Week 9 hw: cell free systems
Homework Part A: General and Lecturer-Specific Questions
- General homework questions Explain the main advantages of cell-free protein synthesis over traditional in vivo methods, specifically in terms of flexibility and control over experimental variables. Name at least two cases where cell-free expression is more beneficial than cell production.
Cell-free protein synthesis (CFPS) offers significant advantages over traditional in vivo methods, particularly in terms of flexibility and control over experimental variables. One of the main advantages is that CFPS operates as an open system, allowing direct manipulation of all reaction components. Researchers can precisely control variables such as DNA or mRNA concentration, ion composition (Mg²⁺, K⁺), cofactors, and environmental conditions like temperature and pH. In contrast, others systems are limited by cellular regulation and metabolic constraints.
CFPS also enables independent control of transcription and translation, making it possible to fine-tune each step of protein synthesis. This level of control is not achievable in living cells, where both processes are tightly coupled and regulated, rapid production of proteins and cell viability constraints, allowing the expression of proteins that would otherwise be toxic or harmful to living cells. Additionally, it allows the incorporation of non-natural amino acids and real-time optimization of reaction conditions.
__ Cases where CFPS is more beneficial than in vivo production: __
Toxic proteins
- Antimicrobial peptides: These proteins disrupt cellular membranes, which would damage or kill host cells during in vivo expression. In CFPS, there are no living cells, so toxicity is not a limitation.
Membrane proteins
- Membrane proteins contain hydrophobic regions that tend to aggregate or misfold in cellular environments. In CFPS, membrane mimetics such as liposomes or detergents can be added directly to support proper folding.
- CFPS also enables co-translational insertion into artificial membranes, improving protein stability and functionality compared to in vivo systems.
- Describe the main components of a cell-free expression system and explain the role of each component.
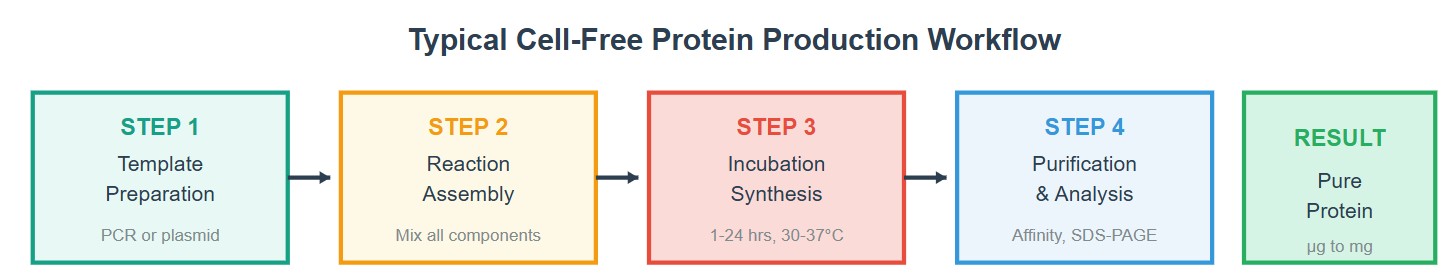

Step 1: Template Preparation
In this step, the genetic material encoding the protein of interest is prepared. This can be in the form of plasmid DNA or PCR products. The quality and concentration of the template are critical, as they directly affect transcription efficiency and overall protein yield.
Step 2: Reaction Assembly
All necessary components of the cell-free system are combined. This includes the cell extract (translation machinery), amino acids, energy sources, cofactors, and the DNA or mRNA template. Because the system is open, each component can be optimized to improve protein expression.
Step 3: Incubation / Synthesis
The reaction mixture is incubated under controlled conditions (typically 30–37°C). During this stage, transcription (if DNA is used) and translation occur, leading to protein synthesis. Reaction time and conditions can be adjusted to balance yield and protein folding.
Step 4: Purification & Analysis
The synthesized protein is purified using methods such as affinity chromatography. Analytical techniques like SDS-PAGE are then used to evaluate protein size, purity, and expression level.
Result: Pure Protein
The final outcome is the production of the target protein, typically in microgram to milligram quantities. The quality and functionality of the protein depend on the optimization of previous steps.
- Why is energy provision regeneration critical in cell-free systems? Describe a method you could use to ensure continuous ATP supply in your cell-free experiment.


Energy provision and regeneration are critical in cell-free protein synthesis (CFPS) because protein production is a highly energy-demanding process. Both transcription and translation require continuous consumption of ATP and GTP for processes such as nucleotide polymerization, tRNA charging, and ribosome translocation.
In a closed reaction environment like CFPS, the initial supply of ATP is rapidly depleted. Without an efficient energy regeneration system, this leads to:
- Premature termination of protein synthesis
- Reduced protein yield
- Accumulation of inhibitory byproducts
Method to Ensure Continuous ATP Supply
One common strategy is the use of a phosphoenolpyruvate (PEP)-based energy regeneration system:
- Phosphoenolpyruvate (PEP) acts as a high-energy phosphate donor.
- In the presence of the enzyme pyruvate kinase, PEP transfers a phosphate group to ADP, regenerating ATP.
Reaction: PEP + ADP → Pyruvate + ATP
This system is effective because it continuously replenishes ATP during the reaction, allowing protein synthesis to proceed for longer periods and increasing overall yield.
- Compare prokaryotic versus eukaryotic cell-free expression systems. Choose a protein to produce in each system and explain why. How would you design a cell-free experiment to optimize the expression of a membrane protein? Discuss the challenges and how you would address them in your setup.
| Feature | Prokaryotic System (E. coli lysate) | Eukaryotic System (HEK293 lysate) |
|---|---|---|
| Yield | High | Moderate to low |
| Speed | Fast (hours) | Slower |
| Cost | Low | Higher |
| Folding | Limited (simple proteins) | Improved folding for complex proteins |
| Post-translational modifications | Not available | Present (e.g., disulfide bonds, glycosylation) |
| Cellular mimicry | Low | High (closer to human environment) |
| Best use | Simple, non-modified proteins | Complex, functional human proteins |
¨System Hybridization and Standardization The future lies in bridging the gap between prokaryotic speed and eukaryotic fidelity. Researchers are increasingly exploring hybrid systems where PTM machinery (e.g., purified glycosylation enzymes or dedicated PDI) is supplemented into the high-yield E. coli lysate.¨
- How would you design a cell-free experiment to optimize the expression of a membrane protein? Discuss the challenges and how you would address them in your setup.
Experimental Design
First, the reaction mixture must be assembled, including the cell extract, amino acids, energy system, and the DNA template encoding the membrane protein. At this stage, it is critical to include membrane mimetics such as liposomes, nanodiscs, or mild detergents (e.g., DDM), which will provide a hydrophobic environment.
Next, the reaction conditions should be optimized. This includes adjusting Mg²⁺ and salt concentrations to stabilize the translation machinery, as well as selecting an appropriate temperature (often lower temperatures) to slow down translation and improve protein folding.
During the synthesis phase, co-translational insertion should be promoted by ensuring that membrane mimetics are present from the beginning. This allows the protein to insert into the artificial membrane as it is being synthesized, reducing aggregation.
Additionally, molecular chaperones can be added to assist folding and increase protein stability.
Finally, the expression rate should be controlled by adjusting DNA concentration or transcriptional activity. Slower expression often leads to better folding and higher functional yield.
On the other hand, one major challenge is the hydrophobic nature of membrane proteins, which causes aggregation in aqueous environments. This can be addressed by adding membrane mimetics that stabilize these regions.
Another challenge is the absence of a natural lipid bilayer, which is essential for correct protein folding. This is overcome by using liposomes or nanodiscs to mimic the membrane environment.
Misfolding is also a concern, especially at high expression rates. This can be mitigated by lowering the temperature and controlling the rate of protein synthesis.
Low solubility and instability can affect protein yield and functionality. The use of chaperones and optimized reaction conditions helps improve protein stability and folding efficiency.
- Imagine you observe a low yield of your target protein in a cell-free system. Describe three possible reasons for this and suggest a troubleshooting strategy for each.
1. Codon bias
Reason:
Codon bias occurs when the gene contains codons that are rarely used in the expression system, leading to low availability of the corresponding tRNAs. This slows down translation or can even stop protein synthesis, reducing yield.
Troubleshooting strategy:
- Optimize the gene sequence to replace rare codons with more frequently used ones
- Lower the reaction temperature (e.g., to ~30°C) to slow translation and allow tRNA recruitment
- Use a different expression system with a more compatible tRNA pool (e.g., eukaryotic system)
2. Protein misfolding
Reason:
Even if the protein is produced, it may not fold correctly due to lack of chaperones or post-translational modifications. Misfolded proteins tend to aggregate, reducing the amount of functional protein.
Troubleshooting strategy:
- Add molecular chaperones to assist protein folding
- Use a eukaryotic system ( wheat germ extract) if the protein requires complex folding or PTMs
- Adjust conditions to favor proper folding ( lower temperature)
3. Transcription-translation imbalance (phage polymerase issue)
Reason:
Phage RNA polymerases transcribe mRNA much faster than ribosomes can translate it. This leads to uncoupling, leaving mRNA exposed to degradation or forming secondary structures, which reduces protein yield.
Troubleshooting strategy:
- Reduce reaction temperature ( down to ~20°C) to slow transcription
- Improve coupling between transcription and translation
- Optimize reaction conditions to stabilize mRNA
References:
- Brookwell, A., Oza, J., & Caschera, F. (2021). Biotechnology Applications of Cell-Free Expression Systems. Life, 11. https://doi.org/10.3390/life11121367.
- https://www.cytion.com/es/Acerca-de-Cytion/Centro-de-conocimiento/Blog/Sistemas-sin-celulas-para-la-produccion-de-proteinas-Ventajas-sobre-las-celulas-vivas/
- Gregorio, N., Levine, M., & Oza, J. (2019). A User’s Guide to Cell-Free Protein Synthesis. Methods and Protocols, https://doi.org/10.3390/mps2010024.
- Dondapati, S., Stech, M., Zemella, A., & Kubick, S. (2020). Cell-Free Protein Synthesis: A Promising Option for Future Drug Development. Biodrugs, 34, 327 - 348. https://doi.org/10.1007/s40259-020-00417-y.
- https://id.elsevier.com/as/authorization.oauth2?platSite=SD%2Fscience&additionalPlatSites=GH%2Fgeneralhospital%2CLS%2FLS%2CMDY%2Fmendeley%2CSC%2Fscopus%2CRX%2Freaxys&scope=openid%20email%20profile%20els_auth_info%20els_idp_info%20els_idp_analytics_attrs%20urn%3Acom%3Aelsevier%3Aidp%3Apolicy%3Aproduct%3Ainst_assoc&response_type=code&redirect_uri=https%3A%2F%2Fwww.sciencedirect.com%2Fuser%2Fidentity%2Flanding&authType=SINGLE_SIGN_IN&prompt=none&client_id=SDFE-v4&state=retryCounter%3D0%26csrfToken%3Df4ffded5-e62a-4dfd-b41f-b0d2a17110e1%26idpPolicy%3Durn%253Acom%253Aelsevier%253Aidp%253Apolicy%253Aproduct%253Ainst_assoc%26returnUrl%3D%252Fscience%252Farticle%252Fpii%252FS0021925824023524%26prompt%3Dnone%26cid%3Datn-8cc5fa75-436c-4a48-b4bf-1affa3324a4b
- González-Ponce, K., Celaya-Herrera, S., Mendoza-Acosta, M., & Casados-Vázquez, L. (2025). Cell-Free Systems and Their Importance in the Study of Membrane Proteins. The Journal of Membrane Biology, 258, 15 - 28. https://doi.org/10.1007/s00232-024-00333-0.
- Smolskaya, S., Logashina, Y., & Andreev, Y. (2020). Escherichia coli Extract-Based Cell-Free Expression System as an Alternative for Difficult-to-Obtain Protein Biosynthesis. International Journal of Molecular Sciences, 21. https://doi.org/10.3390/ijms21030928.
Homework question from Kate Adamala
Design an example of a useful synthetic minimal cell as follows:
Synthetic Cell: DePathogen Cell
Pick a function and describe it
The synthetic cell detects pathogenic fungi and neutralizes their virulence by delivering CRISPR-Cas9 complexes that silence specific virulence genes.
What would your synthetic cell do? What is the input and what is the output?
- What it does: It senses a fungal pathogenic signal and responds by releasing CRISPR components that edit the fungal genome.
- Input: Fungal siderophores (e.g., ferrioxamine B)
- Output (SMC): Release of CRISPR-Cas9 + sgRNA complexes
- System output: Fungus continues to grow but loses its ability to infect (reduced virulence)

Could this function be realized by cell-free Tx/Tl alone, without encapsulation?
No. Encapsulation is required to:
- Isolate sensing and response in a defined unit
- Protect CRISPR components
- Enable controlled release only after signal detection
Could this function be realized by genetically modified natural cell?
Partially. Natural cells (Trichoderma) can inhibit fungi, but CRISPR-based targeting provides higher specificity and programmability compared to metabolite-based inhibition.
Describe the desired outcome of your synthetic cell operation
In the presence of pathogenic fungi, the synthetic cells release CRISPR complexes that edit virulence genes, resulting in fungi that grow normally but are no longer pathogenic.
Design of the Synthetic Cell
What would the membrane be made of?
- POPC (phosphatidylcholine)
- Ergosterol (fungal-like stability)
- POPG (charge and membrane functionality)
What would you encapsulate inside?
- Cell-free Tx/Tl system
- Cas9 protein (pre-synthesized)
- sgRNAs targeting fungal virulence genes
- Mg²⁺ (cofactor for Cas9 activity)
- Energy components (ATP regeneration system)
Which organism will your Tx/Tl system come from?
- Fungal system (Aspergillus niger)
- Chosen for compatibility with fungal regulatory elements and gene expression
- A bacterial system would be less suitable due to differences in regulatory mechanisms
How will your synthetic cell communicate with the environment?
Input: Siderophores diffuse into the vesicle
Output: CRISPR complexes are exported through a membrane transporter
Key transporter gene: foxA (A. niger)
Experimental Details
Lipids
- POPC
- Ergosterol
- POPG
Genes
- foxR → siderophore sensor
- cas9 (Streptococcus pyogenes) → CRISPR nuclease
- sgRNA_tsc2 → targets virulence gene (tsc2)
- sgRNA_fox1 → targets toxin-related pathways
- foxA → siderophore transporter/export
- chiA → chitinase (facilitates fungal penetration)
- gfp → reporter
How will you measure the function of your system?
- GFP fluorescence (monitor activation of the system)
- PCR or sequencing to confirm gene editing (e.g., tsc2 disruption)
- Plant infection assays (compare infection vs control)
- Quantification of fungal virulence reduction
References: -Dobrzyński, J., Jakubowska, Z., Kulkova, I., Kowalczyk, P., & Kramkowski, K. (2023). Biocontrol of fungal phytopathogens by Bacillus pumilus. Frontiers in Microbiology, 14. https://doi.org/10.3389/fmicb.2023.1194606. -Ayaz, M., Li, C., Ali, Q., Zhao, W., Chi, Y., Shafiq, M., Ali, F., Yu, X., Yu, Q., Zhao, J., Yu, J., Qi, R., & Huang, W. (2023). Bacterial and Fungal Biocontrol Agents for Plant Disease Protection: Journey from Lab to Field, Current Status, Challenges, and Global Perspectives. Molecules, 28. https://doi.org/10.3390/molecules28186735. -Mayer, F., & Kronstad, J. (2017). Disarming Fungal Pathogens: Bacillus safensis Inhibits Virulence Factor Production and Biofilm Formation by Cryptococcus neoformans and Candida albicans. mBio, 8. https://doi.org/10.1128/mbio.01537-17.
Homework question from Peter Nguyen
Freeze-dried cell-free systems can be incorporated into all kinds of materials as biological sensors or as inducible enzymes to modify the material itself or the surrounding environment. Choose one application field — Architecture, Textiles/Fashion, or Robotics — and propose an application using cell-free systems that are functionally integrated into the material. Answer each of these key questions for your proposal pitch:
Application Field: Textiles / Fashion
One-sentence pitch
A smart textile embedded with freeze-dried cell-free systems that activates upon moisture to produce visible color changes in response to environmental conditions.
How will the idea work?
The textile contains freeze-dried cell-free transcription/translation (Tx/Tl) systems embedded within microcapsules integrated into the fabric fibers. When exposed to moisture (e.g., sweat, rain, or humidity), the system rehydrates and becomes active.
Upon activation, specific environmental signals such as pH changes, pollutants, or metal ions trigger gene expression that leads to the production of pigments or color-changing molecules. This results in a visible and localized color change directly on the textile.
The system can be programmed with different genetic circuits to respond to distinct stimuli, allowing the fabric to function as a wearable biosensor or dynamic aesthetic material.
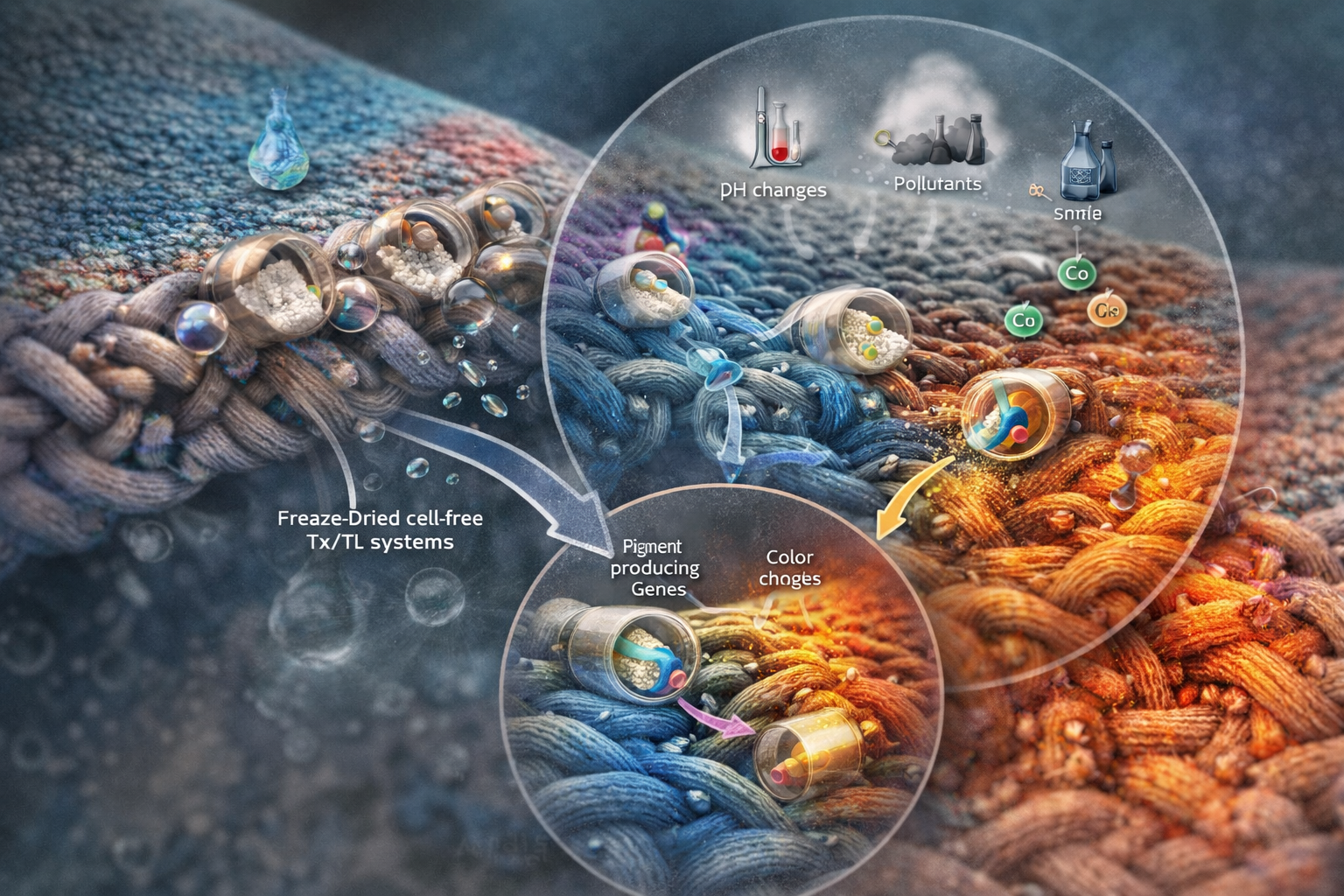

What societal challenge or market need will this address?
This technology addresses the growing demand for sustainable, functional, and responsive textiles. It can be used for:
- Environmental monitoring (pollution detection)
- Health-related wearables (esweat pH indicating stress or dehydration)
- Fashion innovation through dynamic, customizable clothing
Additionally, it reduces reliance on synthetic dyes by enabling on-demand biological pigment production, supporting more sustainable textile practices.
How to address limitations of cell-free systems?
- Activation control: The system is designed to activate only upon hydration (e.g., sweat or water exposure), preventing premature reactions.
- Stability: Freeze-drying (lyophilization) significantly increases shelf life and allows storage at room temperature.
- One-time use limitation: The textile can be designed as either disposable (e.g., medical patches) or modular, with replaceable biosensing patches embedded in garments.
- Protection: Microencapsulation within polymer or hydrogel matrices protects the biological components from degradation and environmental stress.
Homework question from Ally Huang
Freeze-dried cell-free reactions have great potential in space, where resources are constrained. As described in my talk, the Genes in Space competition challenges students to consider how biotechnology, including cell-free reactions, can be used to solve biological problems encountered in space. While the competition is limited to only high school students, your assignment will be to develop your own mock Genes in Space proposal to practice thinking about biotech applications in space!
For this particular assignment, your proposal is required to incorporate the BioBits® cell-free protein expression system, but you may also use the other tools in the Genes in Space toolkit (the miniPCR® thermal cycler and the P51 Molecular Fluorescence Viewer). For more inspiration, check out https://www.genesinspace.org/ .
Background (≤100 words)
During long-duration space missions, astronauts depend on stored, freeze-dried food that undergoes nutrient degradation over time. Vitamin B12 (cobalamin), an essential cofactor for DNA synthesis and neurological function, is particularly susceptible to depletion, increasing the risk of anemia and cognitive impairment. Due to limited resupply capabilities in deep space missions, in situ production of essential micronutrients is highly desirable. Cell-free protein synthesis systems such as BioBits® offer a stable, lyophilized platform for enzymatic production under resource-constrained conditions, enabling decentralized and on-demand biosynthesis of critical compounds in space environments.
Molecular / Genetic Target (≤30 words)
Gene cobA, encoding uroporphyrinogen III methyltransferase, a key enzyme in early cobalamin biosynthesis.
Relevance to the Challenge (≤100 words)
Cobalamin biosynthesis is a complex, multi-step pathway requiring specialized enzymes. The cobA gene encodes a methyltransferase that catalyzes an early and essential step in corrin ring formation. By expressing cobA in a cell-free system, it is possible to reconstruct part of the cobalamin biosynthetic pathway from precursor molecules. Monitoring the activity of this enzyme provides insight into the feasibility of modular vitamin production in space. This approach links molecular enzymatic function to astronaut nutrition, offering a strategy for supplementing degraded food supplies with freshly synthesized cofactors.
Hypothesis / Research Goal (≤150 words)
We hypothesize that a lyophilized BioBits® cell-free system expressing cobA can catalyze the methylation of uroporphyrinogen III, generating intermediates of the cobalamin biosynthesis pathway under space-relevant conditions. This enzymatic activity can be indirectly quantified through fluorescence-based assays targeting pathway intermediates or coupled reporter systems. The rationale is that cell-free systems retain sufficient enzymatic functionality to support partial metabolic pathway reconstruction without the need for living cells. By combining miniPCR® amplification of the cobA gene with cell-free expression, this system enables rapid enzyme production in situ. The P51 Molecular Fluorescence Viewer will be used to detect reaction outputs. The goal is to establish a proof-of-concept for modular, cell-free biosynthesis of essential micronutrients during long-duration space missions.
Experimental Plan (≤100 words)
BioBits® reactions will be prepared with a plasmid encoding cobA and supplied with uroporphyrinogen III as substrate. Controls include reactions without DNA, without substrate, and heat-inactivated systems. Reactions will be incubated at 37°C for 4 hours. Enzymatic activity will be assessed באמצעות fluorescence-based detection of methylated intermediates or coupled reporter assays, visualized using the P51 viewer. Relative fluorescence intensity will be quantified to compare activity across conditions and confirm functional enzyme expression.
Homework Part B: Individual Final Project
We’d like students to start exploring their final project in depth this week! Of your three Aims, for this week you should have at least Aim 1 decided and written down.


Prepare your first DNA order and put it in the “Twist (MIT)” or “Twist (Nodes)” tab of the 2026 HTGAA Ordering: DNA, Reagents, Consumables spreadsheet, as appropriate.
https://benchling.com/mar0502/f/lib_QjyXnjm74R-biodyes/seq_sMk1g9vpft-fusarium/edit