Week 3 HW: Lab Automation
Python Script for Opentrons Artwork
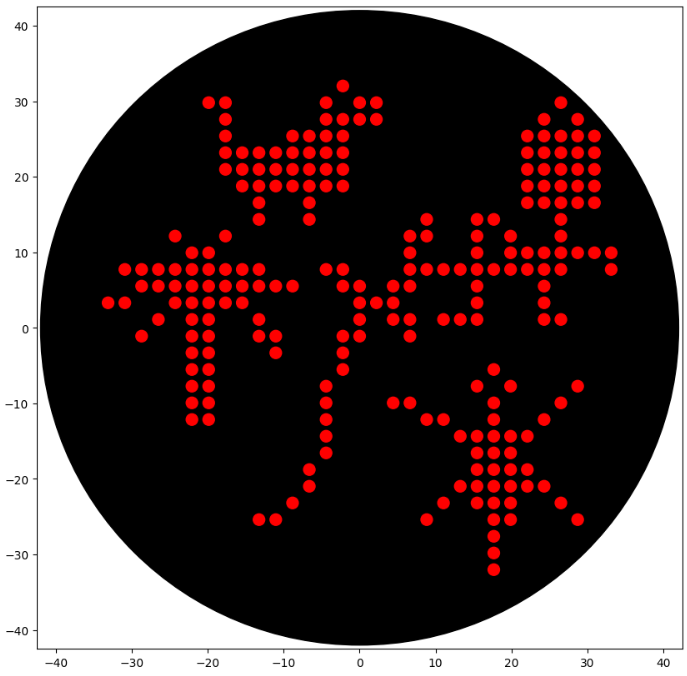
Here’s my HTGAA 2026 Opentrons Art Python Script Submission.
The artistic design I created using the GUI is available here.

I heavily used the “Example 7 Microbial Earth” by Dominika Wawrzyniak, using pixels loaded from an external resource (a CSV file hosted on my GitHub page).
I used Dominika’s well documented Notion page from HTGAA21 to understand the code and replicate it for my case. I used Gemini assistance only to debug minor typos and syntax errors, and to identify which packages to import to execute the code.
Like Dominika Wawrzyniak, I planned to introduce more colors, like in the image I generated in the Automation Art Interface. However, implementing this design into code turned out to be more difficult and tedious than anticipated, so I left it as one color (red).
Post-Lab Questions
Question 1
The paper “High-throughput experimentation for discovery of biodegradable polyesters” (Fransen et al., 2023) uses an Opentrons 1st-generation robot to automate a high-throughput biodegradation assay based on the clear-zone technique.
The researchers synthesized 642 polyesters and polycarbonates and tested their biodegradability using a clear-zone assay with Pseudomonas lemoignei. The Opentrons robot was repurposed as an automated imaging platform to capture time-lapse images of polymer degradation in 12-well plates, enabling consistent, large-scale monitoring over 13 days.
This automation allowed rapid generation of a large biodegradation dataset and supported machine learning models to predict polymer degradability from chemical structure.
Question 2
High-throughput screening of bacterial isolates for PHA production is traditionally extremely time-consuming and labor-intensive, requiring manual handling of hundreds of colonies across multiple conditions. For my final project, I plan to use an Opentrons OT-2 liquid-handling robot to automate this workflow, dramatically increasing throughput, reproducibility, and consistency compared to manual methods I used during my master’s.
Isolates will be spotted in triplicate on 60-sector plates, maintaining identical indexed positions across all plates for direct comparison. Viability will first be confirmed on LB agar, and isolates will then be inoculated onto mineral medium (MM; Ramsay et al., 1990) agar plates supplemented with individual carbon sources at 10% v/v to reach typical screening concentrations.
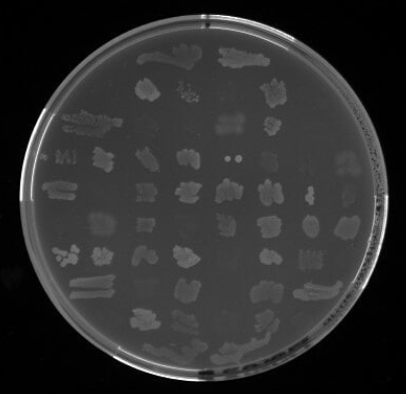
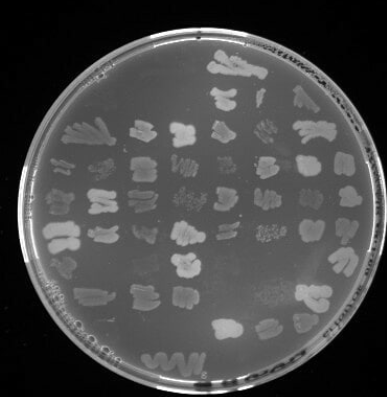
PHA production and bacterial growth will be assessed using a two-step staining workflow. First, Sudan Black B (0.02% in 96% ethanol, followed by ethanol washes) will identify colonies with blue coloration indicative of polymer accumulation. Second, Nile Red A incorporated into MM (0.5 μg/mL) will allow selected isolates to be ranked based on UV fluorescence (312/365 nm).
This automated setup enables rapid testing of hundreds of isolate × carbon source combinations, accelerating the discovery of strains compatible with low-cost feedstocks and efficient bioprocessing while transforming a laborious manual process into a precise, scalable screening platform.
Here’s my draft script for this exercise.
Each “color” would correspond to a different bacterial isolate. I did not implement this in the script yet. The coordinate set is a starting layout and could be refined to achieve a more uniform, regular distribution across the plate (like in the image I drafted using the GUI available below)
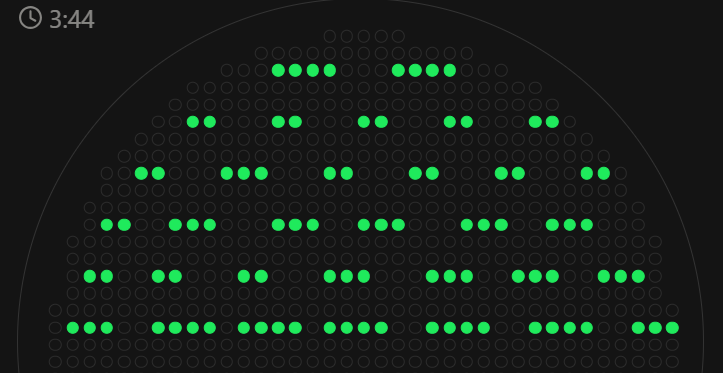

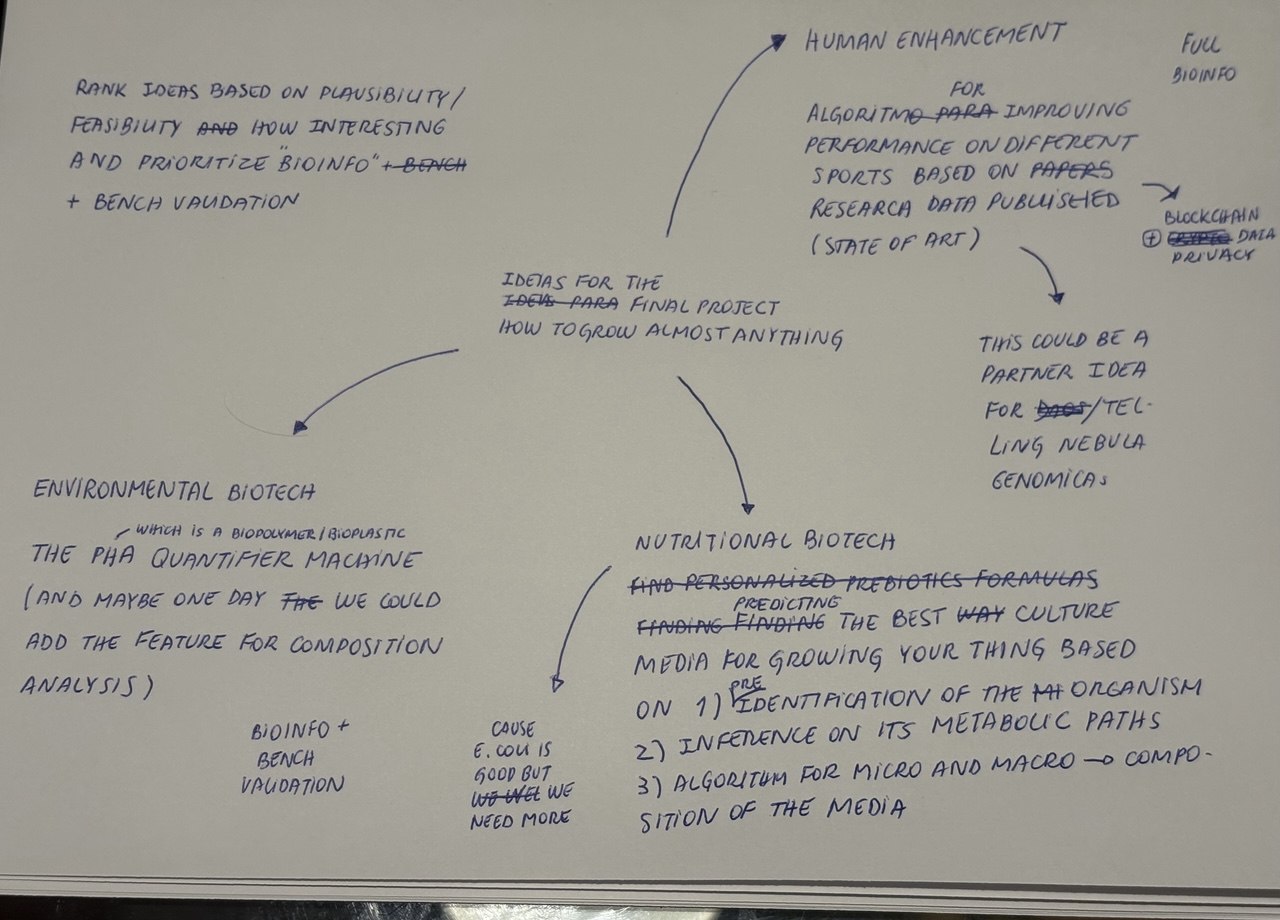
Final Project Ideas
Added 3 slides with 3 ideas for an Individual Final Project in the appropriate slide deck for Commited Listeners here.
Also here’s my analoginal brainstorm