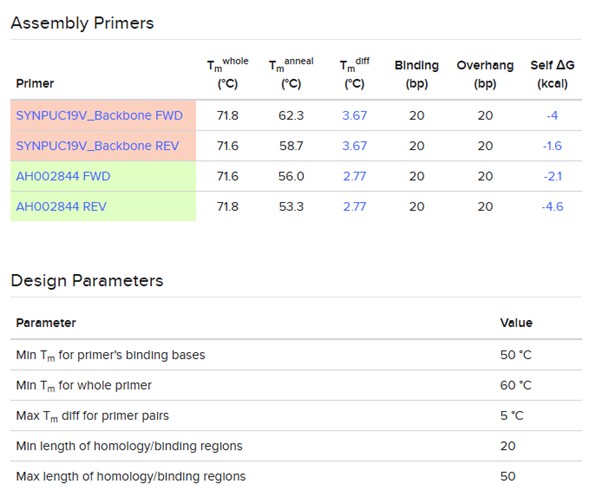
- Assembly primers:
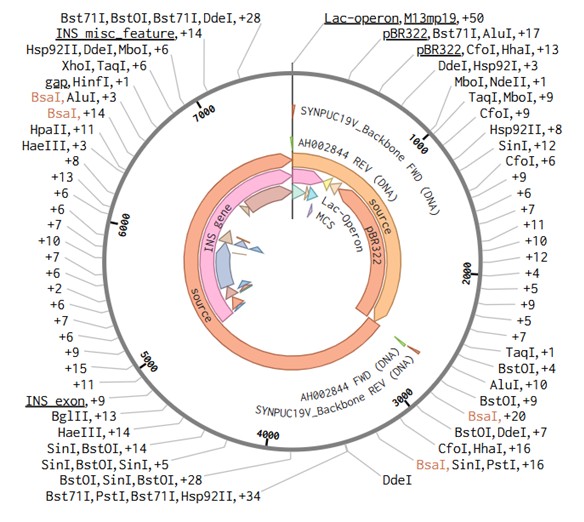
-Plasmid:
- Primers:
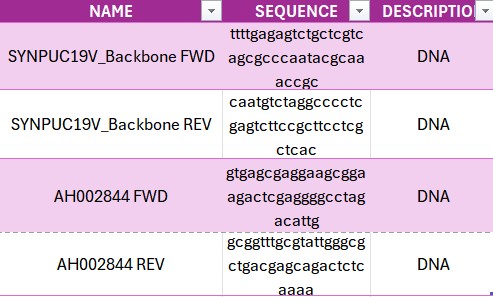
Interpretation/conclusion: The Golden Gate Assembly successfully enabled the construction of a recombinant plasmid using the pUC19 vector as the backbone and the human insulin cDNA as the insert. By incorporating type IIS restriction enzyme recognition sites (such as BsaI) flanking both the vector and insert, compatible 4-base pair overhangs were generated, allowing precise and directional assembly. This method ensured seamless ligation without leaving extra sequences, resulting in an efficient and accurate cloning strategy. Overall, Golden Gate Assembly proved to be a powerful and flexible technique for constructing recombinant DNA, particularly for applications requiring precise control over fragment orientation and sequence design.