Week 5 HW: Protein Design Part 2
Part A: SOD1 Binder Peptide Design (From Pranam)
Part 1: Generate Binders with PepMLM
Question 1
This is human SOD1 sequence from UniProt (P00441) removing the initial Met
ATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ FASTA
introducing the A4V mutant associated with the most aggressive forms of the ALS disease ATKVVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
Question 2 and 3
With the help of ChatGPT and Gemni, I generated 2 new cells ir order to generate four peptides of length 12 amino acids conditioned on the mutant SOD1 sequence.
4 PepMLM-generated 12-aa peptides (conditioned on mutant SOD1):
- HRVPVAGVEWWE
- WSYYVTAVAHKE
- WRYGAAAVEWKE
- WSVPVVAIEHGE
Question 4
- HRVPVAGVEWWE
- WSYYVTAVAHKE
- WRYGAAAVEWKE
- WSVPVVAIEHGE
5. FLYRWLPSRRGG
Question 5
WRYGAAAVEWKE - ppl 4.645 (mean NLL 1.536) WSYYVTAVAHKE - ppl 5.094 (mean NLL 1.628) WSVPVVAIEHGE - ppl 6.423 (mean NLL 1.860) HRVPVAGVEWWE - ppl 7.660 (mean NLL 2.036) Known binder: FLYRWLPSRRGG - ppl 21.391 (mean NLL 3.063)
Interpretation: The perplexity score is PepMLM’s confidence in the peptide under its generative model. PepMLM perplexity can be interpreted this way: lower = higher confidence
PepMLM assigns higher confidence to the four generated peptides than to the known binder under this scoring scheme, with WRYGAAAVEWKE ranked best (lowest perplexity).
The known binder has higher perplexity, suggesting it is less consistent with PepMLM’s learned binder distribution for this target, even though it is experimentally reported to bind. This highlights that PepMLM perplexity is not an experimental binding score. Also, it suggests that perplexity alone is insufficient to validate binding.
As I found this really strange, I decided to find out checks I could run to see whether this was an error/artifact:
Test for missing mask token: negative, so all good.
Conclusion My generated peptides are enriched in W/V/A/Y and look like classic short hydrophobic binders. The known binder FLYRWLPSRRGG has a highly charged tail (RRGG) and a different composition pattern, which the model may assign low probability to even if it binds in reality.
Part 2: Evaluate Binders with AlphaFold3
Evaluate Binders with AlphaFold3
SOD1 Mutant Sequence (A4V mutation) ATKVVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
At first, I mistakenly evaluated all peptides in the same run.
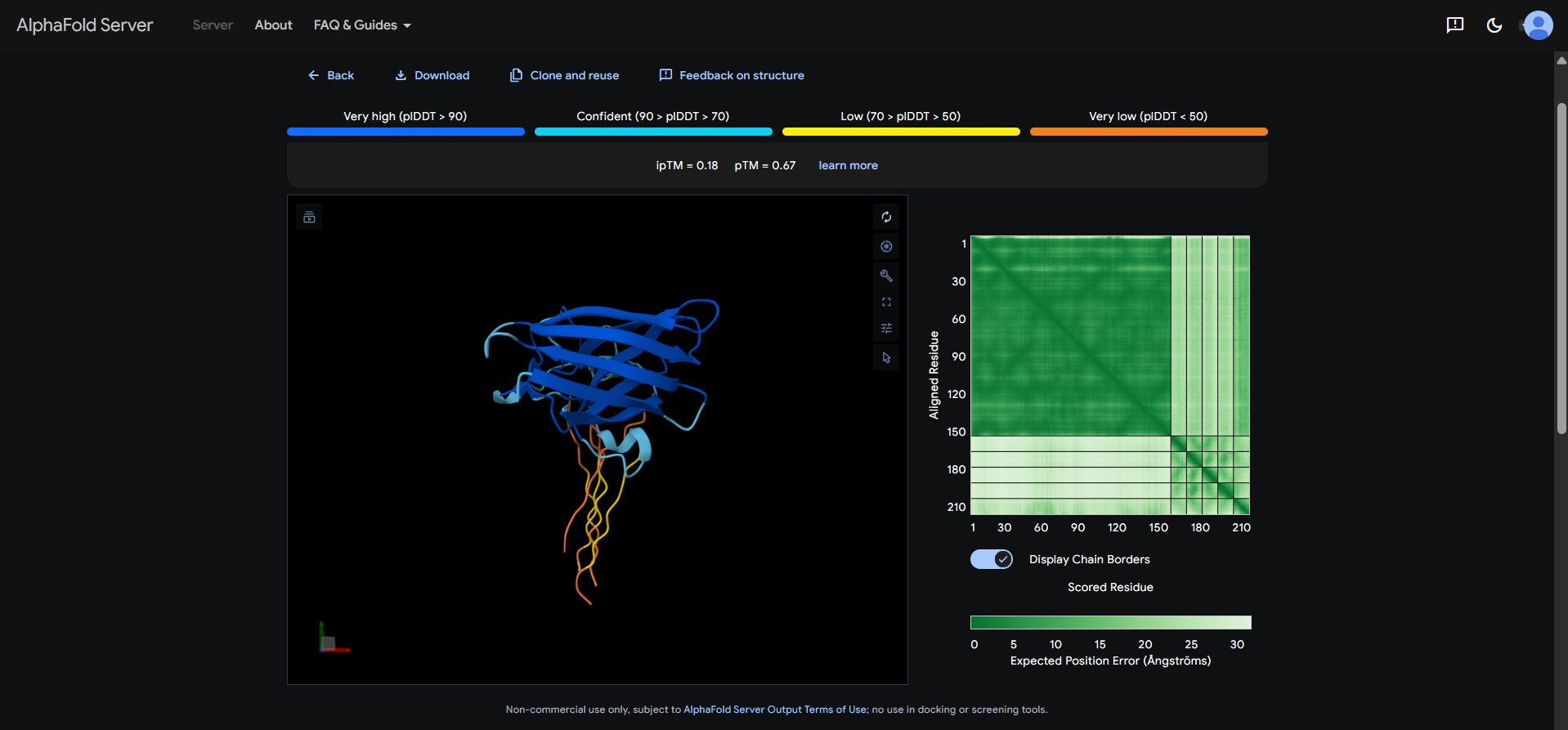
Then I noticed the AlphaFold Server treated that as one multi-chain complex with 6 chains total (SOD1 + 4 generated peptides + the known binder). So to compare them I would had to run 5 separate jobs.
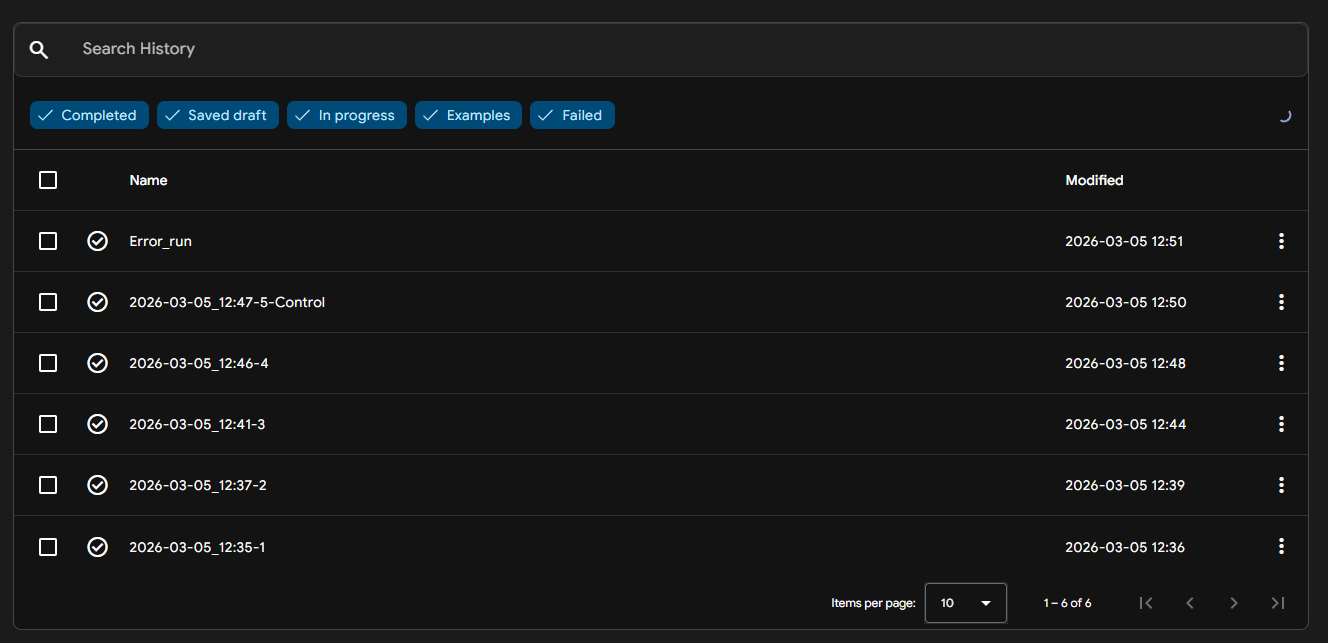
- SOD1 + HRVPVAGVEWWE: ipTM = 0.34; pTM = 0.86
Where the peptide appears to bind? The peptide is positioned along an external surface of the SOD1 β-strand core, contacting a β-sheet edge/adjacent loop (surface-bound).
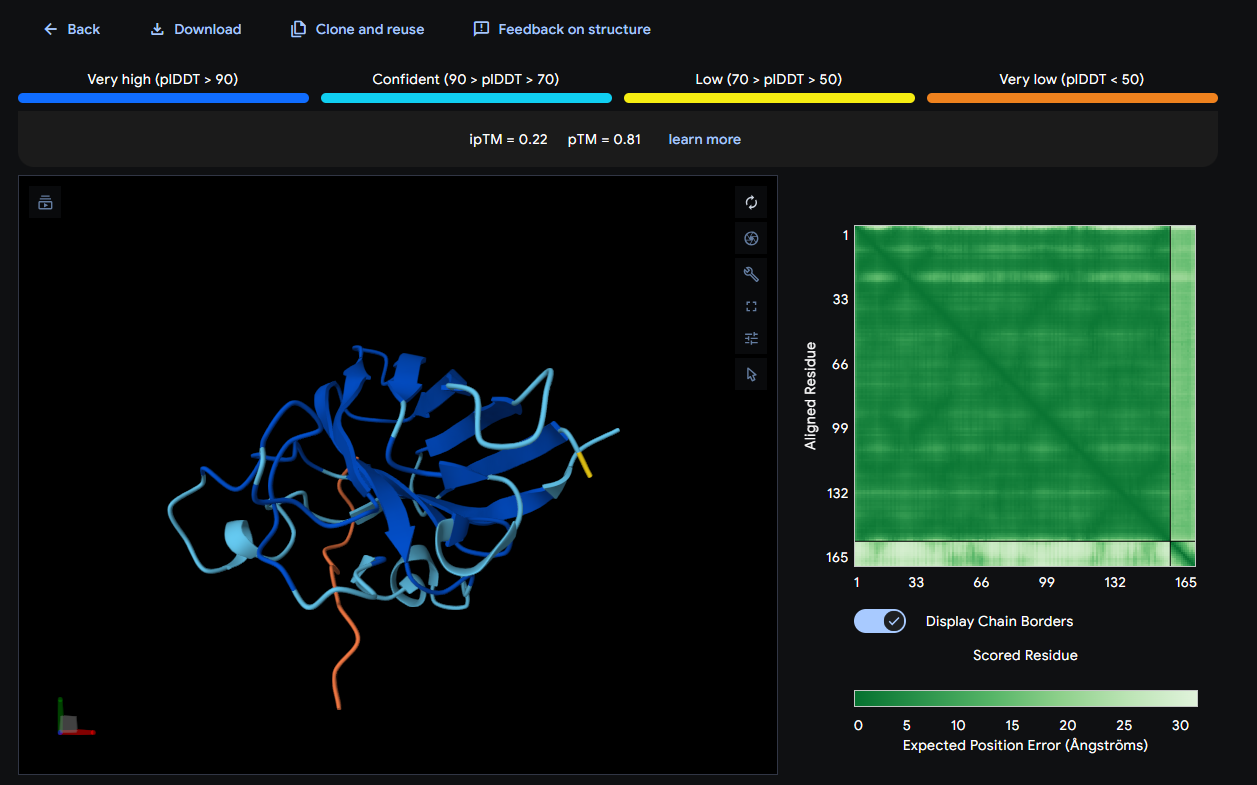
- SOD1 + WSYYVTAVAHKE: ipTM = 0.22; pTM = 0.81
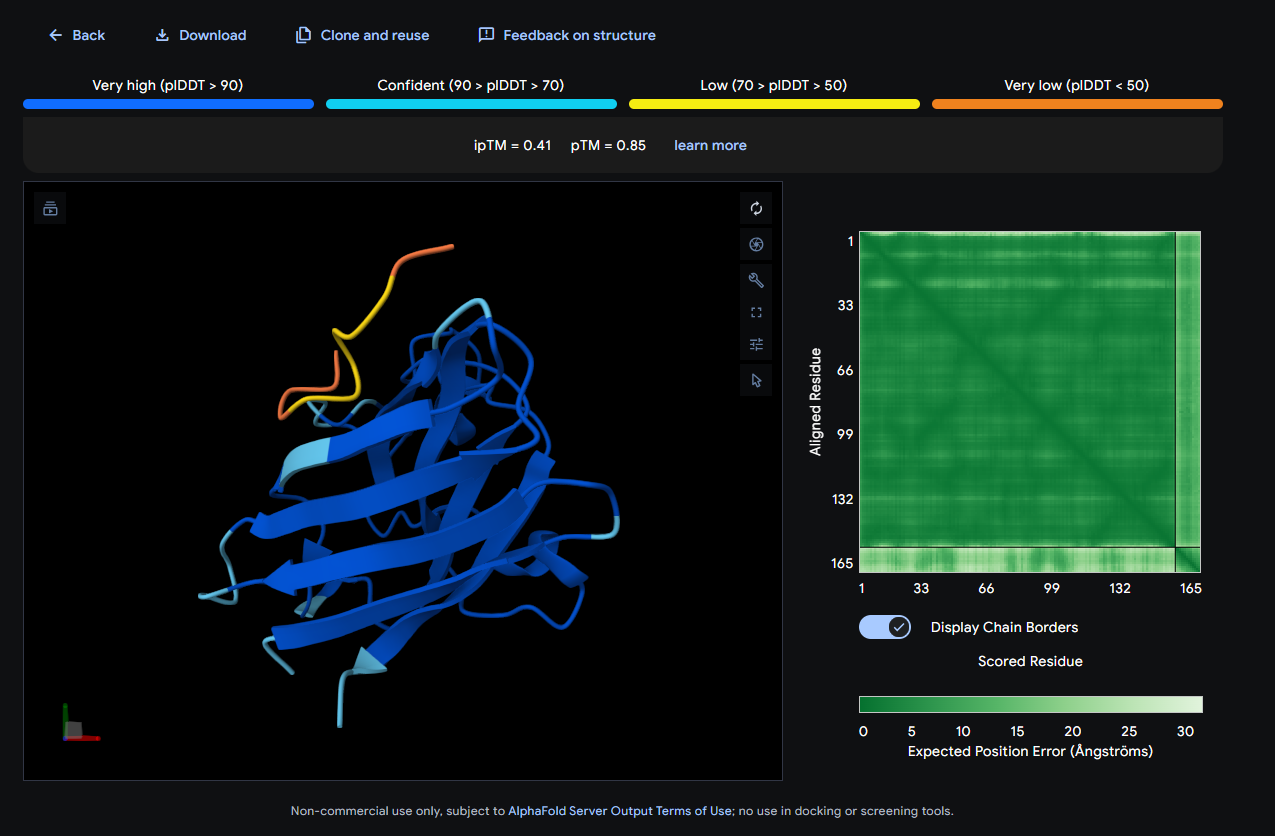
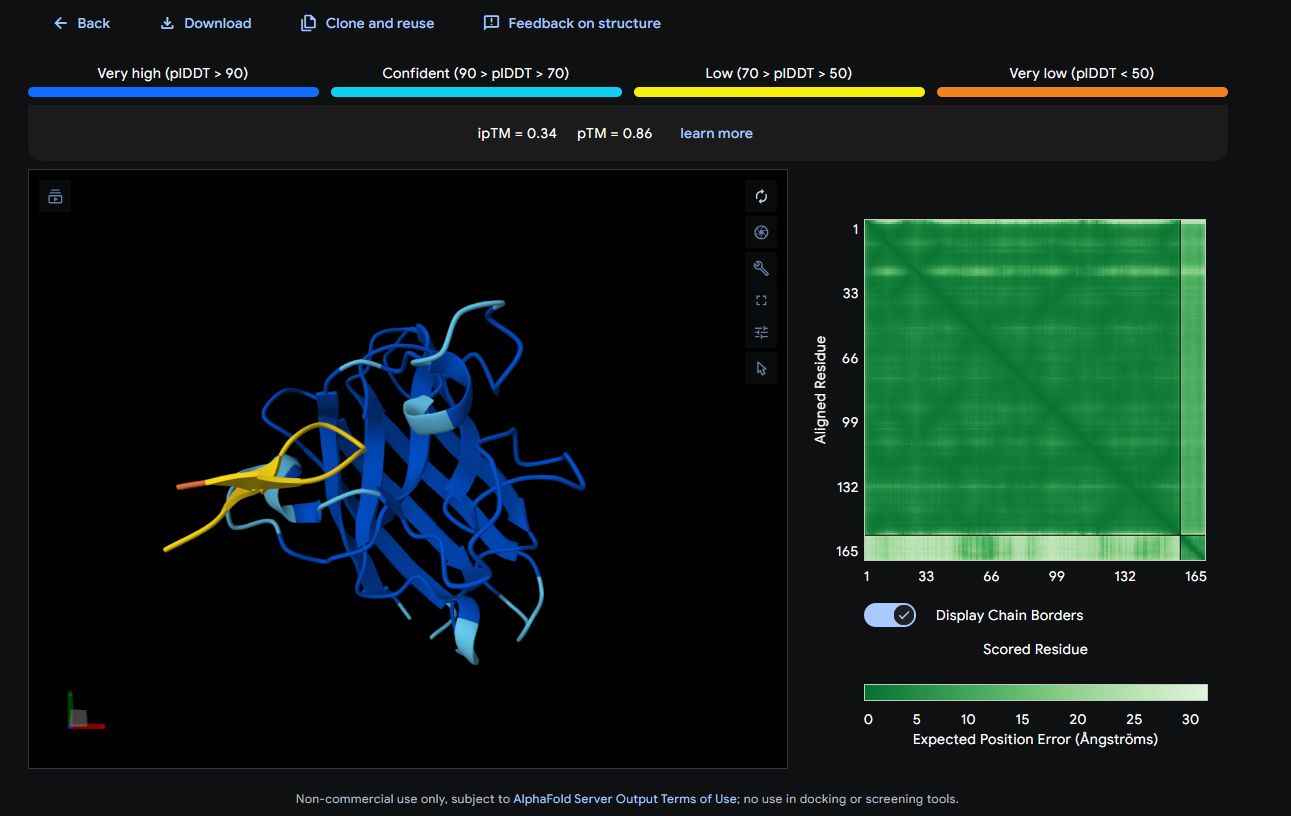
Where the peptide appears to bind? The peptide shows weak localization and appears loosely associated with the protein surface, without a clearly defined contact region.
- SOD1 + WRYGAAAVEWKE: ipTM = 0.41; pTM = 0.85
Where the peptide appears to bind? The peptide is placed near a β-barrel edge/loop region on the outer surface of SOD1 (surface-bound).
- SOD1 + WSVPVVAIEHGE: ipTM = 0.44; pTM = 0.86
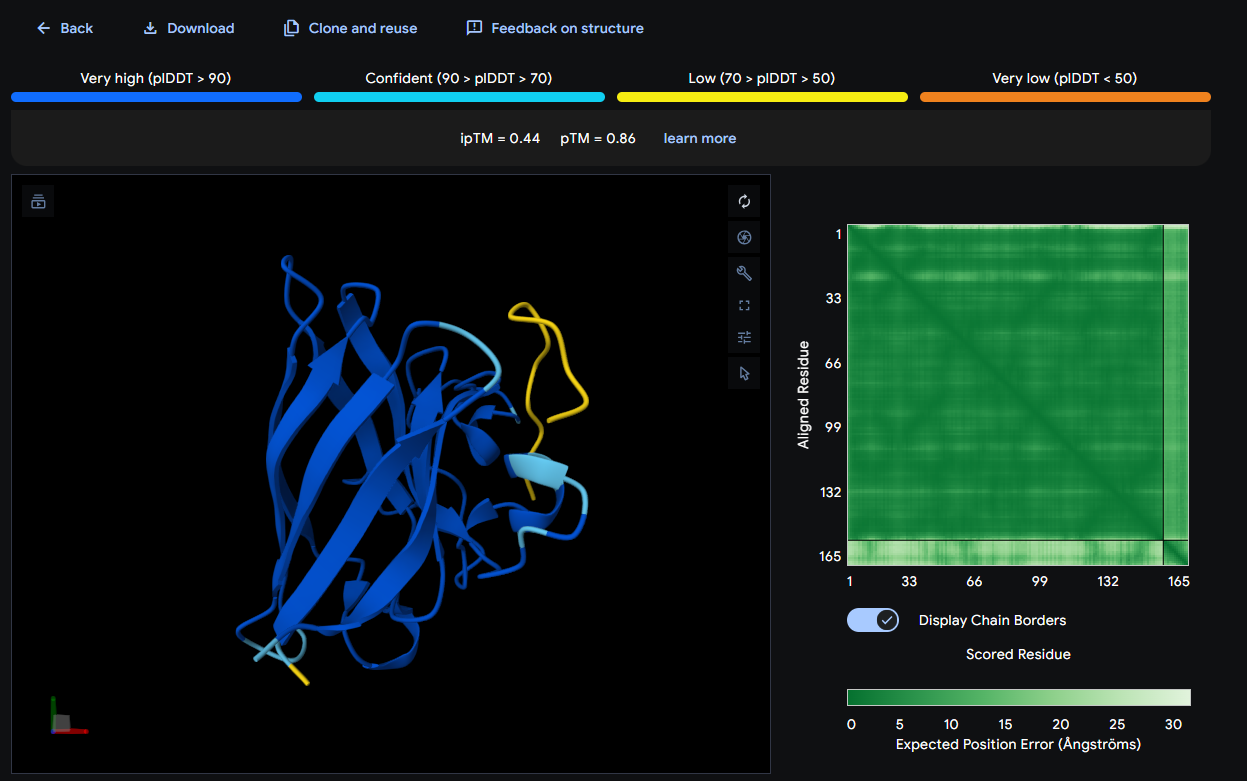
Where the peptide appears to bind? The peptide is positioned on a distinct surface patch on the β-barrel face/edge, appearing more localized than the others (surface-bound).
- SOD1 + FLYRWLPSRRGG (control): ipTM = 0.3; pTM = 0.83
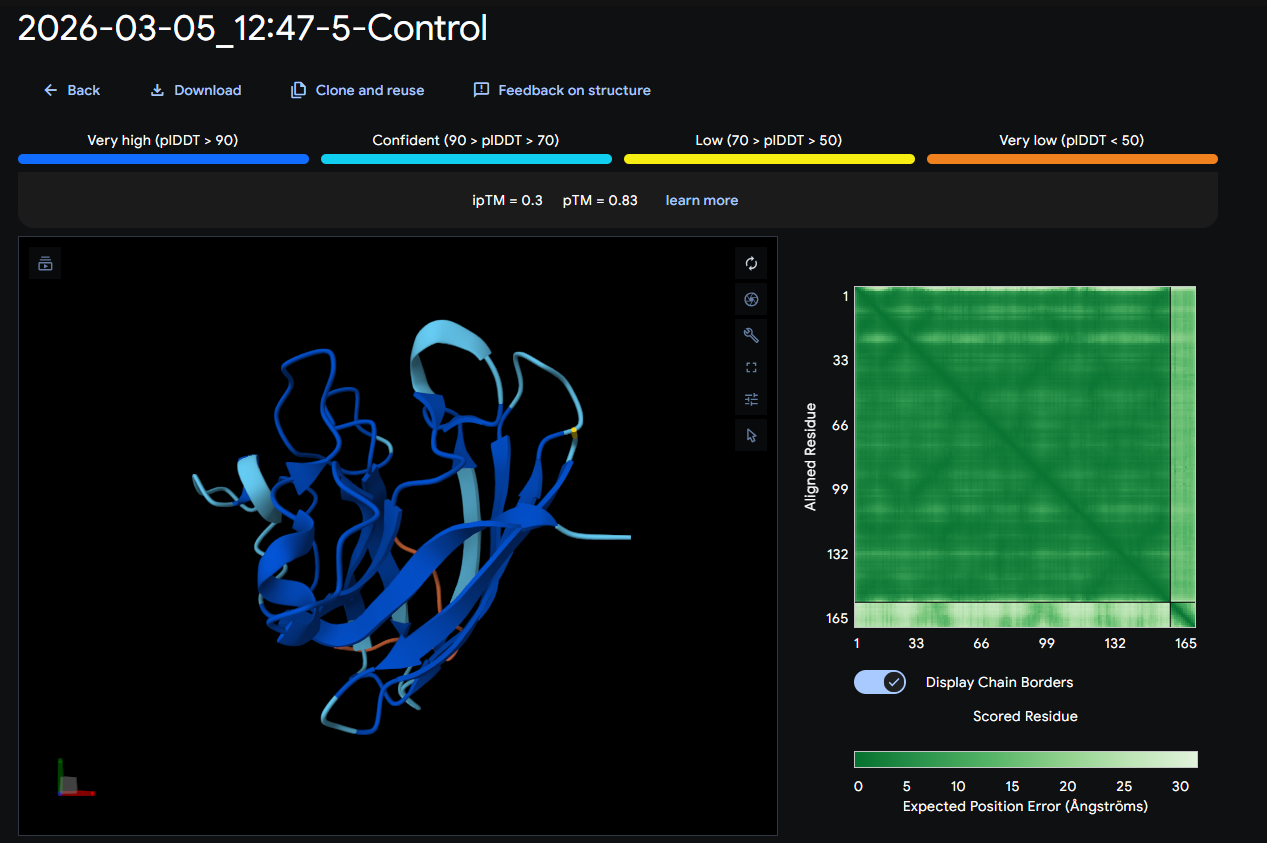
Where the peptide appears to bind? The peptide contacts the protein surface and appears partially inserted into a shallow surface groove/cleft (partially buried relative to the others).
By ipTM ranking: WSVPVVAIEHGE (0.44) > WRYGAAAVEWKE (0.41) > HRVPVAGVEWWE (0.34) > FLYRWLPSRRGG (0.30) > WSYYVTAVAHKE (0.22).
The observed ipTM values are uniformly low (0.22–0.44), indicating limited AlphaFold3 confidence in any specific peptide–SOD1 interface. Among the PepMLM-generated candidates, WSVPVVAIEHGE (ipTM = 0.44) and WRYGAAAVEWKE (ipTM = 0.41) score higher than the known binder FLYRWLPSRRGG (ipTM = 0.30), while HRVPVAGVEWWE (0.34) is slightly higher and WSYYVTAVAHKE (0.22) is lower. Overall, PepMLM-generated peptides match or exceed the known binder by ipTM, but the absolute scores suggest weakly supported, mostly surface-associated binding modes rather than a high-confidence complex.
Part 3: Evaluate Properties of Generated Peptides in the PeptiVerse
- HRVPVAGVEWWE
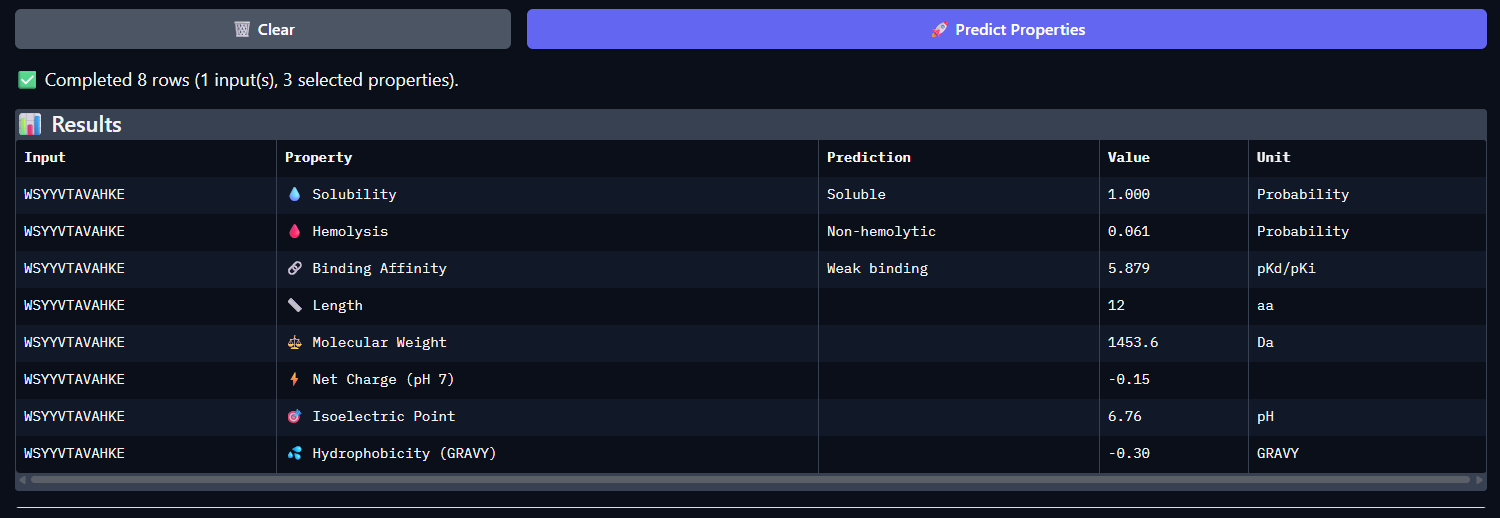
- WSYYVTAVAHKE
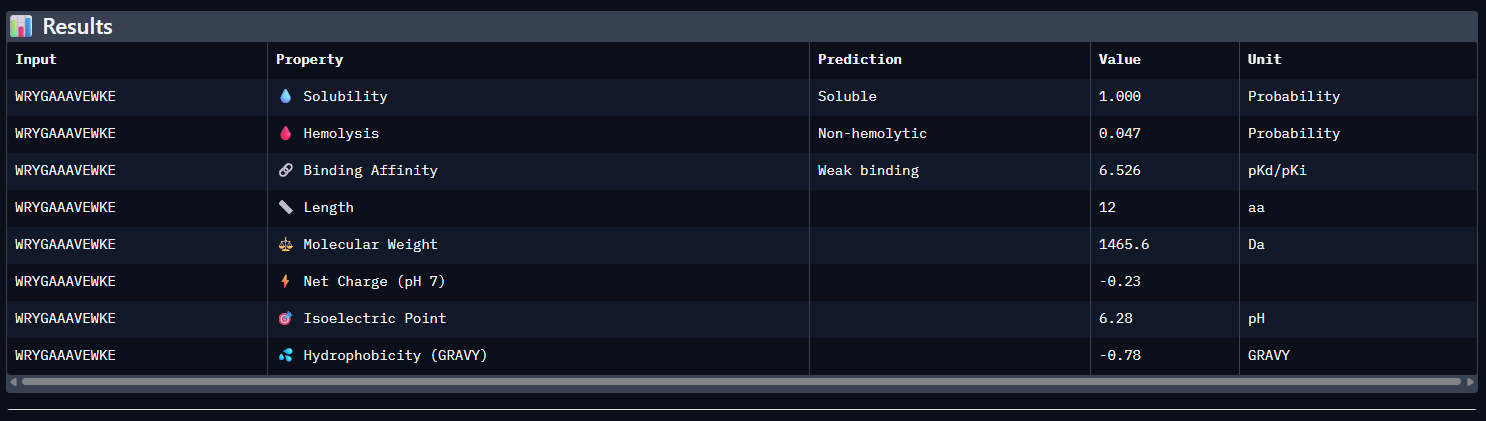
- WRYGAAAVEWKE
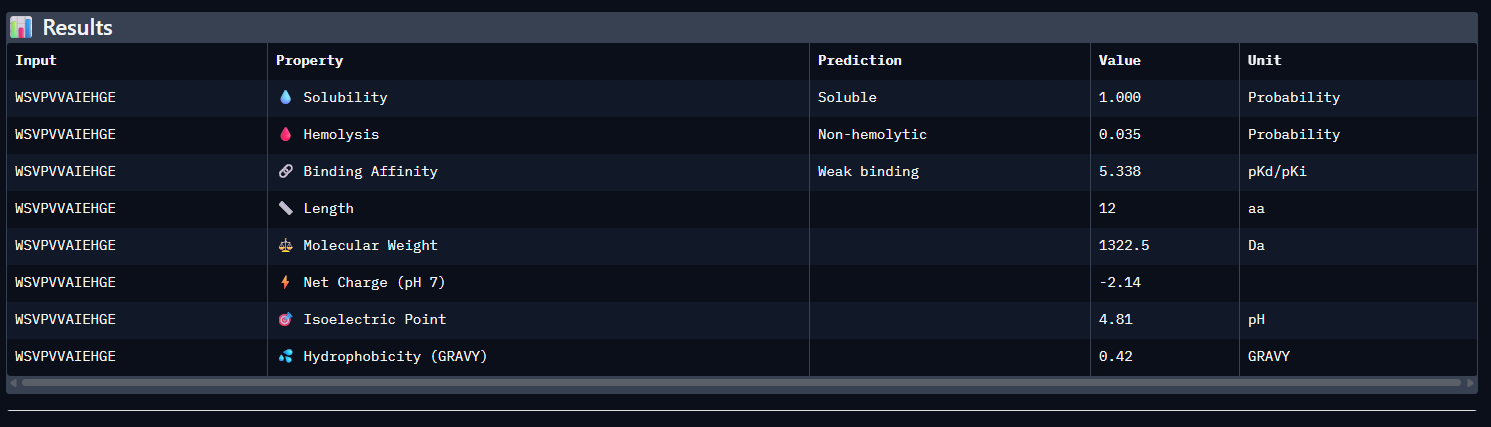
- WSVPVVAIEHGE
- FLYRWLPSRRGG (control)
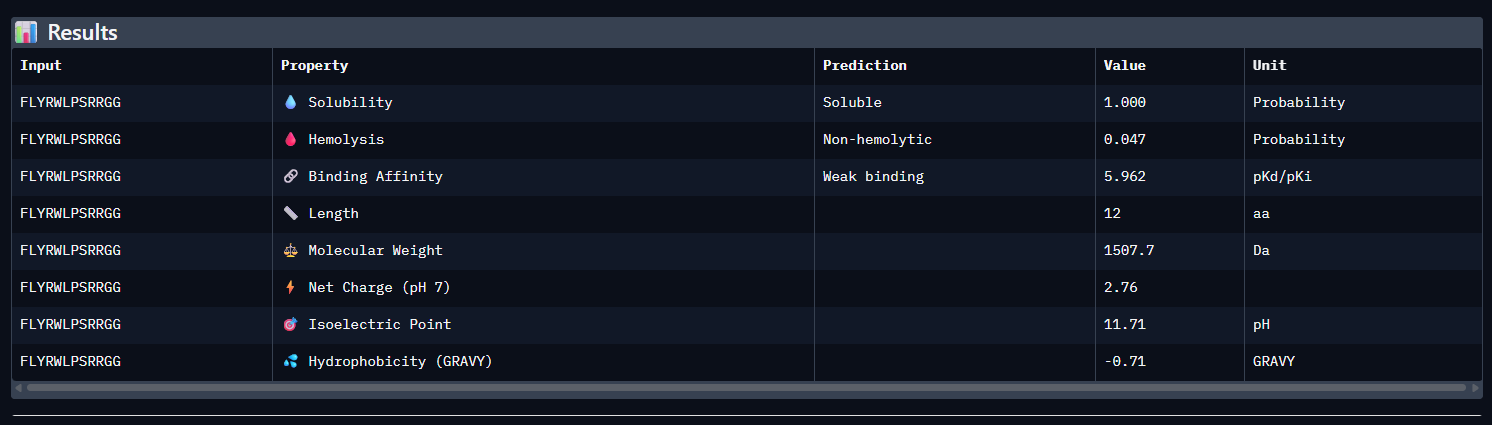
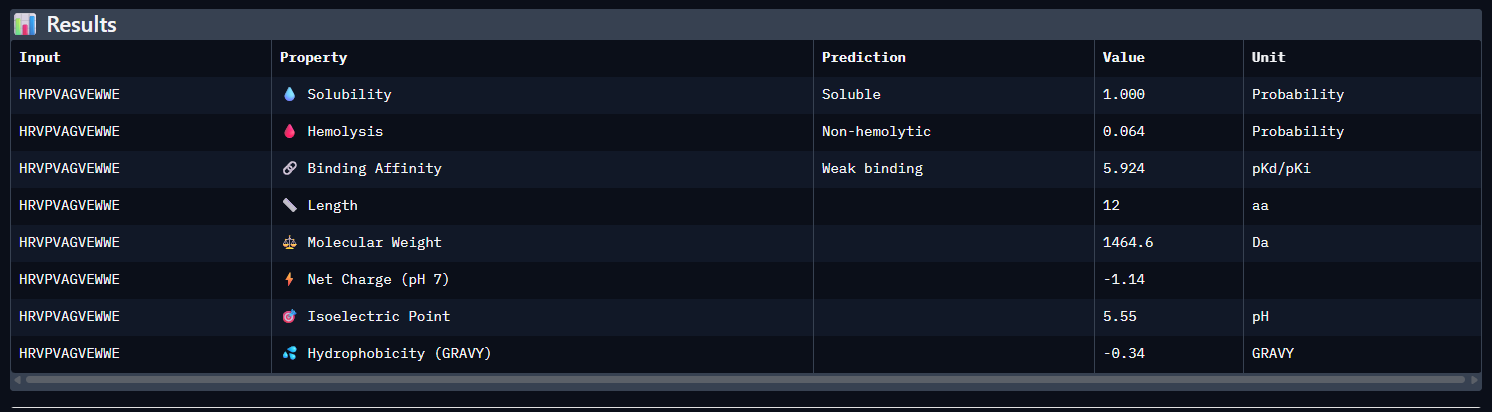
Across all five peptides, PeptiVerse predicts solubility = 1.000 and non-hemolytic behavior (hemolysis probabilities 0.035–0.064), so none of the candidates are flagged as poorly soluble or strongly hemolytic. Predicted binding affinities (pKd/pKi) vary and do not track ipTM: the highest-ipTM peptide (WSVPVVAIEHGE, ipTM 0.44) has the lowest predicted affinity (5.338), while WRYGAAAVEWKE has a higher predicted affinity (6.526) but slightly lower ipTM (0.41).
The known binder (FLYRWLPSRRGG) shows mid-range predicted affinity (5.962) and ipTM (0.30). Considering binding prediction plus safety-like properties, WRYGAAAVEWKE best balances the set: it has the highest predicted affinity (6.526), is predicted soluble (1.000), and has low hemolysis probability (0.047), while still achieving a relatively higher ipTM (0.41) compared to most others.
Peptide to advance: WRYGAAAVEWKE - it is predicted to be soluble, low-hemolysis, and has the strongest predicted binding affinity among the tested peptides, with moderate (though still low-confidence) structural support from AlphaFold3 (ipTM 0.41).
Part 4: Generate Optimized Peptides with moPPIt
I used the moPPIt Colab on a GPU runtime and pasted the A4V mutant SOD1 sequence (mature form without initiator Met). Here’s my collab copy.
I set binder length to 12 aa and generated a pool of candidate peptides using multi-objective guidance. I enabled affinity guidance and included solubility and hemolysis guidance to bias toward more developable peptides.
| Binder (12-aa) | Solubility | Half-life | Affinity |
|---|---|---|---|
| EWWRERLRQTLI | 0.5833 | 0.5833 | 6.0163 |
| EDWLATLRAATS | 0.5000 | 5.9279 | 5.7517 |
| EEEWRQLQSQYE | 0.8333 | 4.4313 | 6.8902 |
| TEEEGVRWKRGV | 0.7500 | 4.0548 | 6.4628 |
| ELLQWILGITIE | 0.4167 | 13.4681 | 6.1644 |
Compared to PepMLM, moPPIt produces peptides shaped by explicit objectives. PepMLM peptides were more diverse but less controlled with respect to developability properties whereas moPPIt candidates tend to show stronger biases in composition, more consistent physicochemical properties across candidates, and often a narrower “design family” reflecting the guidance constraints. On this run, the moPPIt outputs are more compositionally biased toward charged residues (E/D and R/K), consistent with explicit optimization for solubility and half-life alongside affinity. Here’s a summary interpretation of the results:
- Best predicted affinity: EEEWRQLQSQYE (6.8902)
- Best predicted solubility: EEEWRQLQSQYE (0.8333)
- Best predicted half-life: ELLQWILGITIE (13.4681)
- Most “balanced” if you prioritize binding + solubility: EEEWRQLQSQYE (top on both, but not top half-life)
- Most “balanced” if you prioritize half-life strongly: ELLQWILGITIE (best half-life, but lowest solubility)
Before any clinical consideration, I would follow a staged evaluation: (1) in silico screening for interface plausibility (AlphaFold3 ipTM/PAE consistency across seeds) plus basic developability predictions (solubility, hemolysis, aggregation risk); (2) in vitro binding assays (SPR/BLI or competition ELISA), stability in serum, and cytotoxicity/hemolysis assays; (3) cell-based assays for functional effect and off-target toxicity; (4) only after robust preclinical evidence, proceed to in vivo PK/PD and safety studies. In other words, moPPIt designs are hypotheses that must be filtered by structural consistency and validated experimentally before any translational claims.
Part B: BRD4 Drug Discovery Platform Tutorial (Gabriele) Since this was an optional part, I decided to skip for now.
Part C: Final Project: L-Protein Mutants
Phage Lysis Protein Design Challenge
L-Protein Engineering | Option 1: Mutagenesis
I ran the mutational scoring notebook to obtain per-substitution LLR scores and shortlisted mutations with positive scores.
| Position | Wild_Type_AA | Mutation_AA | LLR_Score |
|---|---|---|---|
| 50 | K | L | 2.561464 |
| 29 | C | R | 2.395425 |
| 39 | Y | L | 2.241777 |
| 29 | C | S | 2.043149 |
| 9 | S | Q | 2.014323 |
| 29 | C | Q | 1.997047 |
| 29 | C | P | 1.971026 |
| 29 | C | L | 1.960644 |
| 50 | K | I | 1.928798 |
| 53 | N | L | 1.864930 |
| 61 | E | L | 1.818097 |
| 52 | T | L | 1.813966 |
| 50 | K | F | 1.802066 |
| 29 | C | T | 1.797245 |
| 29 | C | K | 1.795876 |
| 5 | F | Q | 1.795244 |
| 5 | F | R | 1.659717 |
| 29 | C | A | 1.648654 |
| 27 | Y | R | 1.628060 |
| 22 | F | R | 1.602028 |
| 5 | F | P | 1.596888 |
| 50 | K | V | 1.594572 |
| 50 | K | S | 1.574555 |
| 5 | F | T | 1.559023 |
| 5 | F | S | 1.556416 |
| 45 | A | L | 1.539248 |
| 39 | Y | S | 1.517457 |
| 27 | Y | S | 1.497052 |
| 40 | V | L | 1.477630 |
| 27 | Y | L | 1.474637 |
I then intended to cross-check each shortlisted mutation against the experimental mutant dataset (L-Protein Mutants) to see whether the experimental lysis phenotype is directionally consistent with the LLR score.
Only 6 substitutions from my scored shortlist overlapped with the experimental table (C29R, C29S, K50I, K50S, Y27S, Y39S). In the experimental dataset, all overlapping substitutions were labeled as non-lytic (Lysis = 0) despite having positive LLR scores in the notebook. This suggests that, for MS2 L-protein, sequence-only language-model scores may not reliably capture key determinants of lysis (likely influenced by membrane insertion, oligomerization, and host-factor dependence). We therefore should treated LLR scores as a hypothesis generator, not a predictor of functional lysis.
I selected five single-point variants, including two mutations in the soluble region (positions 1–40) and three in the transmembrane region (TM) (positions 41–75), as required.
WT (MS2 L-protein, 75 aa): METRFPQQSQQTPASTNRRRPFKHEDYPCRRQQRSSTLYVLIFLAIFLSKFTNQLLLSLLEAVIRTVTTLQQLLT
I selected five single substitutions with positive LLR scores. I enforced the assignment constraint by choosing two mutations in the soluble region (positions 1–40) and three in the transmembrane region (positions 41–75).
Here are the 5 mutants I choose:
Mutant 1 - S9Q (soluble, LLR = 2.014)
Sequence: METRFPQQQQQTPASTNRRRPFKHEDYPCRRQQRSSTLYVLIFLAIFLSKFTNQLLLSLLEAVIRTVTTLQQLLT Rationale: High positive score in the soluble region (putative DnaJ-interaction domain). Ser→Gln increases hydrogen-bonding potential and may alter surface chemistry without strongly destabilizing the fold.
Mutant 2 - C29R (soluble, LLR = 2.395)
Sequence: METRFPQQSQQTPASTNRRRPFKHEDYPRRRQQRSSTLYVLIFLAIFLSKFTNQLLLSLLEAVIRTVTTLQQLLT Rationale: One of the strongest positive-scoring substitutions in the soluble region. Adds a positive charge that could reshape chaperone-recognition or interaction surfaces.
Mutant 3 - A45L (TM, LLR = 1.539)
Sequence: METRFPQQSQQTPASTNRRRPFKHEDYPCRRQQRSSTLYVLIFLLIFLSKFTNQLLLSLLEAVIRTVTTLQQLLT Rationale: Hydrophobic substitution in the transmembrane segment. Ala→Leu increases hydrophobicity and may stabilize membrane helix packing/insertion and oligomer stability.
Mutant 4 - T52L (TM, LLR = 1.814)
Sequence: METRFPQQSQQTPASTNRRRPFKHEDYPCRRQQRSSTLYVLIFLAIFLSKFLNQLLLSLLEAVIRTVTTLQQLLT Rationale: Polar→hydrophobic change in the TM region. Thr→Leu may increase membrane compatibility and reduce local insertion/misfolding penalties.
Mutant 5 - N53L (TM, LLR = 1.865)
Sequence: METRFPQQSQQTPASTNRRRPFKHEDYPCRRQQRSSTLYVLIFLAIFLSKFTLQLLLSLLEAVIRTVTTLQQLLT Rationale: Polar→hydrophobic change in the TM region with a strong positive score. Selected as an additional TM-stabilizing candidate.