Week 3 HW: Lab automation
- Python Script for Opentrons Artwork — DUE BY YOUR LAB TIME
Portrait
opentrons-art.rcdonovan.com/?id=s49o23t6m4pr6v4

Option 2. Flower opentrons-art.rcdonovan.com/?id=g5u4j61fb0v70e7
from opentrons import types
metadata = { ‘protocolName’: ‘HTGAA Opentrons Lab’, ‘author’: ‘MARIAORTIZ’, ‘source’: ‘HTGAA 2022’, ‘apiLevel’: ‘2.20’ }
##############################################################################
Robot deck setup constants - don’t change these
##############################################################################
TIP_RACK_DECK_SLOT = 9 COLORS_DECK_SLOT = 6 AGAR_DECK_SLOT = 5 PIPETTE_STARTING_TIP_WELL = ‘A1’
well_colors = { ‘A1’ : ‘Red’, ‘B1’ : ‘Yellow’, ‘C1’ : ‘Green’, ‘D1’ : ‘Cyan’, ‘E1’ : ‘Blue’ # if in a 24-well plate, this needs to be moved to e.g. D2 }
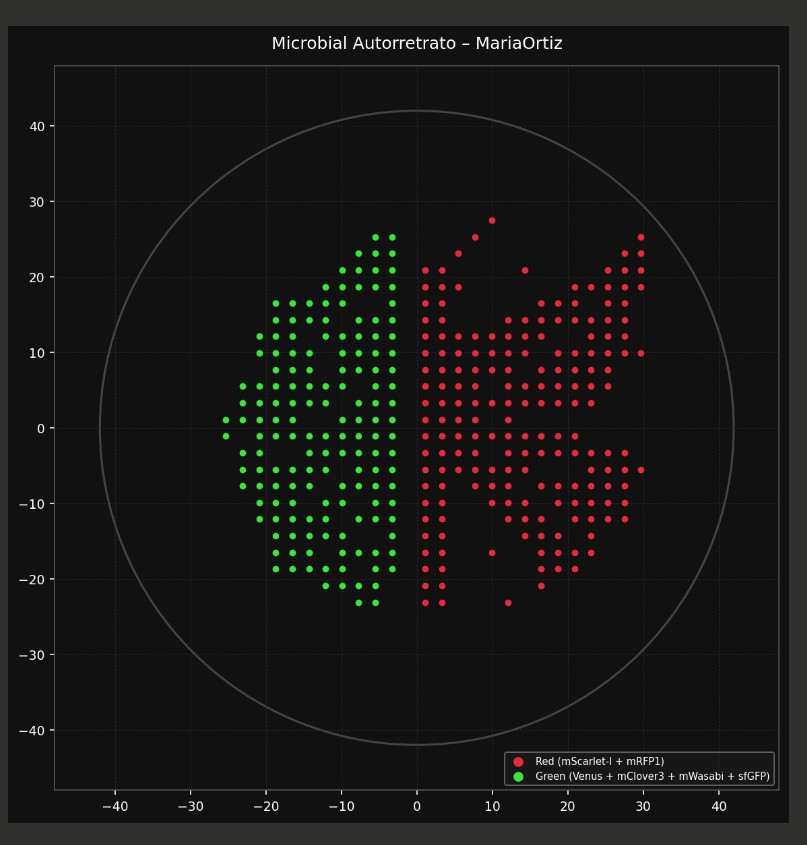
red_points = [(-5, 25), (0, 25), (5, 25), (-15, 20), (-10, 20), (-5, 20), (10, 20), (15, 20), (-10, 15), (-5, 15), (10, 15), (15, 15), (-15, 10), (-10, 10), (-5, 10), (0, 10), (5, 10), (10, 10), (15, 10), (-20, 5), (-15, 5), (15, 5), (-20, 0), (20, 0)]
green_points = [(-30, -15), (-25, -15), (-20, -15), (-15, -15), (15, -15), (20, -15), (25, -15), (30, -15), (-30, -20), (-25, -20), (-20, -20), (-15, -20), (-10, -20), (-5, -20), (0, -20), (5, -20), (10, -20), (15, -20), (20, -20), (25, -20), (30, -20), (-20, -25), (-15, -25), (-10, -25), (-5, -25), (0, -25), (5, -25), (10, -25), (15, -25), (20, -25)]
red1_points = [(-5, 30), (0, 30), (5, 30), (-15, 25), (-10, 25), (10, 25), (15, 25), (-20, 20), (20, 20), (-20, 15), (20, 15), (-25, 10), (25, 10), (-25, 5), (-10, 5), (-5, 5), (0, 5), (5, 5), (10, 5), (25, 5), (-25, 0), (-15, 0), (-5, 0), (5, 0), (15, 0), (25, 0), (-20, -5), (-15, -5), (-5, -5), (5, -5), (15, -5), (20, -5), (-15, -10), (15, -10), (-10, -15), (-5, -15), (0, -15), (5, -15), (10, -15)]
def run(protocol): ##############################################################################
Load labware, modules and pipettes
##############################################################################
##############################################################################
Patterning
##############################################################################
Helper functions for this lab
pass this e.g. ‘Red’ and get back a Location which can be passed to aspirate()
def location_of_color(color_string): for well,color in well_colors.items(): if color.lower() == color_string.lower(): return color_plate[well] raise ValueError(f"No well found with color {color_string}")
For this lab, instead of calling pipette.dispense(1, loc) use this: dispense_and_jog(pipette, 1, loc)
def dispense_and_jog(pipette, volume, location): """ Dispense and then move up 5mm and back down to shake all dispensed fluid off the tip; this also ensures it’s not moving laterally before the dispense is done. """ assert(isinstance(volume, (int, float))) pipette.dispense(volume, location) currLoc = pipette._get_last_location_by_api_version() pipette.move_to(currLoc.move(types.Point(z=5))) pipette.move_to(currLoc)
YOUR CODE HERE to create your design
pipette_20ul.pick_up_tip() red_points = [(29.7,25.3),(29.7,23.1),(14.3,20.9),(25.3,20.9),(27.5,20.9),(5.5,18.7),(20.9,18.7),(23.1,18.7),(25.3,18.7),(27.5,18.7),(16.5,16.5),(18.7,16.5),(20.9,16.5),(25.3,16.5),(12.1,14.3),(14.3,14.3),(16.5,14.3),(18.7,14.3),(20.9,14.3),(23.1,14.3),(25.3,14.3),(1.1,12.1),(7.7,12.1),(12.1,12.1),(14.3,12.1),(16.5,12.1),(23.1,12.1),(25.3,12.1),(9.9,9.9),(12.1,9.9),(14.3,9.9),(18.7,9.9),(20.9,9.9),(23.1,9.9),(27.5,9.9),(29.7,9.9),(5.5,7.7),(9.9,7.7),(12.1,7.7),(16.5,7.7),(18.7,7.7),(20.9,7.7),(23.1,7.7),(7.7,5.5),(12.1,5.5),(14.3,5.5),(16.5,5.5),(18.7,5.5),(20.9,5.5),(23.1,5.5),(14.3,3.3),(16.5,3.3),(18.7,3.3),(20.9,3.3),(12.1,1.1),(5.5,-1.1),(7.7,-1.1),(12.1,-1.1),(14.3,-1.1),(16.5,-1.1),(18.7,-1.1),(20.9,-1.1),(7.7,-3.3),(12.1,-3.3),(14.3,-3.3),(16.5,-3.3),(18.7,-3.3),(27.5,-3.3),(5.5,-5.5),(12.1,-5.5),(14.3,-5.5),(29.7,-5.5),(7.7,-7.7),(9.9,-7.7),(12.1,-7.7),(16.5,-7.7),(18.7,-7.7),(20.9,-7.7),(1.1,-9.9),(9.9,-9.9),(12.1,-9.9),(14.3,-9.9),(12.1,-12.1),(14.3,-12.1),(16.5,-12.1),(25.3,-12.1),(14.3,-14.3),(9.9,-16.5),(23.1,-16.5),(1.1,-18.7),(3.3,-18.7),(16.5,-18.7),(18.7,-18.7),(16.5,-20.9),(12.1,-23.1),(9.9,27.5),(7.7,25.3),(5.5,23.1),(27.5,23.1),(1.1,20.9),(3.3,20.9),(29.7,20.9),(1.1,18.7),(3.3,18.7),(29.7,18.7),(1.1,16.5),(3.3,16.5),(27.5,16.5),(1.1,14.3),(3.3,14.3),(27.5,14.3),(3.3,12.1),(5.5,12.1),(9.9,12.1),(27.5,12.1),(1.1,9.9),(3.3,9.9),(5.5,9.9),(7.7,9.9),(25.3,9.9),(1.1,7.7),(3.3,7.7),(7.7,7.7),(25.3,7.7),(1.1,5.5),(3.3,5.5),(5.5,5.5),(25.3,5.5),(1.1,3.3),(3.3,3.3),(5.5,3.3),(7.7,3.3),(9.9,3.3),(12.1,3.3),(23.1,3.3),(1.1,1.1),(3.3,1.1),(5.5,1.1),(7.7,1.1),(1.1,-1.1),(3.3,-1.1),(9.9,-1.1),(1.1,-3.3),(3.3,-3.3),(5.5,-3.3),(20.9,-3.3),(23.1,-3.3),(25.3,-3.3),(1.1,-5.5),(3.3,-5.5),(7.7,-5.5),(9.9,-5.5),(23.1,-5.5),(25.3,-5.5),(27.5,-5.5),(1.1,-7.7),(3.3,-7.7),(23.1,-7.7),(25.3,-7.7),(27.5,-7.7),(3.3,-9.9),(18.7,-9.9),(20.9,-9.9),(23.1,-9.9),(25.3,-9.9),(27.5,-9.9),(1.1,-12.1),(3.3,-12.1),(20.9,-12.1),(23.1,-12.1),(27.5,-12.1),(1.1,-14.3),(3.3,-14.3),(16.5,-14.3),(18.7,-14.3),(23.1,-14.3),(1.1,-16.5),(3.3,-16.5),(16.5,-16.5),(18.7,-16.5),(20.9,-16.5),(20.9,-18.7),(1.1,-20.9),(3.3,-20.9),(1.1,-23.1),(3.3,-23.1)] for x, y in red_points: adjusted_location = center_location.move(types.Point(x=x, y=y)) if pipette_20ul.current_volume == 0: pipette_20ul.aspirate(1, location_of_color(‘Red’)) dispense_and_jog(pipette_20ul, 1, adjusted_location) pipette_20ul.drop_tip()
pipette_20ul.pick_up_tip() green_points = [(-7.7,20.9),(-5.5,20.9),(-12.1,18.7),(-9.9,18.7),(-5.5,18.7),(-3.3,18.7),(-14.3,16.5),(-12.1,16.5),(-3.3,16.5),(-16.5,14.3),(-12.1,14.3),(-5.5,14.3),(-3.3,14.3),(-16.5,12.1),(-12.1,12.1),(-9.9,12.1),(-7.7,12.1),(-5.5,12.1),(-3.3,12.1),(-18.7,9.9),(-14.3,9.9),(-9.9,9.9),(-7.7,9.9),(-5.5,9.9),(-18.7,7.7),(-16.5,7.7),(-9.9,7.7),(-7.7,7.7),(-5.5,7.7),(-3.3,7.7),(-20.9,5.5),(-16.5,5.5),(-14.3,5.5),(-12.1,5.5),(-9.9,5.5),(-3.3,5.5),(-16.5,3.3),(-14.3,3.3),(-12.1,3.3),(-7.7,3.3),(-5.5,3.3),(-3.3,3.3),(-16.5,1.1),(-9.9,1.1),(-5.5,1.1),(-12.1,-1.1),(-9.9,-1.1),(-7.7,-1.1),(-5.5,-1.1),(-3.3,-1.1),(-9.9,-3.3),(-7.7,-3.3),(-5.5,-3.3),(-5.5,-5.5),(-3.3,-5.5),(-5.5,-7.7),(-3.3,20.9),(-7.7,18.7),(-14.3,14.3),(-18.7,5.5),(-20.9,3.3),(-18.7,3.3),(-20.9,1.1),(-3.3,1.1),(-20.9,-1.1),(-16.5,-1.1),(-14.3,-3.3),(-12.1,-3.3),(-20.9,-5.5),(-18.7,-5.5),(-16.5,-5.5),(-14.3,-5.5),(-12.1,-5.5),(-16.5,-7.7),(-14.3,-7.7),(-9.9,-7.7),(-7.7,-7.7),(-12.1,-9.9),(-5.5,-9.9),(-3.3,-9.9),(-16.5,-12.1),(-12.1,-12.1),(-7.7,-12.1),(-3.3,-12.1),(-14.3,-14.3),(-9.9,-14.3),(-3.3,-14.3),(-9.9,-16.5),(-7.7,-16.5),(-5.5,-16.5),(-3.3,-16.5),(-9.9,-18.7),(-5.5,-20.9),(-9.9,16.5),(-7.7,14.3),(-16.5,9.9),(-3.3,9.9),(-14.3,7.7),(-5.5,5.5),(-18.7,1.1),(-7.7,1.1),(-18.7,-1.1),(-14.3,-1.1),(-20.9,-3.3),(-3.3,-3.3),(-7.7,-5.5),(-18.7,-7.7),(-3.3,-7.7),(-16.5,-9.9),(-9.9,-9.9),(-14.3,-12.1),(-5.5,-12.1),(-12.1,-14.3),(-5.5,-18.7),(-3.3,-18.7),(-5.5,25.3),(-3.3,25.3),(-7.7,23.1),(-5.5,23.1),(-3.3,23.1),(-9.9,20.9),(-18.7,16.5),(-16.5,16.5),(-18.7,14.3),(-20.9,12.1),(-18.7,12.1),(-20.9,9.9),(-23.1,5.5),(-23.1,3.3),(-25.3,1.1),(-23.1,1.1),(-25.3,-1.1),(-23.1,-3.3),(-23.1,-5.5),(-23.1,-7.7),(-20.9,-7.7),(-20.9,-9.9),(-18.7,-9.9),(-20.9,-12.1),(-18.7,-12.1),(-18.7,-14.3),(-16.5,-14.3),(-18.7,-16.5),(-16.5,-16.5),(-14.3,-16.5),(-18.7,-18.7),(-16.5,-18.7),(-14.3,-18.7),(-12.1,-18.7),(-12.1,-20.9),(-9.9,-20.9),(-7.7,-20.9),(-7.7,-23.1),(-5.5,-23.1)] for x, y in green_points: adjusted_location = center_location.move(types.Point(x=x, y=y)) if pipette_20ul.current_volume == 0: pipette_20ul.aspirate(1, location_of_color(‘Green’)) dispense_and_jog(pipette_20ul, 1, adjusted_location) pipette_20ul.drop_tip()

- Post-Lab Questions — DUE BY START OF FEB 24 LECTURE
Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.

 The articule present a robotic automation platform designed to standardize and scale the production of three-dimensional epithelial tissues. The study addresses a key limitation in tissue engineering: the high variability and labor-intensive nature of manual organotypic tissue fabrication. By implementing a programmable liquid-handling robotic system compatible with platforms such as Opentrons, the authors automated critical steps including cell seeding, media exchange, hydrogel handling, and layered tissue assembly.
The articule present a robotic automation platform designed to standardize and scale the production of three-dimensional epithelial tissues. The study addresses a key limitation in tissue engineering: the high variability and labor-intensive nature of manual organotypic tissue fabrication. By implementing a programmable liquid-handling robotic system compatible with platforms such as Opentrons, the authors automated critical steps including cell seeding, media exchange, hydrogel handling, and layered tissue assembly.
The novelty of this work lies in demonstrating that accessible laboratory automation can be successfully applied to complex biofabrication workflows, not just routine pipetting tasks. The ReBiA system enabled the reproducible generation of multilayered epithelial tissues with functional characteristics comparable to manually produced models. This significantly reduces operator-dependent variability and increases throughput, which is essential for applications such as drug screening, disease modeling, and regenerative medicine research.
Overall, the study illustrates how low-cost, programmable automation tools can democratize advanced tissue engineering, improve reproducibility, and facilitate scalable biological manufacturing.
Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more. You may reference this week’s recitation slide deck for lab automation details.
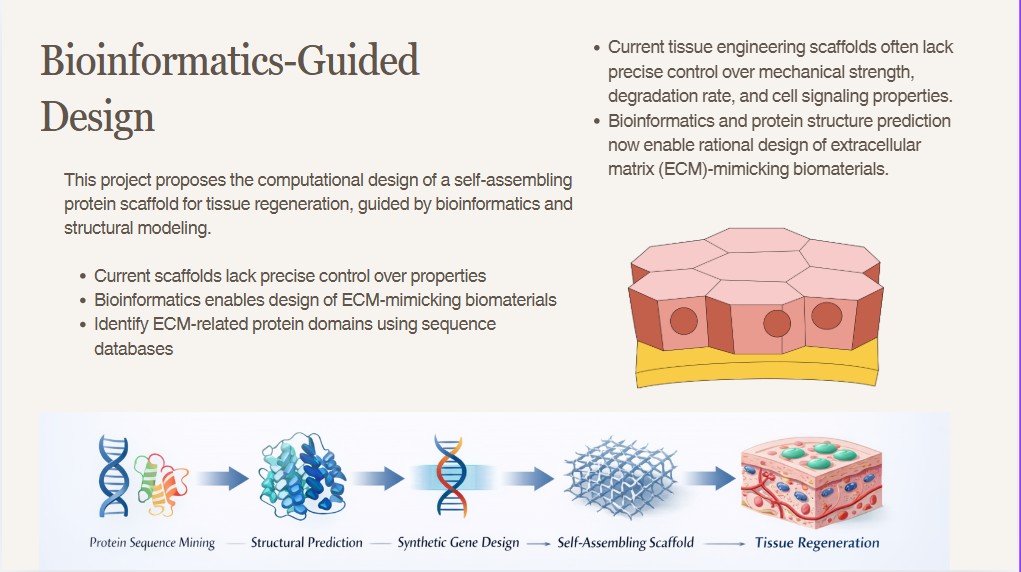
For my final project, I intend to use laboratory automation to standardize and optimize the production of fungal-based biodyes. One of the main challenges in working with filamentous fungi is variability in pigment yield due to small changes in media composition, pH, nutrient availability, and growth conditions. By implementing an automated liquid-handling system such as Opentrons, I aim to systematically control these variables and perform high-throughput screening of culture conditions to identify combinations that maximize and stabilize pigment production.
The automation workflow will include robotic preparation of media gradients, controlled inoculation of fungal spores into multi-well plates, and scheduled sampling for pigment quantification using absorbance measurements. Using the Opentrons Python API, I will program reproducible experimental matrices and spatially controlled dispensing patterns. This same precision can be extended to textile applications, where fungal inoculum or pigment precursors could be deposited in defined areas of fabric to enable localized growth and pattern formation. Custom 3D-printed holders may be designed to stabilize both culture vessels and textile substrates during automated handling. Experimental data will be digitally recorded to support structured analysis and optimization. Overall, automation will transform fungal biodye production into a scalable, reproducible, and spatially programmable biofabrication workflow.

Research sources: I used AI assistance ( Claude) to help organize the coordinate data extracted from the image. Specifically, the AI helped me structure the (x, y) coordinate lists correctly, convert them into a usable data format (e.g., arrays/DataFrame), and ensure they were properly scaled and centered for the experimental setup. Königer, L., Malkmus, C., Mahdy, D., Däullary, T., Götz, S., Schwarz, T., Gensler, M., Pallmann, N., Cheufou, D., Rosenwald, A., Möllmann, M., Groneberg, D., Popp, C., Groeber-Becker, F., Steinke, M., & Hansmann, J. (2024). ReBiA—Robotic Enabled Biological Automation: 3D Epithelial Tissue Production. Advanced Science, 11. https://doi.org/10.1002/advs.202406608.