Week 4 HW: Protein Design Part 1
Homework: Protein Design I
Part A. Conceptual Questions
1) How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons)
~ 21% of meat is protein content (Smith et al. 2022) therefore, 500g meet contains about 105g of protein.
Using the approximation of average amino acid ≈ 100 Da ≈ 100 g/mol for ~100 g protein: 100/100=1.00 mol
Avogadro’s number: 1 mole = 6.02214076×10²³ 1.00 mol × 6.022×10²³ ≈ 6.02×10²³ amino-acid molecules
2) Why do humans eat beef but do not become a cow, eat fish but do not become fish?
Beef/fish supplies raw materials and energy, but it doesn’t transfer “cow/fish identity”. What we eat is digested first meaning the proteins, fats, and carbohydrates are broken down into small building blocks (amino acids, fatty acids, sugars), absorbed, and then reassembled into human molecules under human genetic and hormonal control.
3) Why are there only 20 natural amino acids?
Doig (2017) hypothesizes that the canonical set of 20 standard amino acids is best understood as an evolved “alphabet” that became fixed early because this set is sufficient and practical for building stable, soluble proteins. This set enables soluble folded structures with close-packed hydrophobic cores and ordered binding pockets, rather than being selected because each amino acid was needed for catalysis (since RNA catalysts were already effective enough). Once early life standardized a working translation system around this set, changing the alphabet would have been costly, so it became effectively locked in (“frozen”). Other references, such as Freeland et al. (2000), suggest that 20 is a good number for minimizing damage from errors (mutation/mistranslation).
4) Where did amino acids come from before enzymes that make them, and before life started?
Amino acids could plausibly have come from abiotic chemistry on early Earth. Proposed routes include cyanosulfidic protometabolism and amino-acid formation from electrical discharges in simple “primitive Earth” gas mixtures (the classic Miller experiment).
5) Can you discover additional helices in proteins?
Beyond the α-helix, proteins commonly contain 3₁₀ helices and π helices (less frequent helical variants), as well as polyproline II helices (common in Pro-rich/disordered regions) and the specialized collagen triple helix.
6) Why are most molecular helices right-handed?
Right-handed helices dominate because natural biomolecules are made from single-handed monomers, and the right-handed twist is the lowest-energy way to repeat their geometry without clashes.
7) Why do β-sheets tend to aggregate?
β-sheet aggregation buries exposed hydrophobic side chains and releases ordered water from their surfaces, which is strongly favorable, lowering enthalpy.
8) What is the driving force for β-sheet aggregation?
β-sheet aggregation is driven mainly by the hydrophobic effect and stabilized/propagated by intermolecular backbone H-bonding in the cross-β structure (often reinforced by tight steric-zipper packing).
9) Why do many amyloid diseases form β-sheets?
β-sheet architecture is an unusually generic, stable, and self-templating way for polypeptide backbones to stick together when normal folding fails. In a β-sheet, the peptide backbone forms regular hydrogen bonds. This conformation makes amyloid fibrils thermodynamically stable and hard to clear, because once a small β-sheet nucleus forms, it can seed further growth by recruiting more monomers and templating the same β-rich structure.
Part B: Protein Analysis and Visualization
Question 1
I selected poly(3-hydroxyalkanoate) depolymerase (PhaZ) because it is the key enzyme that degrades PHB, which directly controls whether a microbe accumulates bioplastic (useful for biotechnology) or breaks it down (relevant for environmental fate). phaZ inactivation is commonly discussed as a strategy to reduce PHA mobilization and increase polymer retention.
Question 2
MPEPYIFRTVELDDQSIRTAVRPGKPHLTPLLIFNGIGANLELVFPFIEALDPDLEVIAFDVPGVGGSSTPRHPYRFPGLAKLTARMLDYLDYGQVSAIGVSWGGALAQQFAHDYPERCKKLVLAATAAGAVMVPGKPKVLWMMASPRRYVQPSHVIRIAPLIYGGAFRRDPDLAMHHASKVRSGGKLGYYWQLFAGLGWTSIHWLHKIHQPTLVLAGDDDPLIPLVNMRLLAWRIPNAQLHIIDDGHLFLITRAEAVAPIIMKFLQEERQRAVMHPRPASGG
BLAST Result Lenght: 283 aa Most frequent amino acid: Leucine (L), 32/283 = 11.3%
250 hits Reviewed (Swiss-Prot) homologs: 1
It belongs to the PHA depolymerase (PhaZ) family, which is part of the broader α/β-hydrolase enzyme superfamily.
Question 3
AF_AFP26495F1 - COMPUTED STRUCTURE MODEL OF POLY(3-HYDROXYALKANOATE) DEPOLYMERASE
This is not an experimentally solved structure, so there is no X-ray/EM “resolution” value. RCSB explicitly states: “There are no experimental data to verify the accuracy of this computed structure model. See Model Confidence metrics below for all regions of the polypeptide chain.” Instead, quality is reported by AlphaFold confidence. Global pLDDT: 91.95 (very high confidence overall)
RCSB lists 1 unique protein chain (monomer A1) and no ligands/non-protein entities.
Structure classification family: InterPro annotations classify it as Poly(3-hydroxyalkanoate) depolymerase (IPR011942) and an alpha/beta hydrolase fold protein (Alpha/beta hydrolase fold-1 domain, AB hydrolase superfamily).
Question 4
I opened AF-Q9R9W3-F1-model_v6 in PyMOL and visualized it in cartoon, ribbon, and ball-and-stick representations.
Colored by secondary structure, it shows a mixed α/β fold with more helices than β-sheets.
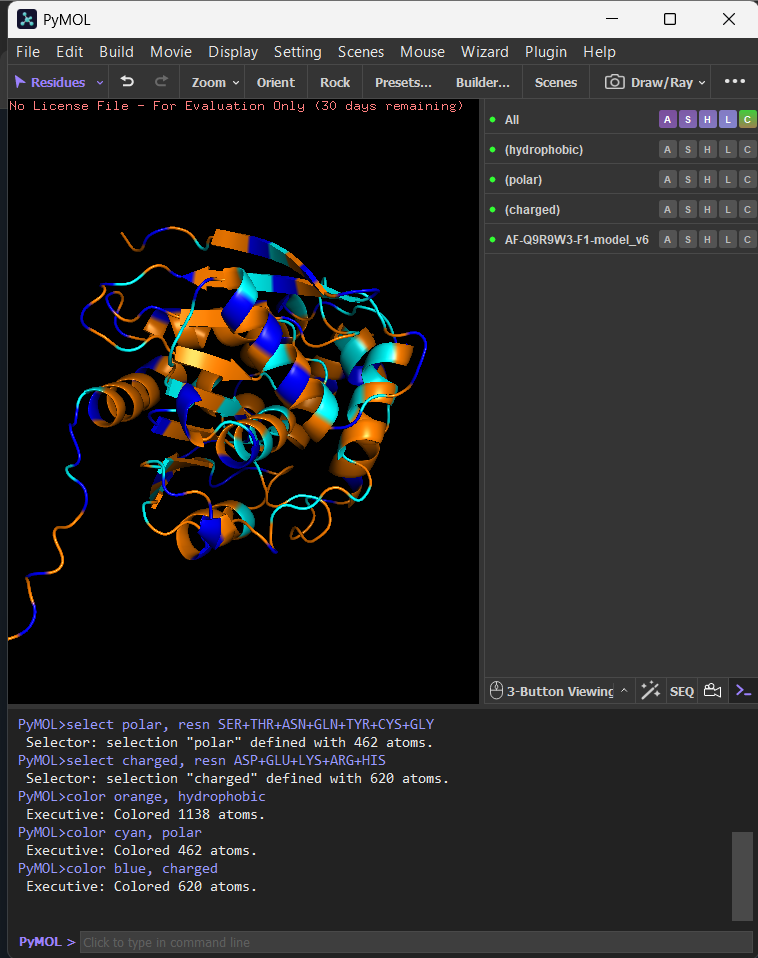
Colored by residue type, hydrophobic residues are enriched in the core (and in a few surface patches), while polar/charged residues are mostly surface-exposed, consistent with solubility.
The surface view shows clear cavities/clefts, consistent with potential binding pockets (e.g., a substrate-binding groove typical of hydrolases).
Part C. Using ML-Based Protein Design Tools
C1. Protein Language Modeling
Question 1
a)
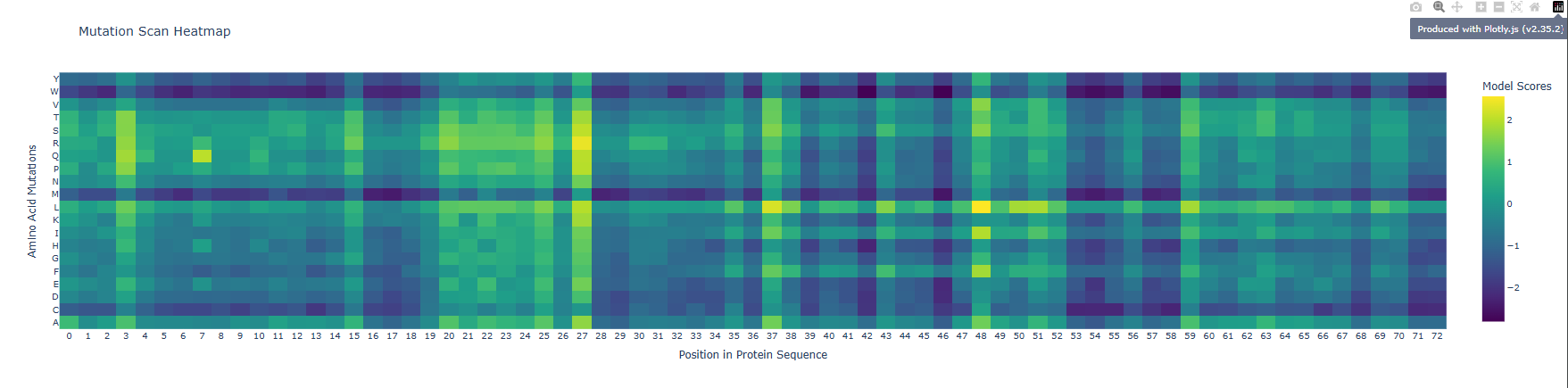
b) The vertical darker columns at certain positions are highly constrained residues where most substitutions are penalized. That usually indicates structural importance (core packing, tight turns, or residues critical for fold stability). Positions with mostly neutral colors across many substitutions are likely surface-exposed or in flexible loops, where the model predicts more tolerance
Question 2
Latent Space Analysis Use the provided sequence dataset to embed proteins in reduced dimensionality. Analyze the different formed neighborhoods: do they approximate similar proteins? Place your protein in the resulting map and explain its position and similarity to its neighbors.
In progress…
Part D. Group Brainstorm on Bacteriophage Engineering
GROUP MEMBERS
GROUP MEMBERS: Diogo Custodio; Flo Razoux; Katharine Kolin; Mariana Kanbe; Marisa Satsia.
PROJECT MAIN GOAL in discussion: Increased stability (easiest), higher titers (medium), higher toxicity of lysis protein (hard)
My group and I are conducting research for the group phage project. We have set up a shared Google Docs (screenshot below).
