Week 2: DNA Read, Write, & Edit
Homework
Part 1: Benchling & In-silico Gel Art
Opened https://benchling.com/ and signed up. Found the Lambda sequence from https://www.neb.com/en/-/media/nebus/page-images/tools-and-resources/interactive-tools/dna-sequences-and-maps/text-documents/lambdafsa.txt?rev=c0c6669b9bd340ddb674ebfd9d55c691&hash=B4188C171E5A42A1CF6FD257F98B97A1 and copied the sequence (without the header). Pasted this sequence into Benchling through “Create” > “DNA / RNA Sequence” > “New DNA / RNA Sequence”. Then I just pasted the sequence in the “Bases” field, titled it “Lambda,” and selected the topology as “Linear.”
Clicked “Digest” (the scissors icon in the right menu), selected “All enzymes,” found all seven using the search tool, and clicked “Run Digest.”

Part 3: DNA Design Challenge
3.1. Choose your protein: Poly(3-hydroxyalkanoate) polymerase subunit PhaC
I chose Polyhydroxyalkanoate synthase (PhaC) because it is involved in the catalysis of the reaction that polymerizes (R)-3-hydroxybutyryl-CoA to produce polyhydroxybutyrate (PHB), which is an important bioproduct of interest due to its plastic/polyethylene-like properties.
Biologically, PHB serves as an intracellular energy reserve material when cells grow under conditions of nutrient limitation.
Sequence of Polyhydroxyalkanoate Synthase (PhaC): MATGKGAAASTQEGKSQPFKVTPGPFDPATWLEWSRQWQGTEGNGHAAASGIPGLDALAGVKIAPAQLGDIQQRYMKDFSALWQAMAEGKAEATGPLHDRRFAGDAWRTNLPYRFAAAFYLLNARALTELADAVEADAKTRQRIRFAISQWVDAMSPANFLATNPEAQRLLIESGGESLRAGVRNMMEDLTRGKISQTDESAFEVGRNVAVTEGAVVFENEYFQLLQYKPLTDKVHARPLLMVPPCINKYYILDLQPESSLVRHVVEQGHTVFLVSWRNPDASMAGSTWDDYIEHAAIRAIEVARDISGQDKINVLGFCVGGTIVSTALAVLAARGEHPAASVTLLTTLLDFADTGILDVFVDEGHVQLREATLGGGAGAPCALLRGLELANTFSFLRPNDLVWNYVVDNYLKGNTPVPFDLLFWNGDATNLPGPWYCWYLRHTYLQNELKVPGKLTVCGVPVDLASIDVPTYIYGSREDHIVPWTAAYASTALLANKLRFVLGASGHIAGVINPPAKNKRSHWTNDALPESPQQWLAGAIEHHGSWWPDWTAWLAGQAGAKRAAPANYGNARYRAIEPAPGRYVKAKA Source: UniProt at https://www.uniprot.org/uniprotkb/P23608/entry#sequences
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence. reh:H16_A1437 K03821 poly(R)-3-hydroxyalkanoate polymerase subunit PhaC EC:2.3.1.304 | (GenBank) phaC1; Poly(3-hydroxybutyrate) polymerase (N) atggcgaccggcaaaggcgcggcagcttccacgcaggaaggcaagtcccaaccattcaaggtcacgccggggccattcgatccagccacatggctggaatggtcccgccagtggcagggcactgaaggcaacggccacgcggccgcgtccggcattccgggcctggatgcgctggcaggcgtcaagatcgcgccggcgcagctgggtgatatccagcagcgctacatgaaggacttctcagcgctgtggcaggccatggccgagggcaaggccgaggccaccggtccgctgcacgaccggcgcttcgccggcgacgcatggcgcaccaacctcccatatcgcttcgctgccgcgttctacctgctcaatgcgcgcgccttgaccgagctggccgatgccgtcgaggccgatgccaagacccgccagcgcatccgcttcgcgatctcgcaatgggtcgatgcgatgtcgcccgccaacttccttgccaccaatcccgaggcgcagcgcctgctgatcgagtcgggcggcgaatcgctgcgtgccggcgtgcgcaacatgatggaagacctgacacgcggcaagatctcgcagaccgacgagagcgcgtttgaggtcggccgcaatgtcgcggtgaccgaaggcgccgtggtcttcgagaacgagtacttccagctgttgcagtacaagccgctgaccgacaaggtgcacgcgcgcccgctgctgatggtgccgccgtgcatcaacaagtactacatcctggacctgcagccggagagctcgctggtgcgccatgtggtggagcagggacatacggtgtttctggtgtcgtggcgcaatccggacgccagcatggccggcagcacctgggacgactacatcgagcacgcggccatccgcgccatcgaagtcgcgcgcgacatcagcggccaggacaagatcaacgtgctcggcttctgcgtgggcggcaccattgtctcgaccgcgctggcggtgctggccgcgcgcggcgagcacccggccgccagcgtcacgctgctgaccacgctgctggactttgccgacacgggcatcctcgacgtctttgtcgacgagggccatgtgcagttgcgcgaggccacgctgggcggcggcgccggcgcgccgtgcgcgctgctgcgcggccttgagctggccaataccttctcgttcttgcgcccgaacgacctggtgtggaactacgtggtcgacaactacctgaagggcaacacgccggtgccgttcgacctgctgttctggaacggcgacgccaccaacctgccggggccgtggtactgctggtacctgcgccacacctacctgcagaacgagctcaaggtaccgggcaagctgaccgtgtgcggcgtgccggtggacctggccagcatcgacgtgccgacctatatctacggctcgcgcgaagaccatatcgtgccgtggaccgcggcctatgcctcgaccgcgctgctggcgaacaagctgcgcttcgtgctgggtgcgtcgggccatatcgccggtgtgatcaacccgccggccaagaacaagcgcagccactggactaacgatgcgctgccggagtcgccgcagcaatggctggccggcgccatcgagcatcacggcagctggtggccggactggaccgcatggctggccgggcaggccggcgcgaaacgcgccgcgcccgccaactatggcaatgcgcgctatcgcgcaatcgaacccgcgcctgggcgatacgtcaaagccaaggcatga Source: KEGG at https://www.genome.jp/dbget-bin/www_bget?reh:H16_A1437
3.3. Codon optimization. I optimized the phaC coding sequence for E. coli because it is a widely used chassis for recombinant protein expression and for rapid prototyping of metabolic engineering constructs.
I did this using the Benchling tool. I’ve selected the region of the AA sequence I wish to back translate and right clicked on the highlighted region. From the the codon optimization tab:
- Host: E. coli K-12
- Method: Match codon usage
- GC content: Medium (0.33 to 0.66) cause the extremes may be inconvenient. High GC can create strong secondary structures and low GC can cause instability/repeats and can make synthesis harder.
- Uridine depletion: off (not relevant for bacterial expression)
- Hairpin parameters: Stem size: 8 and Window 50
- Restriction sites: avoid BsaI, BsmBI, BbsI (Type IIS enzymes for Golden Gate compatibility since I would have to clone phaA and phaB also, not phaC single gene in one vector)
- Patterns to reduce: AAAAAA and ATATATATA
I clicked on “Optimization preview” and got this result:
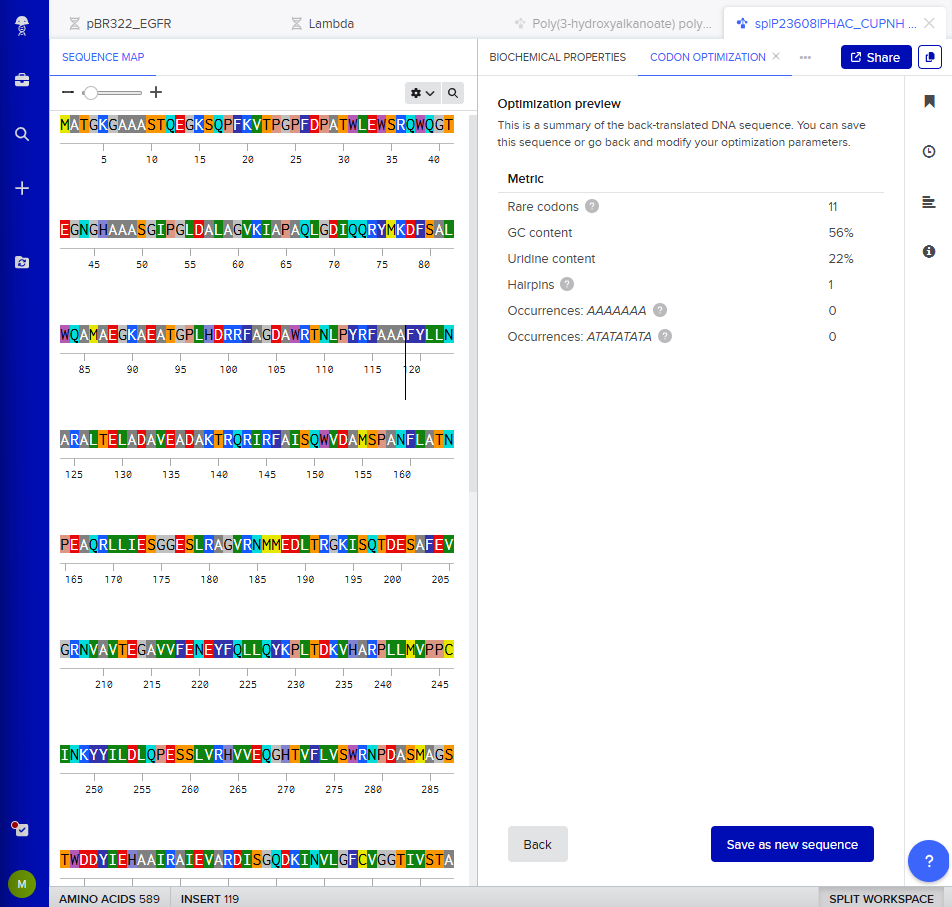
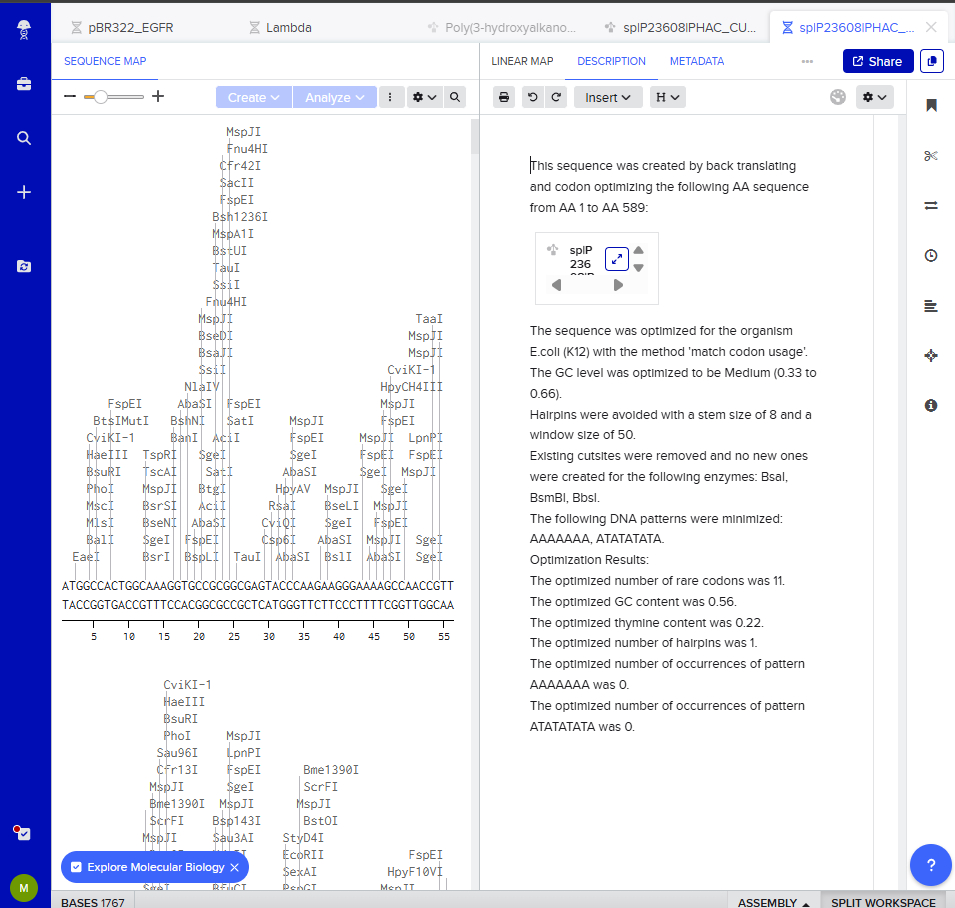
3.4. You have a sequence! Now what?
PhaC alone will not produce PHB. A minimal PHB pathway typically includes PhaA (β-ketothiolase) and PhaB (acetoacetyl-CoA reductase) in addition to PhaC (PHA synthase). PhaA and PhaB convert central metabolites (via acetyl-CoA) into (R)-3-hydroxybutyryl-CoA, which is the direct substrate that PhaC polymerizes into PHB. You will also need a host capable of supplying sufficient acetyl-CoA and NADPH.
Therefore, for PHB production in E. coli, phaA, phaB, and phaC are commonly co-expressed on the same plasmid (as a single operon with one promoter and RBSs for each gene) and grown under appropriate culture conditions (e.g., carbon excess and nutrient limitation) that favor polymer accumulation.
Part 4: Prepare a Twist DNA Synthesis Order
Project: pBBR1-MSC5::phaCAB Cell-dependent recombinant expression approach: cloning the codon-optimized phaA, phaB and phaC coding sequences into E. coli K12
Promoter - RBS - phaA - (RBS) - phaB - (RBS) - phaC - Terminator
phaA Sequence MTDVVIVSAARTAVGKFGGSLAKIPAPELGAVVIKAALERAGVKPEQVSEVIMGQVLTAGSGQNPARQAAIKAGLPAMVPAMTINKVCGSGLKAVMLAANAIMAGDAEIVVAGGQENMSAAPHVLPGSRDGFRMGDAKLVDTMIVDGLWDVYNQYHMGITAENVAKEYGITREAQDEFAVGSQNKAEAAQKAGKFDEEIVPVLIPQRKGDPVAFKTDEFVRQGATLDSMSGLKPAFDKAGTVTAANASGLNDGAAAVVVMSAAKAKELGLTPLATIKSYANAGVDPKVMGMGPVPASKRALSRAEWTPQDLDLMEINEAFAAQALAVHQQMGWDTSKVNVNGGAIAIGHPIGASGCRILVTLLHEMKRRDAKKGLASLCIGGGMGVALAVERK Source: UniProt at https://www.uniprot.org/uniprotkb/P14611/entry#sequences
phaB Sequence MTQRIAYVTGGMGGIGTAICQRLAKDGFRVVAGCGPNSPRREKWLEQQKALGFDFIASEGNVADWDSTKTAFDKVKSEVGEVDVLINNAGITRDVVFRKMTRADWDAVIDTNLTSLFNVTKQVIDGMADRGWGRIVNISSVNGQKGQFGQTNYSTAKAGLHGFTMALAQEVATKGVTVNTVSPGYIATDMVKAIRQDVLDKIVATIPVKRLGLPEEIASICAWLSSEESGFSTGADFSLNGGLHMG Source: UniProt at https://www.uniprot.org/uniprotkb/P14697/entry#sequences
phaC Sequence MATGKGAAASTQEGKSQPFKVTPGPFDPATWLEWSRQWQGTEGNGHAAASGIPGLDALAGVKIAPAQLGDIQQRYMKDFSALWQAMAEGKAEATGPLHDRRFAGDAWRTNLPYRFAAAFYLLNARALTELADAVEADAKTRQRIRFAISQWVDAMSPANFLATNPEAQRLLIESGGESLRAGVRNMMEDLTRGKISQTDESAFEVGRNVAVTEGAVVFENEYFQLLQYKPLTDKVHARPLLMVPPCINKYYILDLQPESSLVRHVVEQGHTVFLVSWRNPDASMAGSTWDDYIEHAAIRAIEVARDISGQDKINVLGFCVGGTIVSTALAVLAARGEHPAASVTLLTTLLDFADTGILDVFVDEGHVQLREATLGGGAGAPCALLRGLELANTFSFLRPNDLVWNYVVDNYLKGNTPVPFDLLFWNGDATNLPGPWYCWYLRHTYLQNELKVPGKLTVCGVPVDLASIDVPTYIYGSREDHIVPWTAAYASTALLANKLRFVLGASGHIAGVINPPAKNKRSHWTNDALPESPQQWLAGAIEHHGSWWPDWTAWLAGQAGAKRAAPANYGNARYRAIEPAPGRYVKAKA Source: UniProt at https://www.uniprot.org/uniprotkb/P23608/entry#sequences
For this exercise, I chose pBBR1MCS-5 as the plasmid backbone because it is a broad-host-range vector commonly used for cloning and expression of phaCAB. Source: https://www.teses.usp.br/teses/disponiveis/87/87131/tde-29042010-102817/publico/RogeriodeSousaGomes_Doutorado.pdf
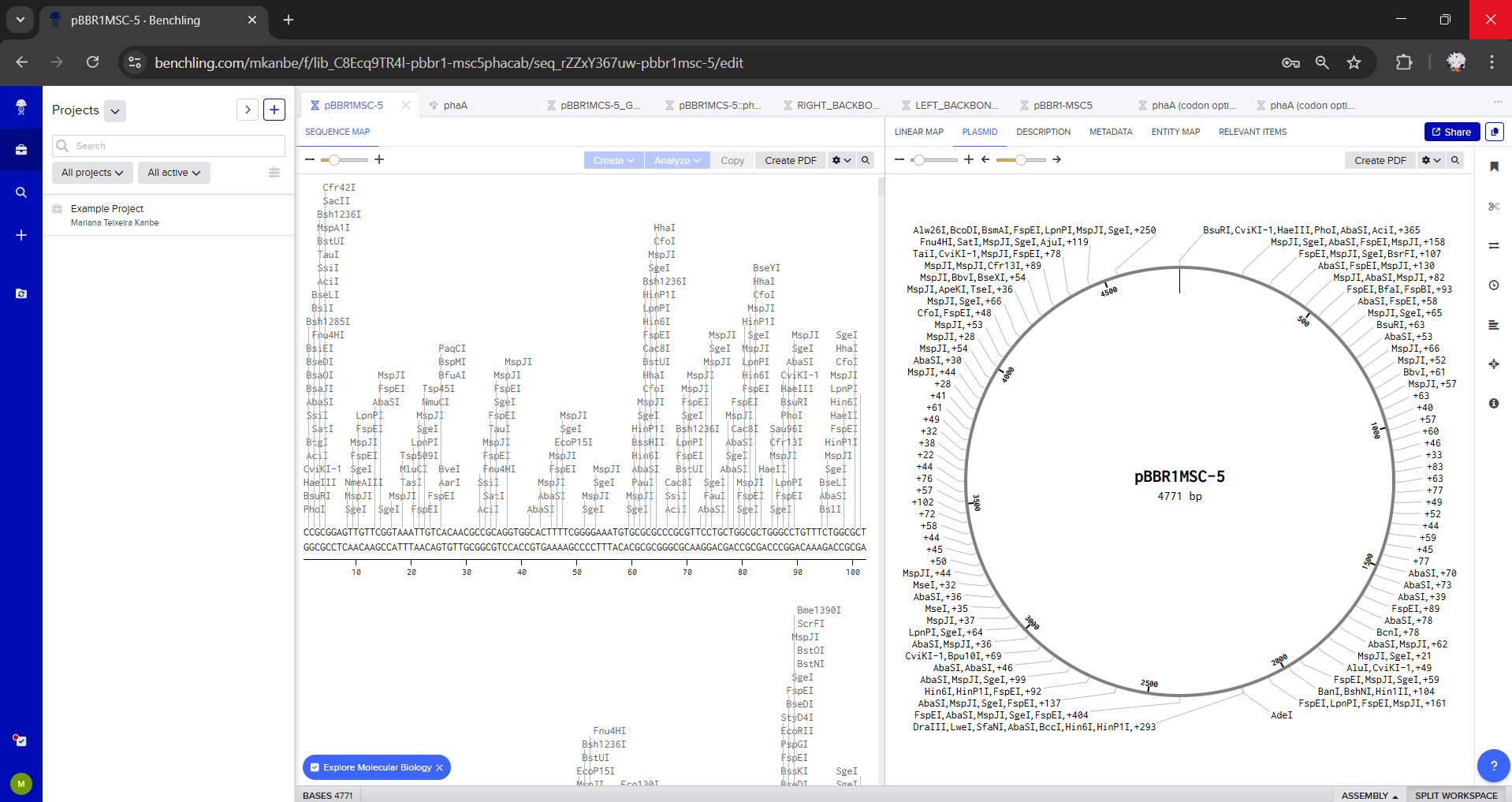
Part 5: DNA Read / Write / Edit
I would sequence DNA used for DNA-based digital data storage, because I’ve never did this before and would feel amazing to be able to instantly interpret the info like reading a book or something like this.
Maybe I’d use Illumina (second-generation, massively parallel short reads) sequencing for high-accuracy base calls and reliable decoding of short oligos and Nanopore (third-generation, single-molecule long reads)to validate longer constructs and integrity.
My input for using the Illumina method would be a DNA pool. This would have to go for a fragmentation stage, adapter ligation (indexes), and PCR amplicication). Throgh Illumina bases are decoded sequencing-by-synthesis with fluorescently labeled reversible terminators and the output is millions to billions of short reads (FASTQ) plus per-base quality scores. To decode that data it is required alignment/consensus and error correction.
I would synthesize a PHA production cassette for E. coli K12 (codon-optimized phaA + phaB + phaC) to enable rapid testing/studing of PHB production. I would use commercial gene synthesis (e.g., Twist) because it is practical, accurate. Essential steps would include oligo synthesis, oligo pooling, assembly into full-length gene/insert, cloning into plasmid. Among the limitations I’d face with this method is error compound since the probability increases with length. So long constructs often require assembly and clonal verification, adding time.
Aiming for increased expression of phaCAB and production of PHA I would edit E. coli metabolic and stress-tolerance genes to increase PHB yield, for example by improving acetyl-CoA/NADPH supply, reducing competing pathways, and increasing tolerance to intracellular polymer accumulation (reducing lysis under high load).
I would use CRISPR-based editing for targeted point mutations without double-strand breaks. RNA is guided direct Cas9 to a locus, DNA is cut and repaired via HDR using a donor template containing the desired edit. In the end I would confirm edits by sequencing. Among the limitations I’d say imprecision (off-target edits) and the fact that multiplex edits increase complexity and screening effort.