Individual Final Project: Melanin-based light-recording bioink/biomaterial
Important links:
- Commited Listener Slide Deck here.
- Benchling (TO BE ADDED)
- Asimov Kernel (TO BE ADDED)

Aim 1: Build a first melanin-producing cell-free DNA module based on melA tyrosinase + Define validation parameters
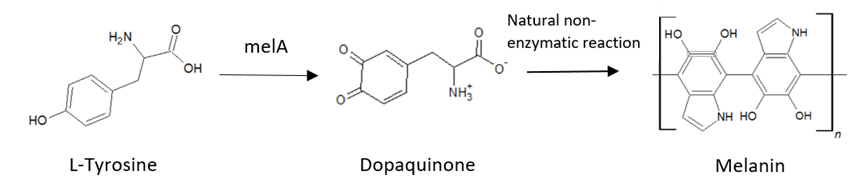
The melA gene is coding sequence of tyrosinase that catalizes the conversion of tyrosine to dopaquinone. Dopaquinone is intermediate product of melanin biosynthesis pathway that polymerizes in an enzyme-independent reaction to form melanin.
The pathway from L-tyrosine to Melanin with the use of the melA tyrosinase.
1. Design MelA expression constructs (Benchling and Twist)
Some of the construct variables to be considered because it will affect melA expression:
- Promoter
- RBS
- Terminator
- Codon usage
- Tag placement
- Vector context
Note: a construct that works in cells may not translate well to cell-free expression.
Previous expression constructs:
iGEM 2004 - using the P(lac)IQ promoter (BBa_I14032), a work designed by Vikram Vijayan, Allen Hsu, Lawrence Fomundam Group of the iGEM04 (2004-08-04 Part:BBa_I14032).
iGEM 2009 - using B0040 RBS (composite part: BBa_K193602) containing pLacIQ(BBa_I14032), RBS(B0030) and melA(BBa_K193600) on low copy vector(BBa_I52001 derived), a work designed by Kazuaki Amikura.
iGEM 2017 - Erin Kelly Group (2017-10-27) have developed a different expression construct specifically for use in E. coli BL21(DE3) which takes advantage of the T7 RNA Polymerase expression construct in the DE3 cassette to provide tighter epxression control and help to prevent leaky expression of the tyrosinase. Additionally, a double terminator B0015 was added to increase control over the system. In order to maximize production of the tyrosinase and limit unnecessary energy expenditure by the cell, a transcriptional terminator ensure energy is not wasted on transcribing an overly-long mRNA transcript. They transformed the melA_pJET plasmid into E. coli BL21(DE3) and attempted to overexpress the MelA tyrosinase (~54kDa) and produce the pigment melanin. Four colonies from the transformation were picked and used to produce pre-cultures, which were then used to incoulate test expression cultures. During the test expression, cultures were also supplied with CuSO4 and extra tyrosine. Cultures were induced with IPTG at OD600 ~1 and a 1OD sample was taken (T0). Another 1OD sample was taken after the cultures were left to grow overnight (TON). The cultures were allowed to grow another three days (supplemented with tyrosine and ampicillin) to see if pigment would form, but we were unable to detect any melanin. The 1OD samples were run on a 12% SDS-PAGE to check for melA overexpression (Figure 3). The MelA tyrosinase is ~54 kDa in size. A faint band of approximately 54 kDa appears in the TON lane of culture 3. This indicated that we were successful in expressing the MelA tyrosinase from the pJET plasmid. Before the Jamboree, we will attempt another overexpression of MelA from the pSB1C3 plasmid.
My take: T7 can maximize protein yield but also overwhelm folding capacity, causing inactive protein accumulation (increase the likehood of tyrosinases misfolds, aggregation, or fail to incorporate copper correctly). I’d replace it by a moderated construct and compare the results in reference to the BBa_K2481108 (control).
Note: MelA expression is not the same as melanin production. Melanin polymerization is messy, dopaquinone polymerizes through non-enzymatic downstream chemistry. This means color output depends not only on MelA, but also on oxygen, pH, time, redox state, and local chemistry.
2. Prepare reagents and workflow (Ginko & Open AI, HW 11C)
Melanin production in E. coli or in a cell-free system is influenced by several parameters that actuate at the level of melA expression and enzyme activity / posterior reactions:
L-tyrosine concentration (substrate, limited solubility)
CuSO4 concentration: since this tyrosinase is a type 3 copper-containing enzyme, Cu2+ is a cofactor of the enzyme. Too much copper can also stress cells or inhibit cell-free reactions.
Magnesium
Energy mix
Molecular oxigen avaliability for tyrosinase reactions
pH: tyrosinase activity and melanin polymerization are pH-dependent. If the reaction acidifies over time, enzyme activity or pigment formation may decrease.
Note: Optimizing for sfGFP may not optimize for MelA.
3. Validation
The 2017 iGEM result is a useful warning: they may have produced a faint ~54 kDa MelA band, but still detected no pigment. For this reason, I am proposing a staged validation workflow that moves from simple expression and pigment checks to more refined mechanistic analyses, depending on the results obtained in each previous round.
| Order | Method | What it tells you | Hypothesis tested |
|---|---|---|---|
| 1 | HW11 protein fluorescence | General cell-free expression capacity | If fluorescence is strong, the TX-TL system is functional and supports protein expression. |
| 2 | Petri dish photos | Visual pigment output (qualitative) | If colonies/spots darken over time, the construct produces visible pigment. |
| 3 | Spectrophotometric OD 400–500 nm | Pigment formation kinetics | If absorbance increases over time, MelA is producing melanin-like pigment. |
| 4 | SDS-PAGE (MelA ~54 kDa) | MelA protein expression | If a ~54 kDa band is present, the construct expresses MelA protein. |
| 5 | LC-MS (tyrosine / L-DOPA) | Pathway-level activity (mechanistic) | If tyrosine decreases and intermediates increase, MelA is catalytically active even without visible pigment. |
Results interpretation framework:
| Observation | Interpretation |
|---|---|
| Strong MelA band + no pigment | Expression works, but enzyme is likely inactive (folding, copper, pH, oxygen, or substrate issue). |
| Weak MelA band + strong pigment | Low expression but high enzymatic efficiency; not expression-limited. |
| Strong MelA band + strong pigment | Optimal case: high expression and active enzyme. |
| Weak MelA band + no pigment | Expression failure (construct, transcription, translation, or stability issue). |
| MelA band + LC-MS intermediates + weak pigment | Enzyme is active, but pigment polymerization or accumulation is limiting. |
| MelA band + no LC-MS intermediates + no pigment | Enzyme is expressed but inactive (likely folding or cofactor issue). |
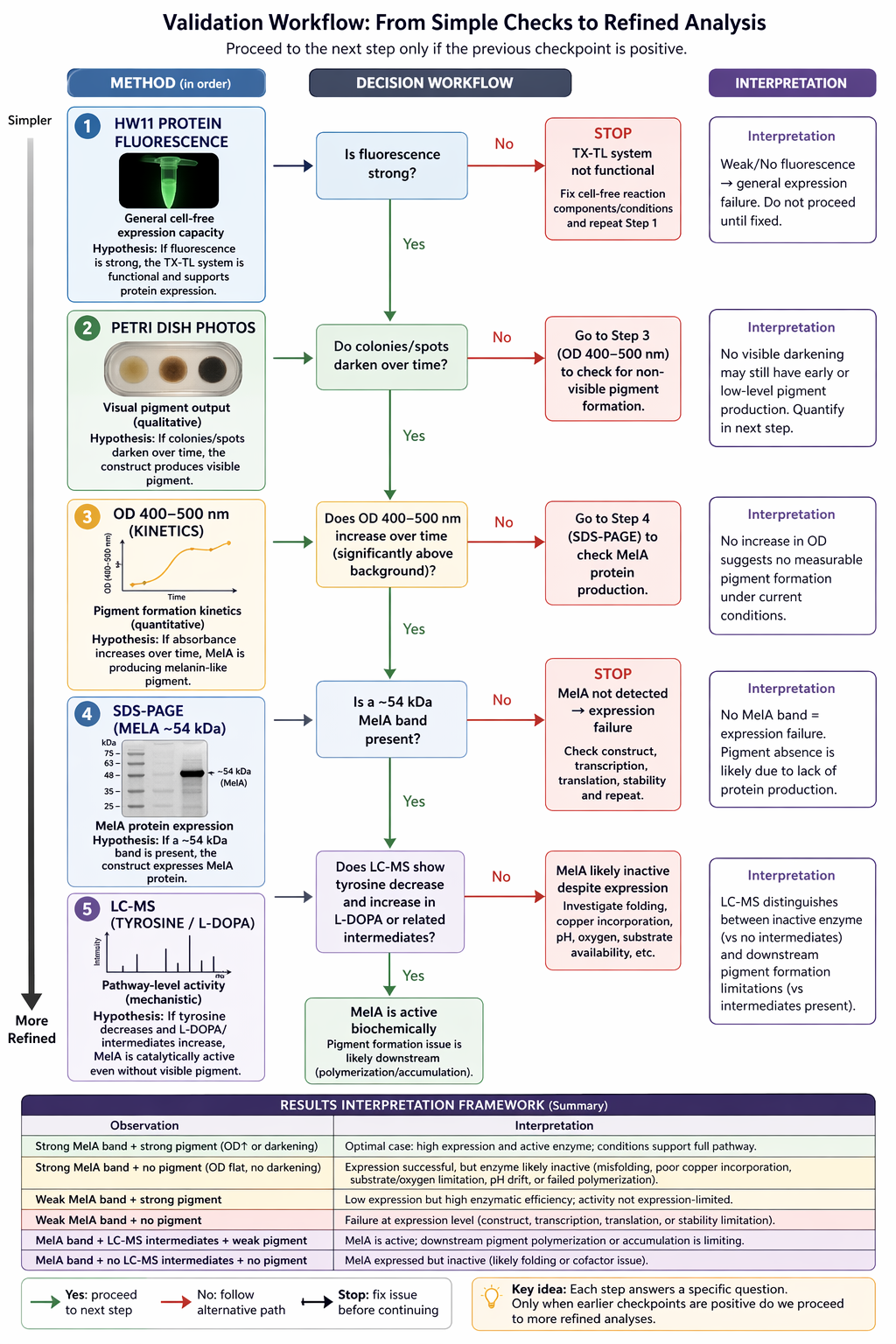
Here’s a diagram of my proposed validation workflow
4. Model a light-activated expression circuit that could later support gradual tonal change in a material system (Asimov Kernel).
Post-Course: Towards the melanin-based light-recording bio-ink
- Refine Asimov Kernel’s output for controlling melanin expression till the system is fine-tuned with my aesthetical needs (most controlled and previsible) or, if not possible develop a experimental design for embedded sensing (in this case map both the quantitative and qualitative workflows available).**
Decision point: Push for maximum molecular control first, even if the material context is still abstract? OR Move earlier into material-scale experiments?
Material engineering: Test different models of integration of the melanin cell-free module towards a intended function/product. Tests for integrating into material though ELM engineer Komagataeibacter rhaeticus + bacterial celulose (BC) or a hybrid system with BC scaffold / other biomaterial that can be embed with cell-free modules with synthetic minimal cells / K. rhaeticus.
Rethink the workflow for measurable readouts for the cell-free system. RGB image analysis in a controlled lighting box or spectral (semi-quantitative) with melanin absorbance ranging from 475 nm to 500 nm;
Optimization rounds looking into every step of the proposed workflow, including economic tests for modeling parameters (such as ink dilution)
Benchmarking partnerships + reach out plan (heavly based on the previous item conclusions (BioFabricate, Cultivarium)