Week 11 HW: Bioproduction & Cloudlabs
Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork
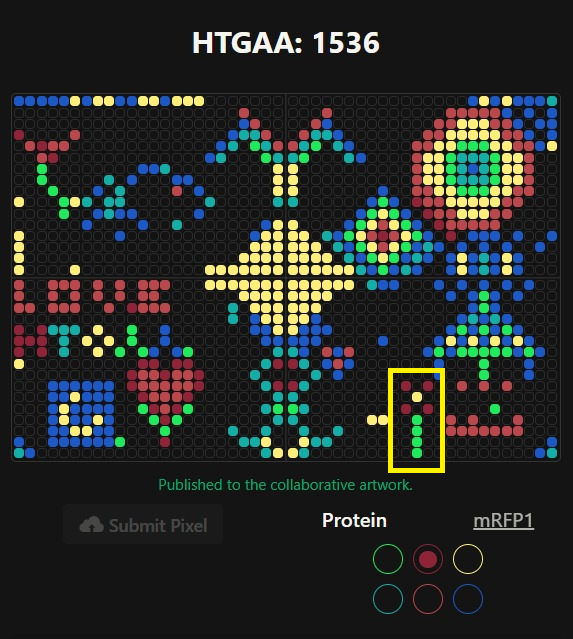

In the initial artwork (before the complete design was changed :()) I made that little flower using mRFP, sfGFP and mKO2.
I actually loved this project because we had the opportunity to contribute to the HTGAA effort. I would also like to say that Ronan’s page is easy to understand and use, thanks Ronan!
One thing I didn’t like was the fact that when this project was launched and we got the email, we started making our own figures but then (close to the deadline) some other students changed everything on the original design (can’t blame them though I know it was colaborative!). Maybe for next year the deadline could be shorter so it stays the original design or there’s not much time to re-design it?
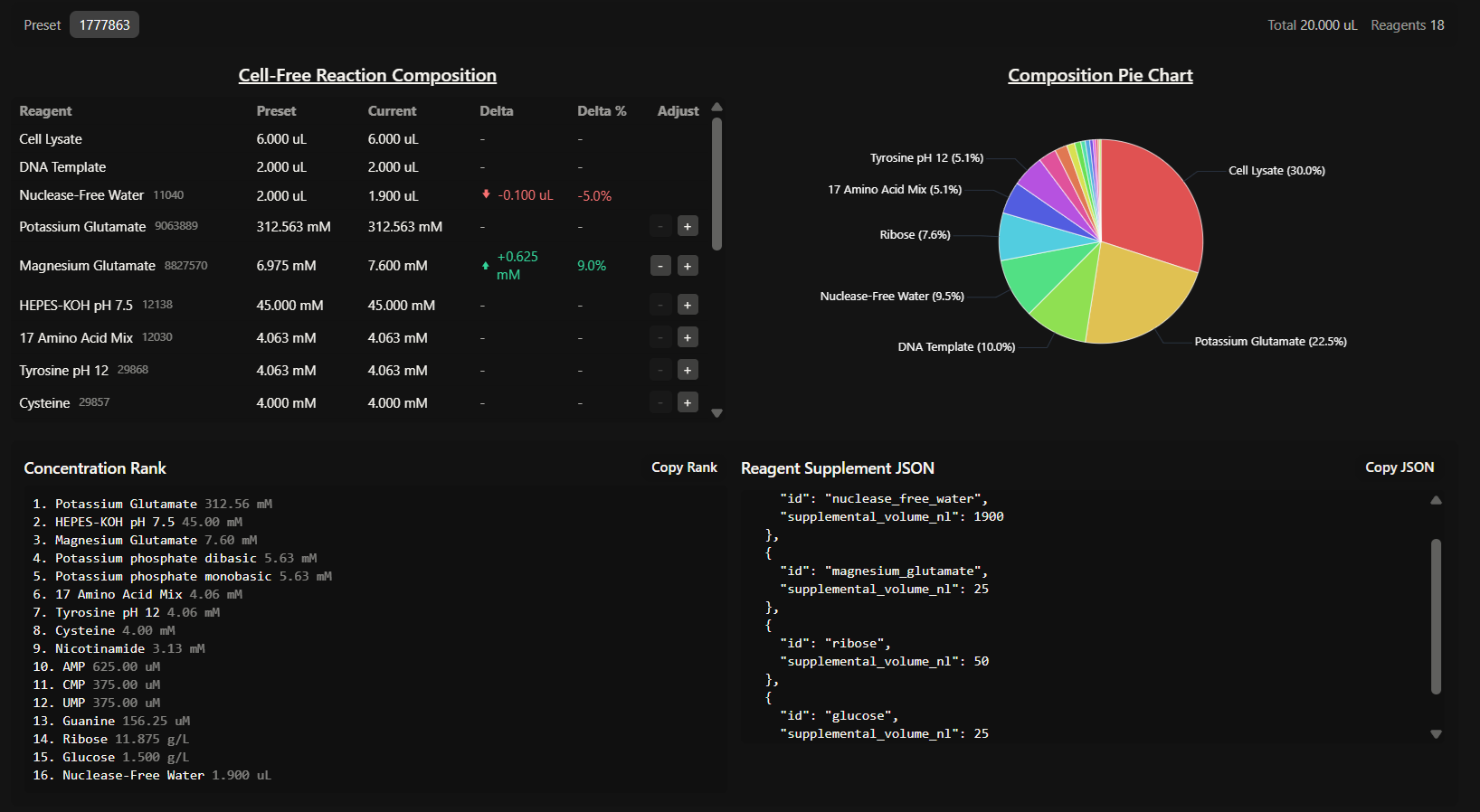
Part B: Cell-Free Protein Synthesis | Cell-Free Reagents
- Referencing the cell-free protein synthesis reaction composition (the middle box outlined in yellow on the image above, also listed below), provide a 1-2 sentence description of what each component’s role is in the cell-free reaction.
a. E. coli Lysate
• BL21 (DE3) Star Lysate (includes T7 RNA Polymerase): bacterial cell extract containing all the molecular machinery needed for protein synthesis
b. Salts/Buffer
• Potassium Glutamate: maintains the ionic strength of the solution
• HEPES-KOH pH 7.5: buffer that keeps pH stable at 7.5 (close to physiological conditions)
• Magnesium Glutamate: provides Mg+2 ions essential for ribosome assembly and enzymatic activity
• Potassium phosphate monobasic
• Potassium phosphate dibasic: these two forms together act a buffer providing phosphate groups
c. Energy / Nucleotide System
• Ribose: backbone for building nucleotides
• Glucose: energy source
• AMP
• CMP
• GMP
• UMP: these four nucleoside monophosphates are necessary for RNA synthesis
• Guanine: can be converted into GMP and then GTP
d. Translation Mix (Amino Acids)
• 17 Amino Acid Mix: provides 17 of the 20 standard amino acids for protein building
• Tyrosine
• Cysteine: tyrosine and cysteine are added separately because are prone to oxidation and poorly soluble at neutral pH
e. Additives
• Nicotinamide: precursor to NAD+, necessary coenzyme for metabolic reactions
f. Backfill
• Nuclease Free Water: to bring the reaction volume to the final desired amount, plus it has no enzymes that could degrade RNA or DNA in the reaction.
- Describe the main differences between the 1-hour optimized PEP-NTP master mix and the 20-hour NMP-Ribose-Glucose master mix shown in the Google Slide above. (2-3 sentences).
The main difference is in the energy/nucleotide system: the 1-hour PEP-NTP mis provides energy through PEP-Mono and Maltodextrin and fully built NTPs ready to use immediately, while the 20-hour NMP-Ribose-Glucose system uses Ribose + Glucose and NMPs as simpler precursors that the enzymes convert into usable NTPs over time. The 1-hour system also includes several additives absent in the 20-hour version: spermidine to stabilize ribosomes, DMSO to help with solubility, cAMP as a gene expression regulator, and folinic acid that supports amino acid synthesis.
- Bonus question: How can transcription occur if GMP is not included but Guanine is?
Enzymes present in the E. coli lysate can take the free guanine and attach it to a ribose-phosphate group to synthesize GMP on its way. That GMP can be phosphorylated to GDP and then GTP.
Part C: Planning the Global Experiment – Cell-Free Master Mix Design
- Given the 6 fluorescent proteins we used for our collaborative painting, identify and explain at least one biophysical or functional property of each protein that affects expression or readout in cell-free systems. (Hint: options include maturation time, acid sensitivity, folding, oxygen dependence, etc) (1-2 sentences each).
• sfGFP: this Green fluorescent protein is very important because it has an exceptional folding robustness, so it was engineered so that its folding is not affected in any conditions.
• mRFP1: the biggest limitation is its low brightness, so it is needed a lot of protein to get a decent readout. In addition, some molecules get stuck during maturation, which makes the signal even lower.
• mKO2: this protein matures very slowly: it takes about 2 hours just to reach half of its final brightness, so it takes much more time to analyze the final results
• mTurquoise2: it is one of the most reliable fluorescent proteins to use as a reporter in cell-free systems because it basically converts almost every proton it absorbs into emitted fluorescence, so even small amounts of expressed protein will produce a strong signal.
• mScarlet_I: this protein has been engineered to have a marked maturation acceleration in cells, so compared to its sister protein mScarlet, this one is faster to light up in a cell-free reaction
• Electra2: this blue fluorescent protein tends to aggregate forming non-fluorescent aggregates instead of properly folding. It requires molecular oxygen for chromophore maturation, which must be present in sufficient quantity in the reaction.
- Create a hypothesis for how adjusting one or more reagents in the cell-free mastermix could improve a specific biophysical or functional property you identified above, in order to maximize fluorescence over a 36-hour incubation. Clearly state the protein, the reagent(s), and the expected effect.
My hypothesis is that increasing the concentrations of Ribose, Glucose and Magnesium Glutamate in the NMP-Ribose mastermix will maximize mKO2 fluorescence over a 36-hour incubation. In a long-term reaction like this, the main issues are the energy depletion and byproduct inhibition, specifically the accumulation of inorganic phosphate which can inhibit key enzymes and slow down the reaction. By increasing ribose and glucose we provide a larger and longer-lasting carbon and energy soiurce, preventing early termination of transcription and translation. Additionally, increasing magnesium glutamate is crucial because Mg+2 ions are essential cofactors for ribosome stability and RNA polymerase activity, and in extended incubation periods free Mg+2 becomes sequestered by those accumulated phosphate groups, reducing its availability in the reaction. mKO2 is particularly useful for rthis extended format because of its slow maturation half-time and its long-lived non-fluorescent intermediate state, meaning a significant fraction of the protein will still be completing chromophore formation past the 20-hour mark. The expected effect is a slow-burn metabolic rate that keeps the lysate active long enough for mKO2 to fully mature, resulting in a higher cumulative fluorescent yield at 36 hours compared to the standard 20-hour mix.
- he second phase of this lab will be to define the precise reagent concentrations for your cell-free experiment. You will be assigned artwork wells with specific fluorescent proteins and receive an email with instructions this week (by April 24). You can begin composing master mix compositions here.
I tested and changed the reagent master mix compositions according to my hypothesis, increasing concentrations of Glucose, Ribose and Magnesium Glutamate as shown in the image: