Week 11 hw: Building genomes
Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork Next year
- Part B: Cell-Free Protein Synthesis | Cell-Free Reagents
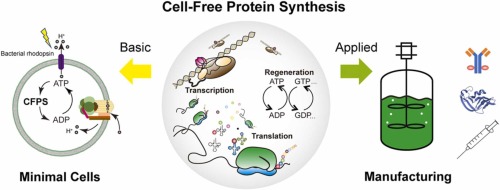

Referencing the cell-free protein synthesis reaction composition (the middle box outlined in yellow on the image above, also listed below), provide a 1-2 sentence description of what each component’s role is in the cell-free reaction.
E. coli Lysate
BL21 (DE3) Star Lysate (includes T7 RNA Polymerase): Provides the cellular machinery (ribosomes, enzymes, cofactors) required for transcription and translation. T7 RNA polymerase enables efficient transcription of genes under a T7 promoter.
Salts/Buffer
Potassium Glutamate: Maintains ionic strength and mimics intracellular conditions, supporting proper enzyme function and protein synthesis.
HEPES-KOH pH 7.5: Maintains a stable physiological pH, preventing loss of enzymatic activity and preserving ribosomal integrity during prolonged incubation.
Magnesium Glutamate: Provides Mg²⁺ ions required for ribosome assembly, tRNA charging, and stabilization of ATP and NTPs; directly impacts translation efficiency and fidelity.
Potassium phosphate monobasic: Acts as the acidic component of the phosphate buffer system, helping maintain pH stability during the reaction. It also supplies inorganic phosphate required for ATP regeneration and metabolic reactions.
Potassium phosphate dibasic: Acts as the basic component of the phosphate buffer system, balancing the monobasic form to tightly regulate pH. It also contributes phosphate ions and supports optimal ionic conditions for enzyme activity.
Energy / Nucleotide System
Ribose: Feeds into the pentose phosphate and nucleotide salvage pathways to generate ribose-5-phosphate, a precursor for de novo and salvage nucleotide synthesis.
Glucose: Drives glycolytic activity in the lysate, enabling continuous ATP regeneration and maintaining redox balance through endogenous metabolic enzymes.
AMP: Serves as a precursor for ATP synthesis through phosphorylation pathways, helping regenerate the main energy currency required for transcription and translation.
CMP: Acts as a precursor for CTP, which is required for RNA synthesis during transcription.
GMP: Functions as a precursor for GTP, which is essential for both RNA synthesis and translation elongation steps.
UMP: Serves as a precursor for UTP, another nucleotide required for RNA synthesis during transcription.
Guanine: Provides an additional substrate for purine salvage pathways, enhancing GMP/GTP pools required for transcription and translation elongation.
Translation Mix (Amino Acids)
17 Amino Acid Mix: Supplies most amino acids required for polypeptide elongation, supporting continuous translation.
Tyrosine: Added separately due to its low solubility and tendency to precipitate, ensuring it does not become rate-limiting.
Cysteine: Included independently because it is chemically reactive and prone to oxidation, which can otherwise limit protein synthesis.
Additives
Nicotinamide: Acts as a precursor to NAD⁺, supporting redox cycling and metabolic reactions required for sustained ATP generation in long-duration reactions
Backfill: Used to adjust final reaction volume while preventing nucleic acid degradation by contaminating nucleases.
Nuclease Free Water
- Describe the main differences between the 1-hour optimized PEP-NTP master mix and the 20-hour NMP-Ribose-Glucose master mix shown in the Google Slide above. (2-3 sentences)
The PEP–NTP system directly supplies high-energy phosphate donors (PEP) and fully phosphorylated nucleotides (NTPs), enabling immediate and high-rate transcription and translation. However, this leads to rapid accumulation of inhibitory byproducts (e.g., inorganic phosphate) and depletion of energy sources, limiting reaction lifespan to ~1 hour.
In contrast, the NMP–Ribose–Glucose system relies on endogenous metabolic pathways within the lysate to regenerate ATP and NTPs from low-energy precursors (NMPs, ribose, and glucose). This reduces byproduct accumulation and allows continuous energy recycling, resulting in slower but significantly prolonged protein synthesis (~20 hours) with improved overall yield.
- Part C: Planning the Global Experiment | Cell-Free Master Mix Design
Given the 6 fluorescent proteins we used for our collaborative painting, identify and explain at least one biophysical or functional property of each protein that affects expression or readout in cell-free systems. (Hint: options include maturation time, acid sensitivity, folding, oxygen dependence, etc) (1-2 sentences each)
sfGFP: Superfolder GFP is engineered for robust folding, allowing efficient fluorescence even in partially unfavorable conditions. This makes it highly reliable in cell-free systems where folding machinery may be limited.
mRFP1: Has a slow maturation time, meaning chromophore formation takes longer, which can delay fluorescence readout in shorter reactions.
mKO2: Is relatively acid-sensitive, so fluorescence intensity decreases if the reaction pH drops over time due to metabolic byproducts.
mTurquoise2: Has high quantum yield but requires efficient folding, making it sensitive to translation conditions and potentially prone to misfolding in cell-free systems.
mScarlet_I: Is is optimized for fast maturation and high brightness, but its performance can still depend on proper folding and oxygen availability for chromophore formation.
Electra2:Is is oxygen-independent but depends on availability of flavin cofactors (e.g., FMN/FAD) for fluorescence, which may be limiting in cell-free systems.
The amino acid sequences are shown in the HTGAA Cell-Free Benchling folder.
- Create a hypothesis for how adjusting one or more reagents in the cell-free mastermix could improve a specific biophysical or functional property you identified above, in order to maximize fluorescence over a 36-hour incubation. Clearly state the protein, the reagent(s), and the expected effect.
For mKO2, which is sensitive to acidic conditions, increasing the concentration of buffering agents such as HEPES-KOH and potassium phosphate in the cell-free mastermix will help maintain a stable pH throughout the 36-hour incubation, preventing acidification caused by metabolic byproducts. By stabilizing the pH, the protein structure and chromophore environment will be preserved, leading to improved fluorescence intensity and stability over time.
Part D: Build-A-Cloud-Lab | (optional) Bonus Assignment Assignees for this section MIT/Harvard students Optional Committed Listeners Optional Ginkgo Nebula Cloud Laboratory Rendering, 2025 Ginkgo Nebula Cloud Laboratory Rendering, 2025
Use this simulation tool to create an interesting looking cloud lab out of the Ginkgo Reconfigurable Automation Carts. This is just a minimal implementation so far, but I would love to see some fun designs! Tip Note from Ronan: If you are interested in helping me build out future HTGAA cloud lab software, please fill out this form!