Week 2 HW: DNA, READ, WRITE AND EDIT!


Geeking out over protein structures and data banks, DNA storage in plants, clouds and decoding DNA into sound
I love that artist Antoine Bertin has decoded the RNA of SARS COV 2 into this track! check it out.
This is the RNA of the Coronavirus translated into sound (viruses are made of RNA, not exactly DNA). Each nucleotide of the RNA (A,U,G or C) is transformed into a note so the virus sequence can be heard. The tempo of the track follows the rhythm at which the epidemic is growing (exponential curve) and how this curve flattens if we all stay home :) I wanted to create a track that can help with relaxation in times of isolation, and meditate on the fact all life on earth, including viruses, are made of the same material. We (humans, animals, trees, bacteria, viruses) are the continuation of a same common ancestor. Anyway; I hope this will helps everyone explore in their own sonic way what we are going through! Here is an extract of the RNA sequence :)
Wuhan seafood market pneumonia virus isolate Wuhan-Hu-1, complete genome (NC_045512.2)
auuaaagguuuauaccuucccagguaacaaaccaaccaacuuucgaucucuuguagaucuguucucuaaacgaacuuua aaaucuguguggcugucacucggcugcaugcuuagugcacucacgcaguauaauuaauaacuaauuacugucguugaca ggacacgaguaacucgucuaucuucugcaggcugcuuacgguuucguccguguugcagccgaucaucagcacaucuagg uuucguccgggugugaccgaaagguaagauggagagccuugucccugguuucaacgagaaaacacacguccaacucagu uugccuguuuuacagguucgcgacgugcucguacguggcuuuggagacuccguggaggaggucuuaucagaggcacguc aacaucuuaaagauggcacuuguggcuuaguagaaguugaaaaaggcguuuugccucaacuugaacagcccuauguguu caucaaacguucggaugcucgaacugcaccucauggucauguuaugguugagcugguagcagaacucgaaggcauucag uacggucguaguggugagacacuugguguccuugucccucaugugggcgaaauaccaguggcuuaccgcaagguucuuc uucguaagaacgguaauaaaggagcugguggccauaguuacggcgccgaucuaaagucauuugacuuaggcgacgagcu uggcacugauccuuaugaagauuuucaagaaaacuggaacacuaaacauagcagugguguuacccgugaacucaugcgu gagcuuaacggaggggcauacacucgcuaugucgauaacaacuucuguggcccugauggcuacccucuugagugcauua aagaccuucuagcacgugcugguaaagcuucaugcacuuuguccgaacaacuggacuuuauugacacuaagaggggugu auacugcugccgugaacaugagcaugaaauugcuugguacacggaacguucugaaaagagcuaugaauugcagacaccu
I wanna read, write and edit DNA!!!
I had twisted sister in my mind while I was saying this, particularly I WANNA ROCK.


The 2nd week has been again packed with new information but I cannot wait to read, write and edit DNA as this it totally new information.
Week 2- DNA Read, Write, & Edit HW
This week explores the read–write–edit toolkit: sequencing and synthesis workflows, restriction digests and gel electrophoresis, and early genome-editing frameworks.
Make sure to document every step of the in-silico and lab experiments. Make sketches, screenshots, notes, drawings… anything that helps you - and others - understand the experiment.
Part 0: Basics of Gel Electrophoresis
Gel electrophoresis separates DNA fragments based on size using:
Negatively charged DNA backbone Electric field Agarose matrix Size-dependent migration
I attended and watched all lecture and recitation videos apart from the one last week on Thursday, the first meetup with Tokyo Bioclub node because I was setting up an exhibition and because with the time difference I did not see the email on time but I watched the recording :)
How does gel electrophoresis work?!
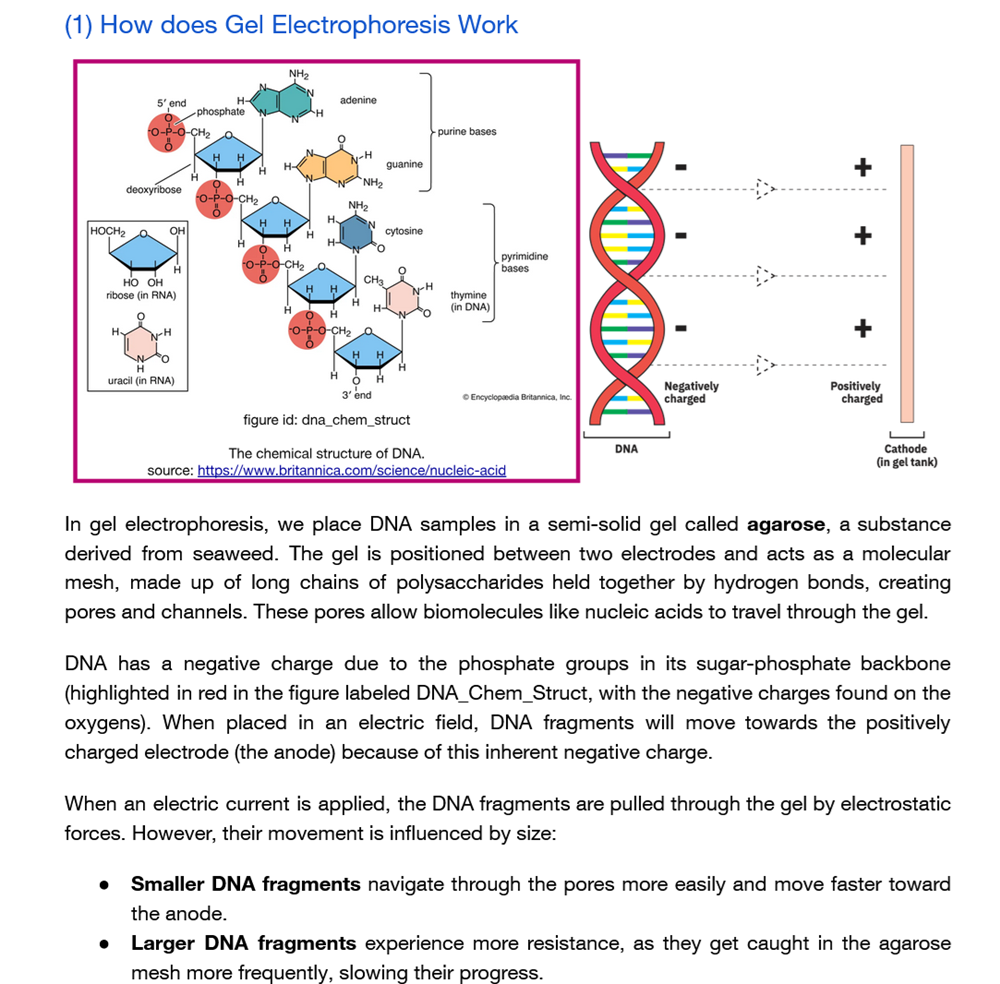



…and what does it look like?
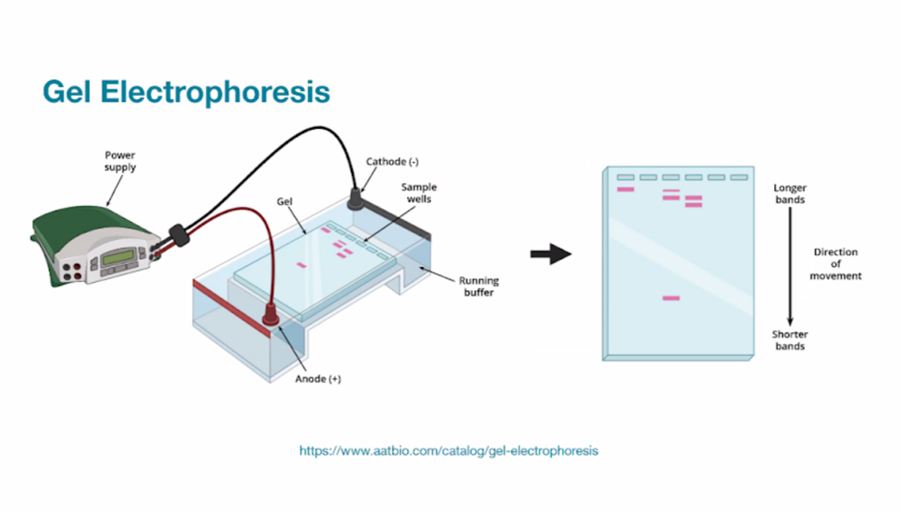

I have known for a while how it looks like but I never really looked properly into it. I have been working with agar for a while now due to making biomaterials for textiles and edible materials too. In addition, I have also worked with other polymers too such as different kinds of alginate, gelatin and different kinds of starch.
Part 1: Benchling & In-silico Gel Art
See the Gel Art: Restriction Digests and Gel Electrophoresis protocol for details.
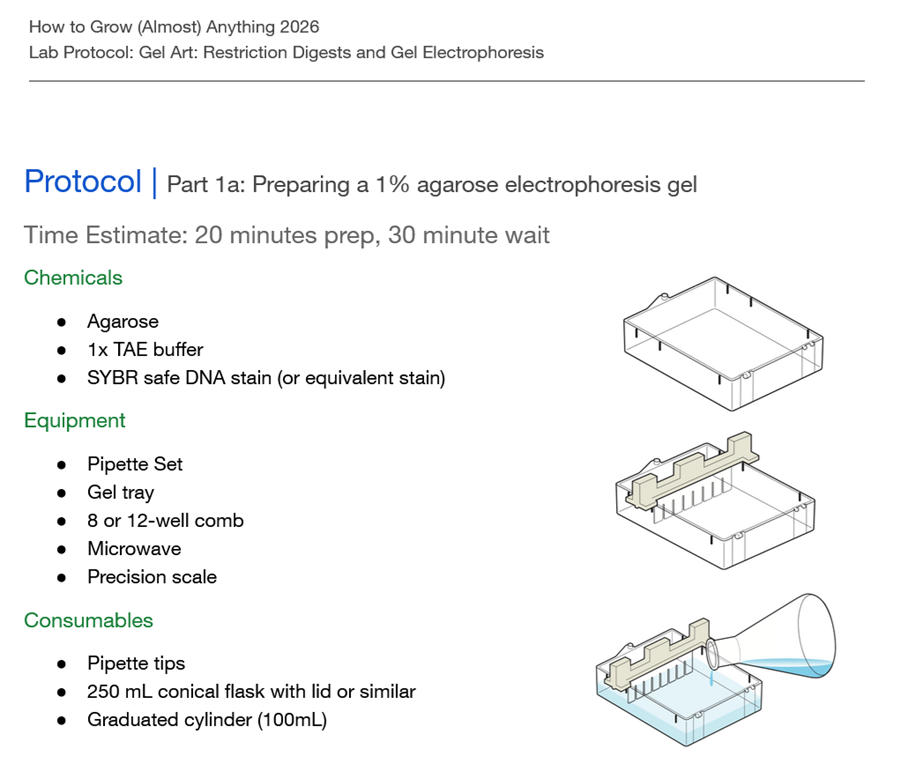

•Overview:
- Make a free account at benchling.com
It was super easy! I logged in with my google account.
- Import the Lambda DNA
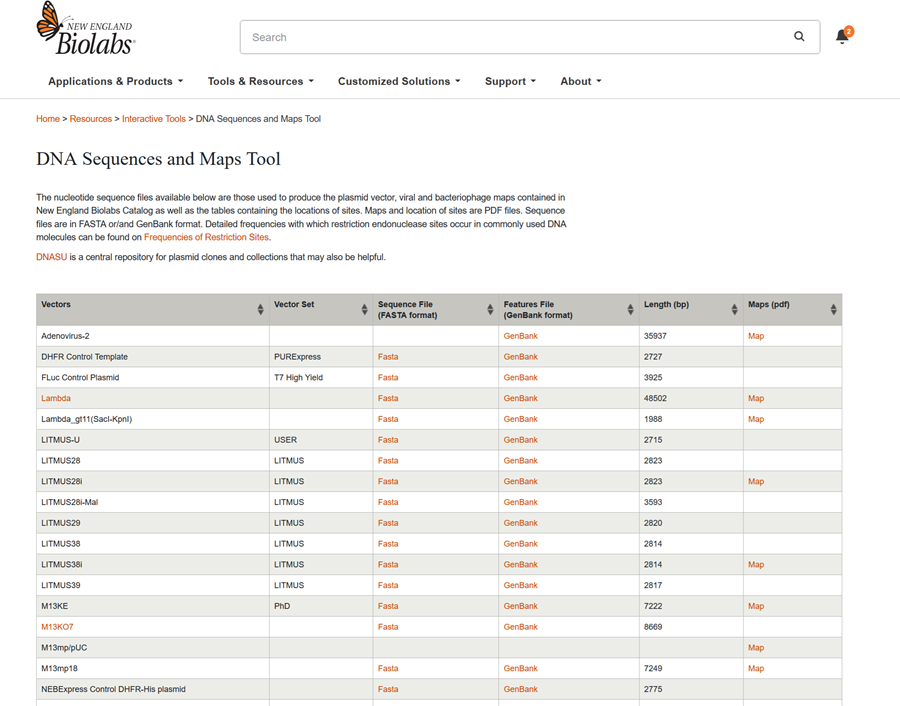

This is what the DNA sequence looks like in FASTA SEQUENCE FORMAT! I saved the file in a file document because it was the only available option on the the neb.com website. I did right click and saved in file format. Let’s see if we can import it like this in benchling!
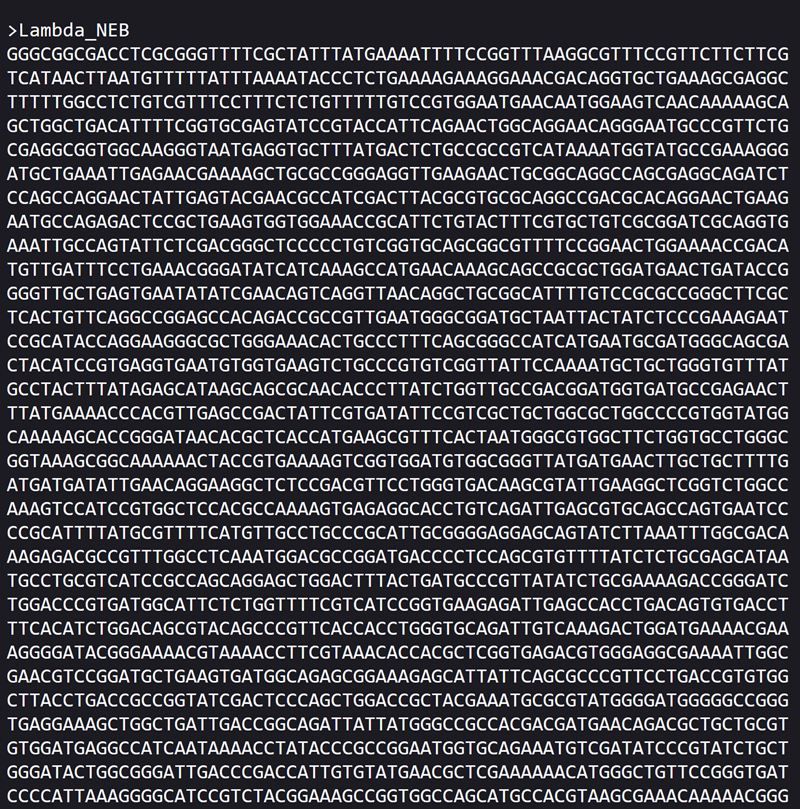

Importing the lambda DNA sequence in benchling
First I created a new project on benchling named ‘htgaa week 2 - MARISA SATSIA’.
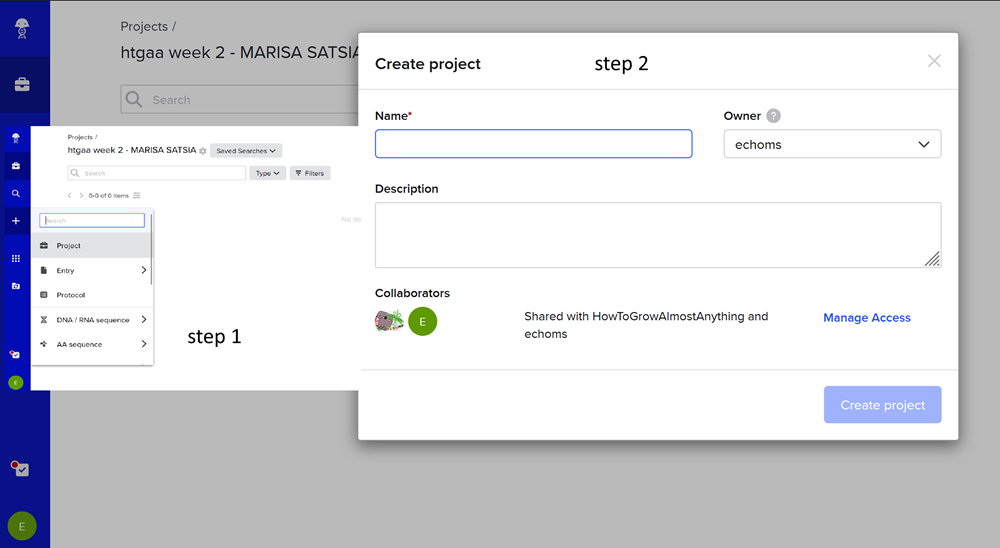

Then i imported the DNA!
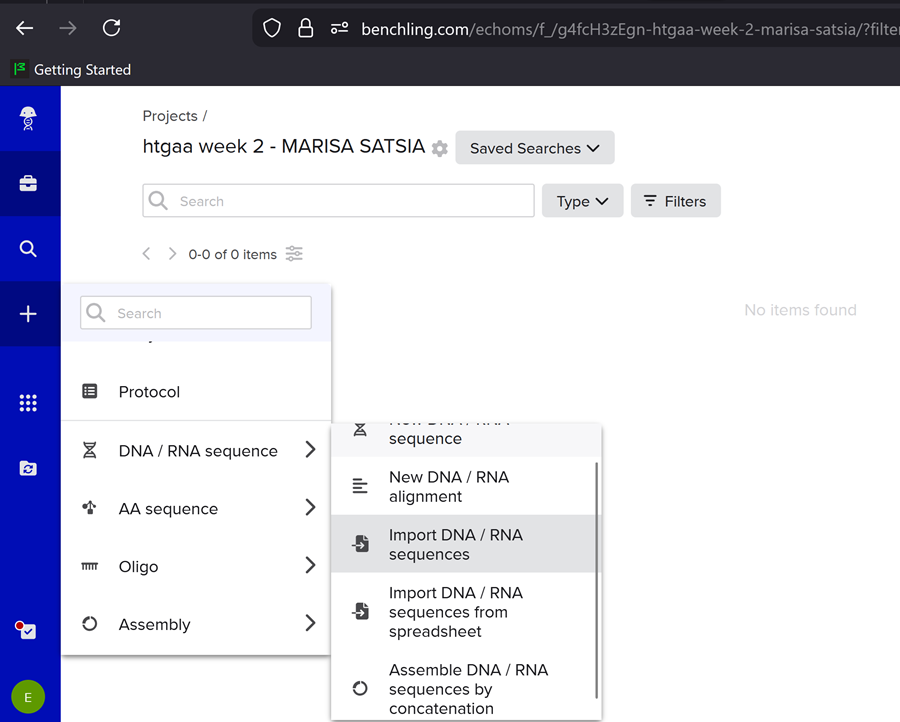

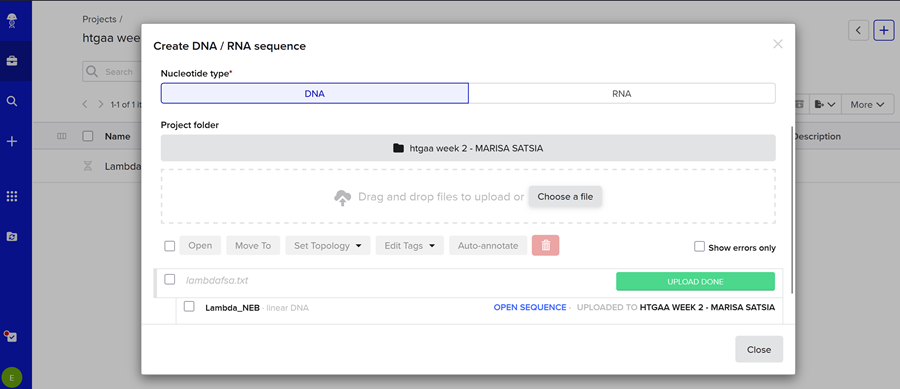

Then I clicked on open sequence and VOILA!
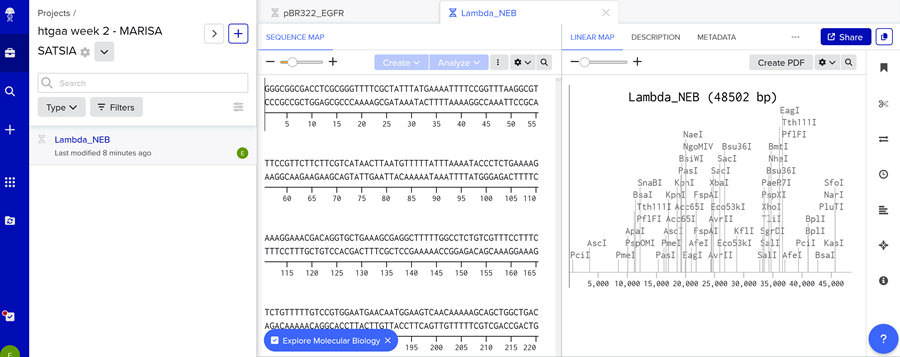

- Simulate Restriction Enzyme
You might wonder what a restriction enzyme is right?!


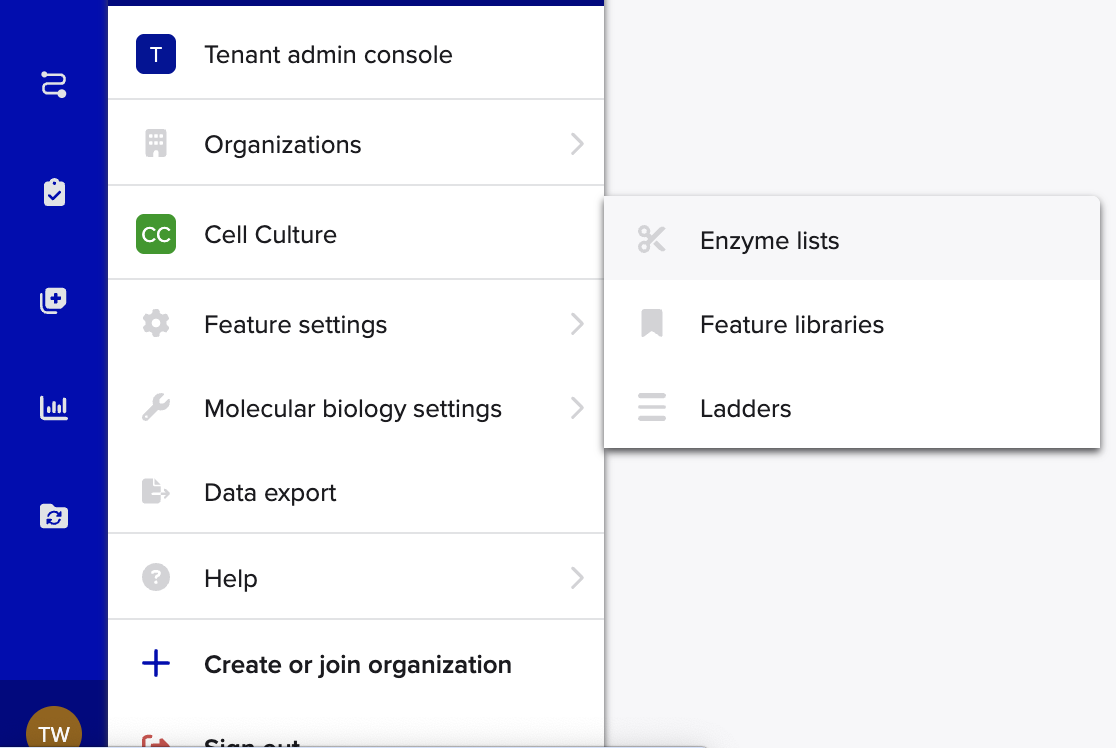

- Simulate Restriction Enzyme Digestion with the following Enzymes:
EcoRI
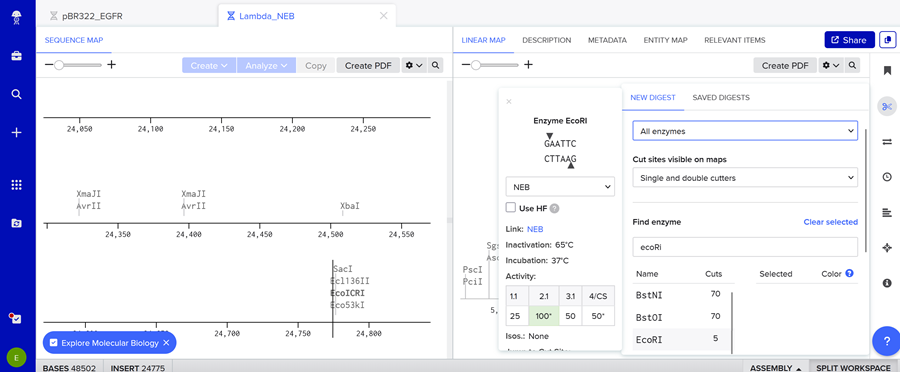

HindIII BamHI KpnI EcoRV SacI SalI
Here is the enzyme digest simulation with all the enzymes!
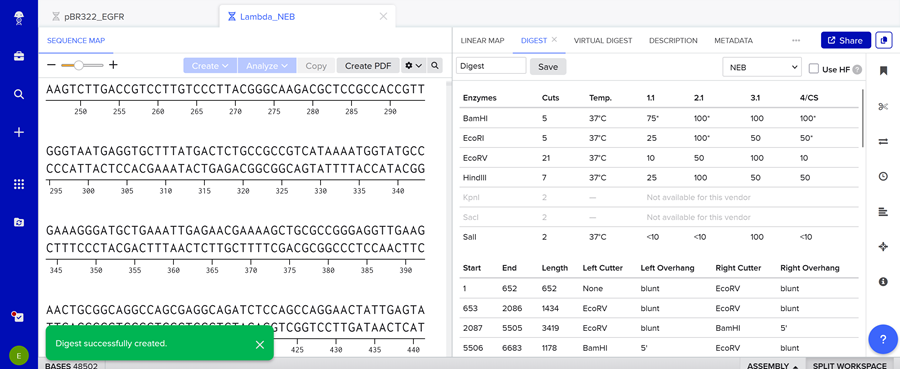

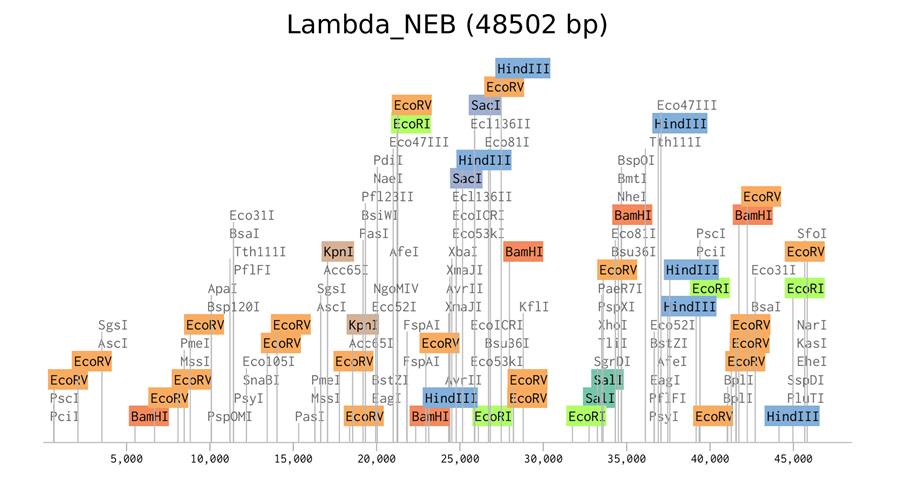

Here is the ladder simulation
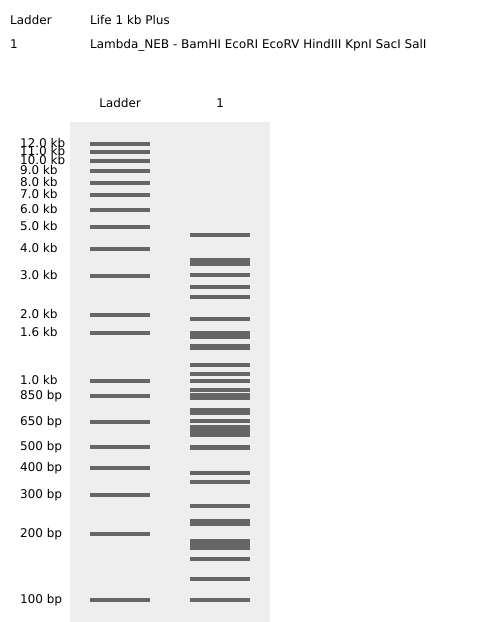

- Create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks and 6. -> You might find Ronan’s website, a helpful tool for quickly iterating on designs!
I made this using Ronan’s website. I think it is pretty cool to simulate this whole process and have a visual because I do not know when I am actually gonna do the lab!
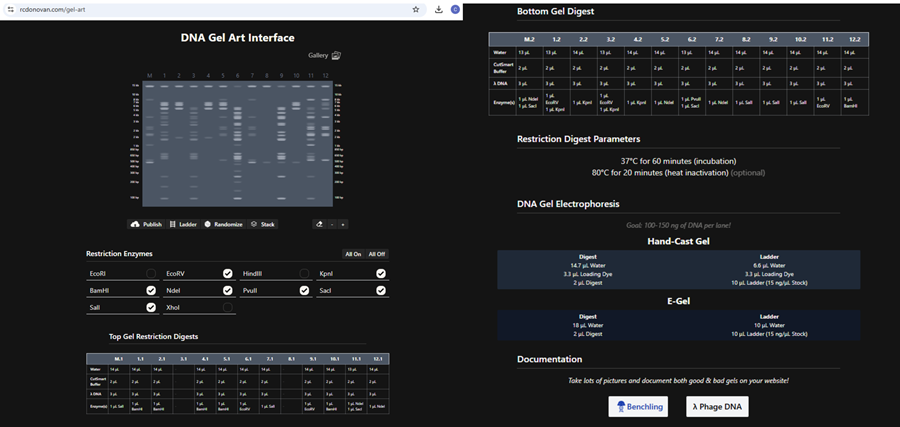

Part 2: Gel Art - Restriction Digests and Gel Electrophoresis
Perform the lab experiment you designed in Part 1 and outlined in the Gel Art: Restriction Digests and Gel Electrophoresis protocol.
Unfortunately I cannot do that here in Cyprus, but I am actively looking for a lab to let me practice a bit.


Part 3: DNA Design


Part 3.1. Choose your protein
In recitation, we discussed that you will pick a protein for your homework that you find interesting. Which protein have you chosen and why? Using one of the tools described in recitation (NCBI, UniProt, google), obtain the protein sequence for the protein you chose.
[Example from our group homework, you may notice the particular format — The example below came from UniProt]
sp|P03609|LYS_BPMS2 Lysis protein OS=Escherichia phage MS2 OX=12022 PE=2 SV=1 METRFPQQSQQTPASTNRRRPFKHEDYPCRRQQRSSTLYVLIFLAIFLSKFTNQLLLSLL EAVIRTVTTLQQLLT
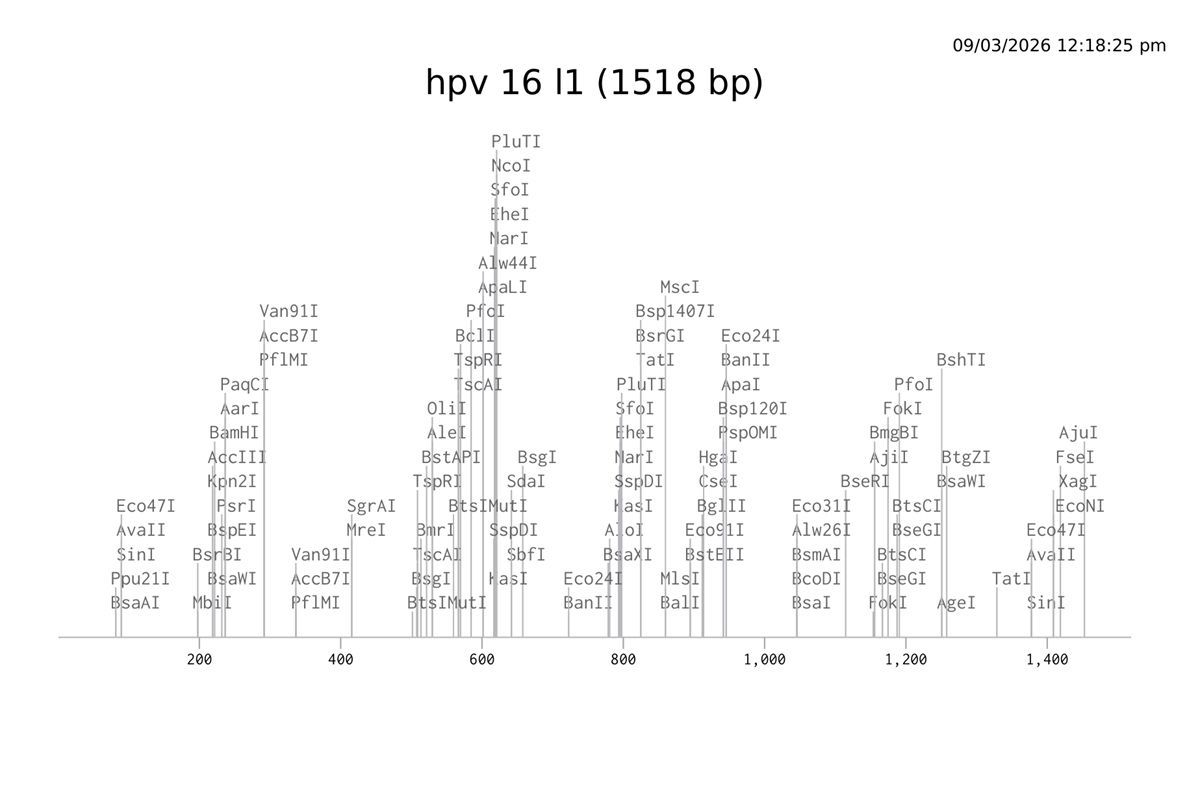
I choose the HPV genome proteins L1 (HPV16-L1) and L2(HPV16-L2). The HPV genome is surrounded by an icosahedral capsid consisting of two structural proteins: the major capsid protein L1 (HPV16-L1) and the minor capsid protein L2 (HPV16-L2). The L1 proteins are highly conserved and aggregate to form 72 fivefold capsomers. The L2 protein binds viral DNA. There are multiple types of HPV unfortunately and each affects us differently. Some types cause cervical cancer and some warts. There is an mRNA vaccine which I got when it first came out in 2007 or 2008 or 2009, when I was 18 or 19, I do not exactly remember.
L1 Protein Lengths by HPV Type
The L1 gene encodes the major capsid protein of the Human Papillomavirus (HPV), which spontaneously self-assembles into virus-like particles (VLPs)).
Because HPV has over 100 different genotypes, the exact sequence length varies slightly:
HPV 16: 505 amino acids (Prototype ID: P03101). HPV 18: 568 amino acids (UniProt ID: T2A5K9). HPV 51: 504 amino acids (UniProt ID: P26536).
This is what AI mode in google mentioned!


Below is the FASTA sequence for the L1 Major Capsid Protein of HPV Type 16, the strain responsible for approximately 50% of all cervical cancer cases worldwide. HPV 16 L1 Protein Sequence (UniProt P03101). This protein is 505 amino acids long and is the primary antigen used in HPV vaccines like Gardasil.
L1 SEQUENCE
sp|P03101|VL1_HPV16 Major capsid protein L1 OS=Human papillomavirus type 16 OX=333760 GN=L1 PE=1 SV=1
MSLWLPSEATVYLPPVPVSKVVSTDEYVARTNIYYHAGTSRLLAVGHPYFPIKKPNNNKI LVPKVSGLQYRVFRIHLPDPNKFGFPDTSFYNPDTQRLVWACVGVEVGRGQPLGVGISGH PLLNKLDDTENASAYAANAGVDNRECISMDYKQTQLCLIGCKPPIGEHWGKGSPCTNVAV NPGDCPPLELINTVIQDGDMVHTGFGAMDFTTLQANKSEVPLDICTSICKYPDYIKMVSE PYGDSLFFYLRREQMFVRHLFNRAGAVGENVPDDLYIKGSGSTATLANNYYPTPSGSMVT SDAQIFNKPYWLQRAQGHNNGICWGNQLFVTVVDTTRSTNMSLCAAISTSETTYKNTNFK EYLRHGEEYDLQFIFQLCKITLTADVMTYIHSMNSTILEDWNFGLQPPPGGTLEDTYRFV TSQAIACQKHTPPAPKEDDPLKKYTFWEVNLKEKFSADLDQFPLGRKFLLQAGLKAKPKF TLGKRKATPTTSSTSTTAKRKKRKL
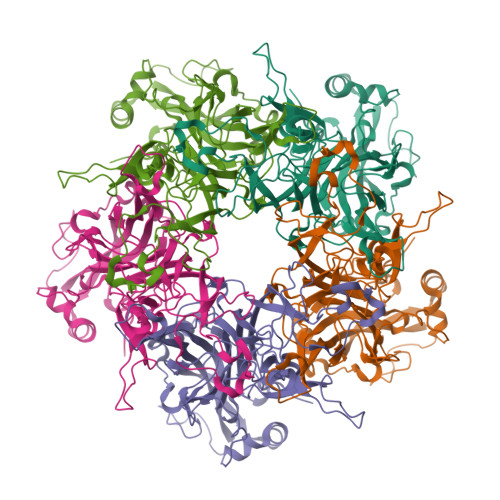
I also got this Pentamer Structure of Major Capsid protein L1 of Human Papilloma Virus type 11 from the 3d viewer from the RCSB PDB I love the 3d visualisation tool and the fact that you can isolate things and make animations and download 3d models.

Part 3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence
The Central Dogma discussed in class and recitation describes the process in which DNA sequence becomes transcribed and translated into protein. The Central Dogma gives us the framework to work backwards from a given protein sequence and infer the DNA sequence that the protein is derived from. Using one of the tools discussed in class, NCBI or online tools (google “reverse translation tools”), determine the nucleotide sequence that corresponds to the protein sequence you chose above.
Example: Get to the original sequence of phage MS2 L-protein from its genome. The LYSIS protein DNA sequence below-
atggaaacccgattccctcagcaatcgcagcaaactccggcatctactaatagacgccggccattcaaacatgaggattacccatgtcgaagacaacaaagaagttcaactctttatgtattgatcttcctcgcgatctttctctcgaaatttaccaatcaattgcttctgtcgctactggaagcggtgatccgcacagtgacgactttacagcaattgcttacttaa
This is the FASTA sequence of phage MS2 DNA genome on the website:


For the HPV 16 L1 Protein Sequence (UniProt P03101) the reverse translation or reverse engineering sequence iiiiisssss:
NC_001526.4:5560-7077 Human papillomavirus type 16 (HPV16), L1 major capsid protein ATGAGCCTGTGGCTGCCCAGCGAGGCCACCGTGTACCTGCCTCCCGTGCCCGTGTCCAAG GTGGTGAGCACCGACGAGTACGTGGCCCGGACCAACATCTACTACCACGCCGGCACCAGC CGCCTGCTGGCCGTGGGCCACCCCTACTTCCCCATCAAGAAGCCCAACAACAACAAGATC CTGGTGCCCAAGGTGAGCGGCCTGCAGTACCGGGTGTTCCGGATCCACCTGCCCGACCCC AACAAGTTCGGCTTCCCCGACACCAGCTTCTACAACCCCGACACCCAGCGGCTGGTGTGG GCCTGCGTGGGCGTGGAGGTGGGCCGGGGCCAGCCCCTGGGCGTGGGCATCAGCGGCCAC CCCCTGCTGAACAAGCTGGACGACACCGAGAACGCCAGCGCCTACGCCGCCAACGCCGGC GTGGACAACCGGGAGTGCATCAGCATGGACTACAAGCAGACCCAGCTGTGCCTGATCGGC TGCAAGCCCCCCATCGGCGAGCACTGGGGCAAGGGCAGCCCCTGCACCAACGTGGCCGTG AACCCCGGCGACTGCCCCCCACTGGAGCTGATCAACACCGTGATCCAGGACGGCGACATG GTGCACACCGGCTTCGGCGCCATGGACTTCACCACCCTGCAGGCCAACAAGAGCGAGGTG CCCCTGGACATCTGCACCAGCATCTGCAAGTACCCCGACTACATCAAGATGGTGAGCGAG CCCTACGGCGACAGCCTGTTCTTCTACCTGCGGCGGGAGCAGATGTTCGTGCGGCACCTG TTCAACCGGGCCGGCGCCGTGGGCGAGAACGTGCCCGACGACCTGTACATCAAGGGCAGC GGCAGCACCGCCACCCTGGCCAACAACTACTACCCCACCCCCAGCGGCAGCATGGTGACC AGCGACGCCCAGATCTTCAACAAGCCCTACTGGCTGCAGCGGGCCCAGGGCCACAACAAC GGCATCTGCTGGGGCAACCAGCTGTTCGTGACCGTGGTGGACACCACCCGGAGCACCAAC ATGAGCCTGTGCGCCGCCATCAGCACCAGCGAGACCACCTACAAGAACACCAACTTCAAG GAGTACCTGCGGCACGGCGAGGAGTACGACCTGCAGTTCATCTTCCAGCTGTGCAAGATC ACCCTGACCGCCGACGTGATGACCTACATCCACAGCATGAACAGCACCATCCTGGAGGAC TGGAACTTCGGCCTGCAGCCCCCCCCCGGCGGCACCCTGGAGGACACCTACCGGTTCGTG ACCAGCCAGGCCATCGCCTGCCAGAAGCACACCCCCCCCGCCCCCAAGGAGGACGACCCC CTGAAGAAGTACACCTTCTGGGAGGTGAACCTGAAGGAGAAGTTCAGCGCCGACCTGGAC CAGTTCCCCCTGGGCCGGAAGTTCCTGCTGCAGGCCGGCCTGAAGGCCAAGCCCAAGTTC ACCCTGGGCAAGCGGAAGGCCACCCCCACCACCAGCAGCACCAGCACCACCGCCAAGCGG AAGAAGCGGAAGCTGTAA
Official Reference Information
Database: NCBI GenBank / RefSeq Accession Number: NC_001526.4 Locus Tag: HPV16gp6 (L1) Coordinates: 5560 to 7077 (1518 base pairs) Function: Major capsid protein; self-assembles into virus-like particles (VLPs) used in vaccines.
Part 3.3. Codon optimization
Once a nucleotide sequence of your protein is determined, you need to codon optimize your sequence. You may, once again, utilize google for a “codon optimization tool”. In your own words, describe why you need to optimize codon usage. Which organism have you chosen to optimize the codon sequence for and why?
Example from from Codon Optimization Tool | Twist Bioscience while avoiding Type IIs enzyme recognition sites BsaI, BsmBI, and BbsI.
Lysis protein DNA sequence with Codon-Optimization
ATGGAAACCCGCTTTCCGCAGCAGAGCCAGCAGACCCCGGCGAGCACCAACCGCCGCCGCCCGTTCAAACATGAAGATTATCCGTGCCGTCGTCAGCAGCGCAGCAGCACCCTGTATGTGCTGATTTTTCTGGCGATTTTTCTGAGCAAATTCACCAACCAGCTGCTGCTGAGCCTGCTGGAAGCGGTGATTCGCACAGTGACGACCCTGCAGCAGCTGCTGACCTAA
For the HPV 16 L1 protein DNA sequence with codon-optimization
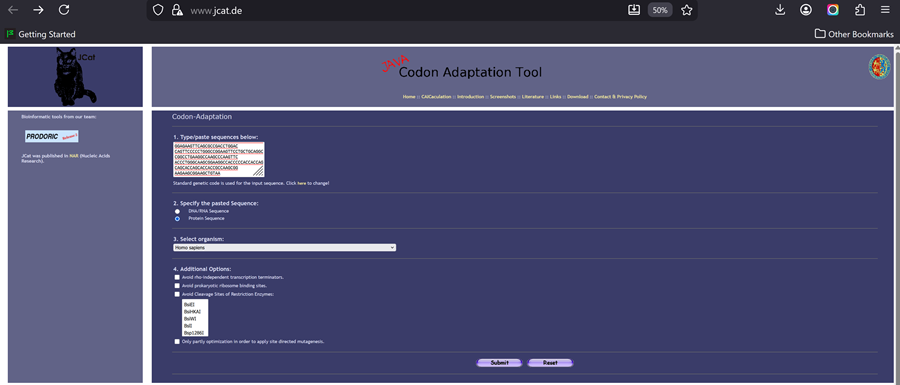
According to AI the preferred codon optimization tool for HPV16 and HPV18, particularly for designing vaccines, is the Java Codon Adaptation Tool (JCat). JCat is used to adapt the codon usage of the HPV genes to the host organism (e.g., E. coli or humans) to improve protein expression.
GC-Content of Homo sapiens: 40.892862223204
Translation ATGAGCCTGTGGCTGCCCAGCGAGGCCACCGTGTACCTGCCTCCCGTGCC 50 CGTGTCCAAGGTGGTGAGCACCGACGAGTACGTGGCCCGGACCAACATCT 100 ACTACCACGCCGGCACCAGCCGCCTGCTGGCCGTGGGCCACCCCTACTTC 150 CCCATCAAGAAGCCCAACAACAACAAGATCCTGGTGCCCAAGGTGAGCGG 200 CCTGCAGTACCGGGTGTTCCGGATCCACCTGCCCGACCCCAACAAGTTCG 250 GCTTCCCCGACACCAGCTTCTACAACCCCGACACCCAGCGGCTGGTGTGG 300 GCCTGCGTGGGCGTGGAGGTGGGCCGGGGCCAGCCCCTGGGCGTGGGCAT 350 CAGCGGCCACCCCCTGCTGAACAAGCTGGACGACACCGAGAACGCCAGCG 400 CCTACGCCGCCAACGCCGGCGTGGACAACCGGGAGTGCATCAGCATGGAC 450 TACAAGCAGACCCAGCTGTGCCTGATCGGCTGCAAGCCCCCCATCGGCGA 500 GCACTGGGGCAAGGGCAGCCCCTGCACCAACGTGGCCGTGAACCCCGGCG 550 ACTGCCCCCCACTGGAGCTGATCAACACCGTGATCCAGGACGGCGACATG 600 GTGCACACCGGCTTCGGCGCCATGGACTTCACCACCCTGCAGGCCAACAA 650 GAGCGAGGTGCCCCTGGACATCTGCACCAGCATCTGCAAGTACCCCGACT 700 ACATCAAGATGGTGAGCGAGCCCTACGGCGACAGCCTGTTCTTCTACCTG 750 CGGCGGGAGCAGATGTTCGTGCGGCACCTGTTCAACCGGGCCGGCGCCGT 800 GGGCGAGAACGTGCCCGACGACCTGTACATCAAGGGCAGCGGCAGCACCG 850 CCACCCTGGCCAACAACTACTACCCCACCCCCAGCGGCAGCATGGTGACC 900 AGCGACGCCCAGATCTTCAACAAGCCCTACTGGCTGCAGCGGGCCCAGGG 950 CCACAACAACGGCATCTGCTGGGGCAACCAGCTGTTCGTGACCGTGGTGG 1000 ACACCACCCGGAGCACCAACATGAGCCTGTGCGCCGCCATCAGCACCAGC 1050 GAGACCACCTACAAGAACACCAACTTCAAGGAGTACCTGCGGCACGGCGA 1100 GGAGTACGACCTGCAGTTCATCTTCCAGCTGTGCAAGATCACCCTGACCG 1150 CCGACGTGATGACCTACATCCACAGCATGAACAGCACCATCCTGGAGGAC 1200 TGGAACTTCGGCCTGCAGCCCCCCCCCGGCGGCACCCTGGAGGACACCTA 1250 CCGGTTCGTGACCAGCCAGGCCATCGCCTGCCAGAAGCACACCCCCCCCG 1300 CCCCCAAGGAGGACGACCCCCTGAAGAAGTACACCTTCTGGGAGGTGAAC 1350 CTGAAGGAGAAGTTCAGCGCCGACCTGGACCAGTTCCCCCTGGGCCGGAA 1400 GTTCCTGCTGCAGGCCGGCCTGAAGGCCAAGCCCAAGTTCACCCTGGGCA 1450 AGCGGAAGGCCACCCCCACCACCAGCAGCACCAGCACCACCGCCAAGCGG 1500 AAGAAGCGGAAGCTGTAA

In your own words, describe why you need to optimize codon usage. Which organism have you chosen to optimize the codon sequence for and why?
For humans and for vaccine development.
Part 3.4. You have a sequence! Now what?
What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both.
AI Overview
The protein used to make HPV vaccines (specifically the L1 major capsid protein) is produced using recombinant DNA technology to create Virus-Like Particles (VLPs). These particles mimic the structure of the actual HPV virus but contain no genetic material, making them non-infectious and incapable of causing disease.
Here is the breakdown of the technology and production systems used:
1. Recombinant Expression Systems
The L1 genes are inserted into host cells that act as "factories" to produce large quantities of the protein. The two main systems are:
+Yeast Cells (Saccharomyces cerevisiae): Used in the production of Gardasil and Gardasil 9.
+Insect Cells (Baculovirus Expression Vector System - BEVS): Trichoplusia ni (Hi-5) cells infected with recombinant baculovirus are used for Cervarix.
2. VLP Self-Assembly and Purification
+Self-Assembly: Once the L1 protein is produced, it spontaneously assembles into VLPs within the host cells.
+Purification: The cells are broken open, and the VLPs are purified through complex physical and chemical processes.
+Adsorption: The purified VLPs are adsorbed onto an aluminum-based adjuvant to improve immune response.
3. Alternative & Emerging Technologies
Research is ongoing to reduce costs and increase production efficiency:
Bacteria-based systems: Escherichia coli (E. coli) is being tested for production of L1 proteins and L1 capsomeres (a cheaper alternative to full VLPs).
Transgenic Plants: Tobacco plants are being researched for plant-based VLP production.
Also!
Sources
Production of virus-like particles for vaccines, J Fuenmayor a,⁎, F Gòdia a, L Cervera b
Recombinant protein vaccines produced in insect cells Manon MJ Cox 1,⁎
L1 Recombinant Proteins of HPV Tested for Antibody Forming Using Sera of HPV Quadrivalent Vaccine
Part 3.5. How does it work in nature/biological systems?
+Describe how a single gene codes for multiple proteins at the transcriptional level.
A single gene can code for multiple proteins known as isoforms at the transcriptional level primarily through a mechanism called alternative splicing, along with alternative promoter usage and alternative polyadenylation. This process allows the 20,000-25,000 human genes to generate over 90,000 different proteins, greatly expanding the coding capacity of the [genome](https://pmc.ncbi.nlm.nih.gov/articles/PMC4360811/).
Here is a detailed description of how a single gene codes for multiple proteins at the transcriptional level:
Alternative Splicing (The Primary Mechanism). Alternative splicing occurs when the pre-mRNA (primary transcript) is processed, and different combinations of exons are joined together while introns are removed.
+Transcriptional Processing: The entire gene, including exons (coding regions) and introns (non-coding regions), is transcribed into pre-mRNA. +Splicing Variations: During maturation, splicing machinery (spliceosomes) can skip certain exons or retain specific introns, resulting in different mature mRNA molecules. +Protein Diversity: These varied mRNA molecules are translated into proteins with different, sometimes opposing, functions.
+Key Types of Alternative Splicing: -Cassette Exon (Exon Skipping): An exon may be included or excluded from the final mRNA (e.g., Exon 1-2-3 or 1-3-4). -Alternative 5’ or 3’ Splice Sites: Changes the length of the exon, affecting the coding sequence. -Intron Retention: An intron is kept in the final mRNA, usually leading to non-functional protein or degradation, but sometimes contributing to diversity.
-Mutually Exclusive Exons: Only one of two adjacent exons is retained.
Alternative Promoter Usage
+A single gene may have multiple promoters (transcription start sites). +Different Transcription Starts: The cell can choose to start transcription at a different location (e.g., a “fast” or “slow” promoter), resulting in different 5’ exons (first exons). +Functional Impact: This can produce N-terminal variants of a protein, which may have different localization or enzymatic properties.
Alternative Polyadenylation Alternative Termination. Genes can have multiple polyadenylation sites at the 3’ end.
+Different 3’ Ends: Transcription can end at different points, producing mRNAs with varying 3’ untranslated regions (UTRs) or different final exons. +Impact on Stability: This affects the length of the transcript and its stability, often altering the binding sites for miRNAs, thus regulating translation.
Trans-splicing
+A more unusual, yet significant, mechanism where exons from two completely different primary transcripts are spliced together to form a new, chimeric, or “fusion” mRNA molecule.
+Try aligning the DNA sequence, the transcribed RNA, and also the resulting translated Protein!!! See example below.
[Example shows the biomolecular flow in central dogma from DNA to RNA to Protein] Special note that all “T” were transcribed into “U” and that the 3-nt codon represents].
I will see if I have time later to do this!
Part 4: Prepare a Twist DNA Synthesis Order
Part 4.1. Create a Twist account and a Benchling account
Part 4.2. Build Your DNA Insert Sequence
For example, let’s make a sequence that will make E. coli glow fluorescent green under UV light by constitutively (always) expressing sfGFP (a green fluorescent protein).
- In Benchling, select New DNA/RNA sequence
I added my protein dna sequence of hpv16 l1 in jcat to get it optimised. Then I got the optimised codon result and added it here on the bases section!
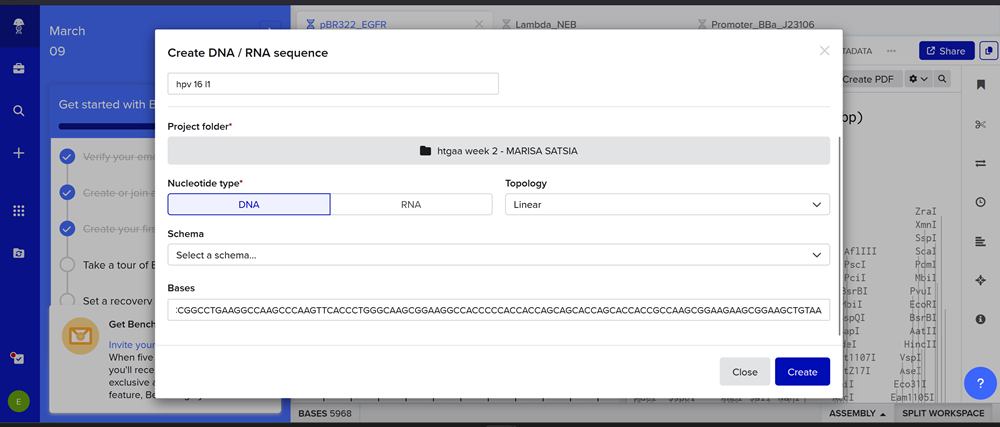

- Give your insert sequence a name and select DNA with a Linear topology (this is a linear sequence that will be inserted into a circular backbone vector of our choosing).



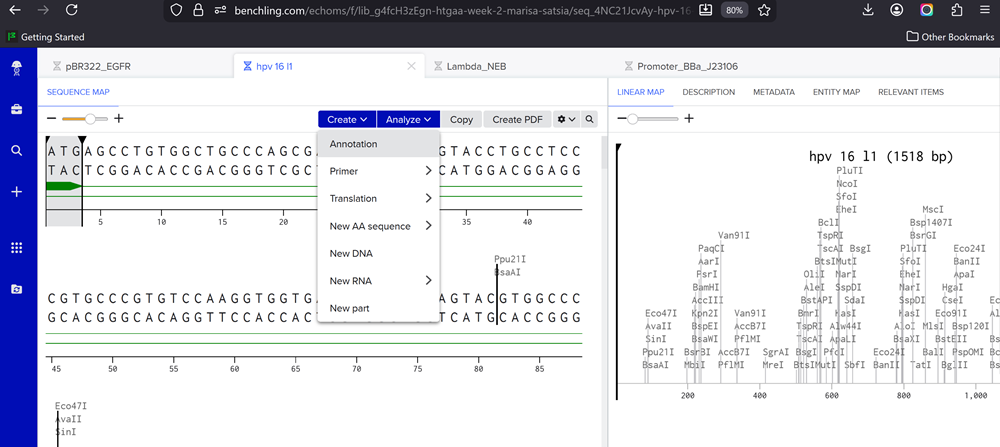
Here you can find my benchling page with the dna insert seq.
- Go through each piece of the given DNA sequences highlighted below (Promoter, RBS, Start Codon, Coding Sequence, His Tag, Stop Codon, Terminator) and paste the sequences into the Benchling file one after the other (replacing the coding sequence with your codon optimized DNA sequence of interest!). Each time you add a new piece of the sequence, make sure to annotate by right clicking over the sequence and creating an annotation that describes what each piece (e.g., Promoter, RBS, etc.) is (see image below).
I have not managed to find a way to make annotations even from the previous section so I will do some more troubleshooting!

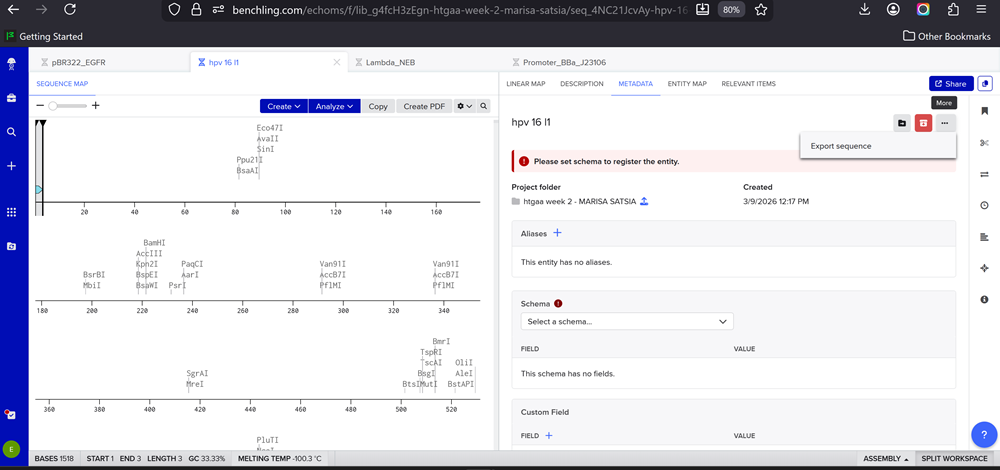
- Once you’ve completed this, click on Linear Map to preview the entire sequence. If you intend to have a TA review a sequence in the future, this is a good way to verify that all sections are annotated!
Still having issues since number 3 is still confusing me! I need someone to explain it to me so I can finish this part -.-
- This insert sequence you built is commonly referred to as an expression cassette in molecular biology (a sequence you can drop into any vector and it’ll perform its function). Go ahead and download the FASTA file for the sequence you made.

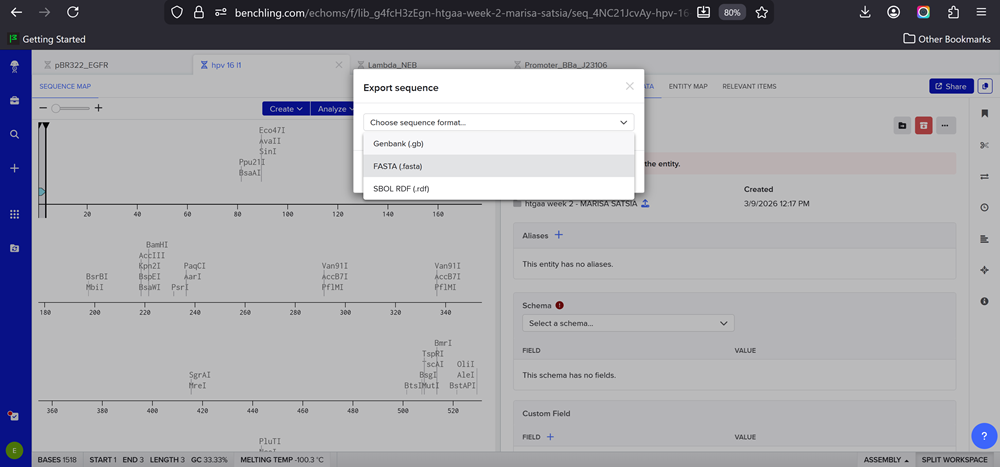

- It’s helpful to visualize DNA designs using SBOL Canvas and Synthetic Biology Open Language to convey your designs. Here’s an example of what you just annotated in Benchling:
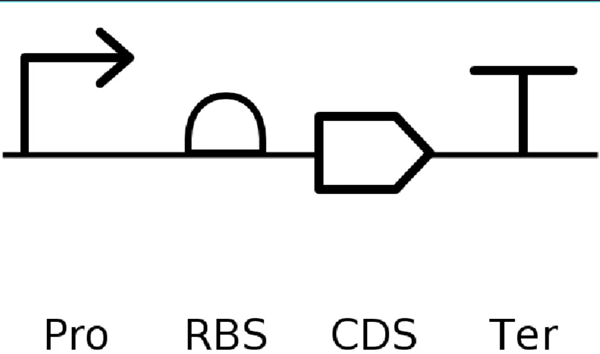

Part 4.3. On Twist, Select The “Genes” Option

Part 4.4. Select clonal genes option

Part 4.5. Import your sequence

Annnd I run into an issue!
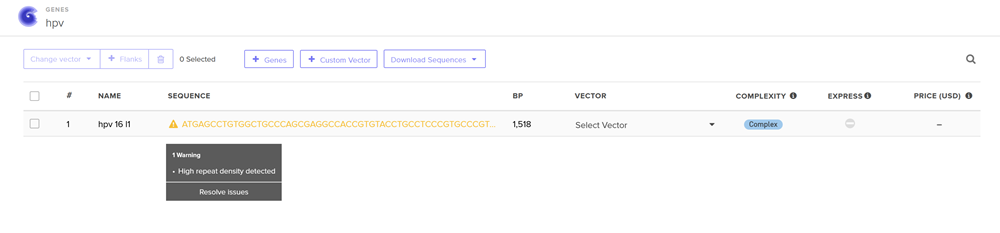

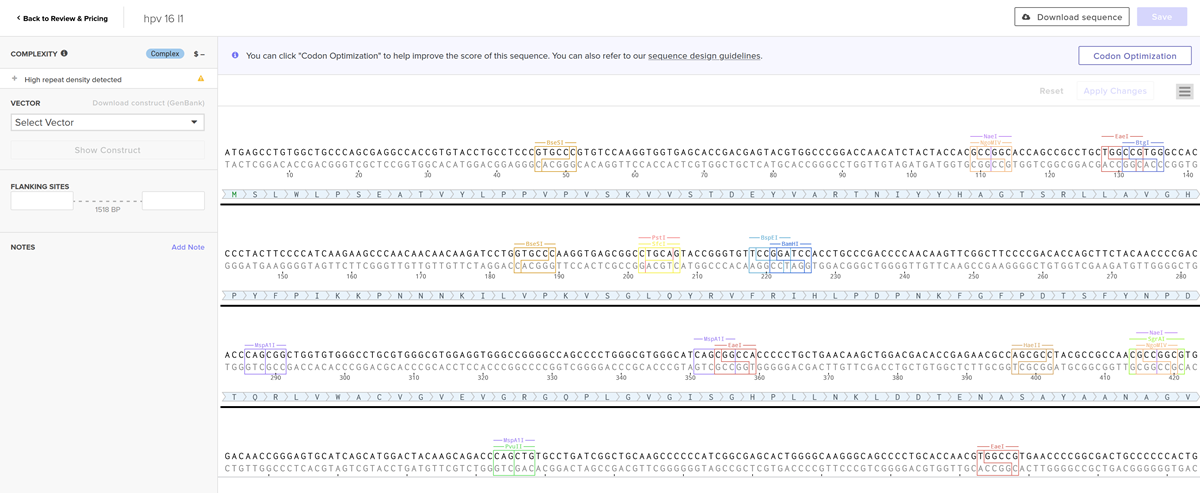

I clicked on codon optimization option on the top right! I think something went wrong from the time I got the nucleotide sequence of the hpv16 l1 protein so I am having a hard time to optimize in jcat and subsequenstly to continue this section on benchling and then twist. It is taking a while so I am gonna document how it went and continue some other time.
I optimised again in Twist and I proceeded.
Part 4.6. Choose Your Vector
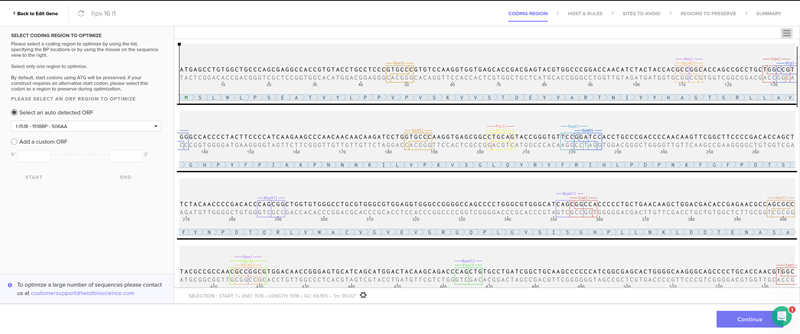





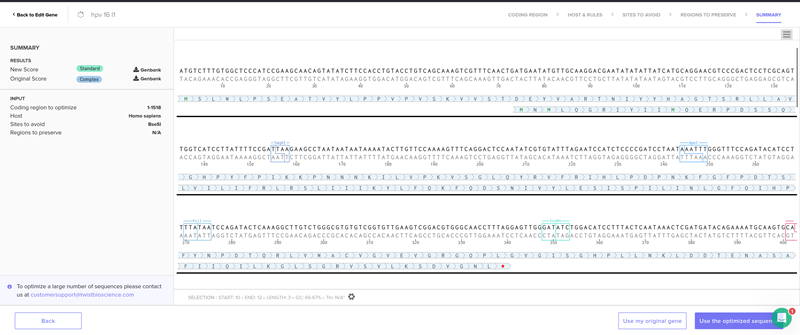

I chose the cloning vector.
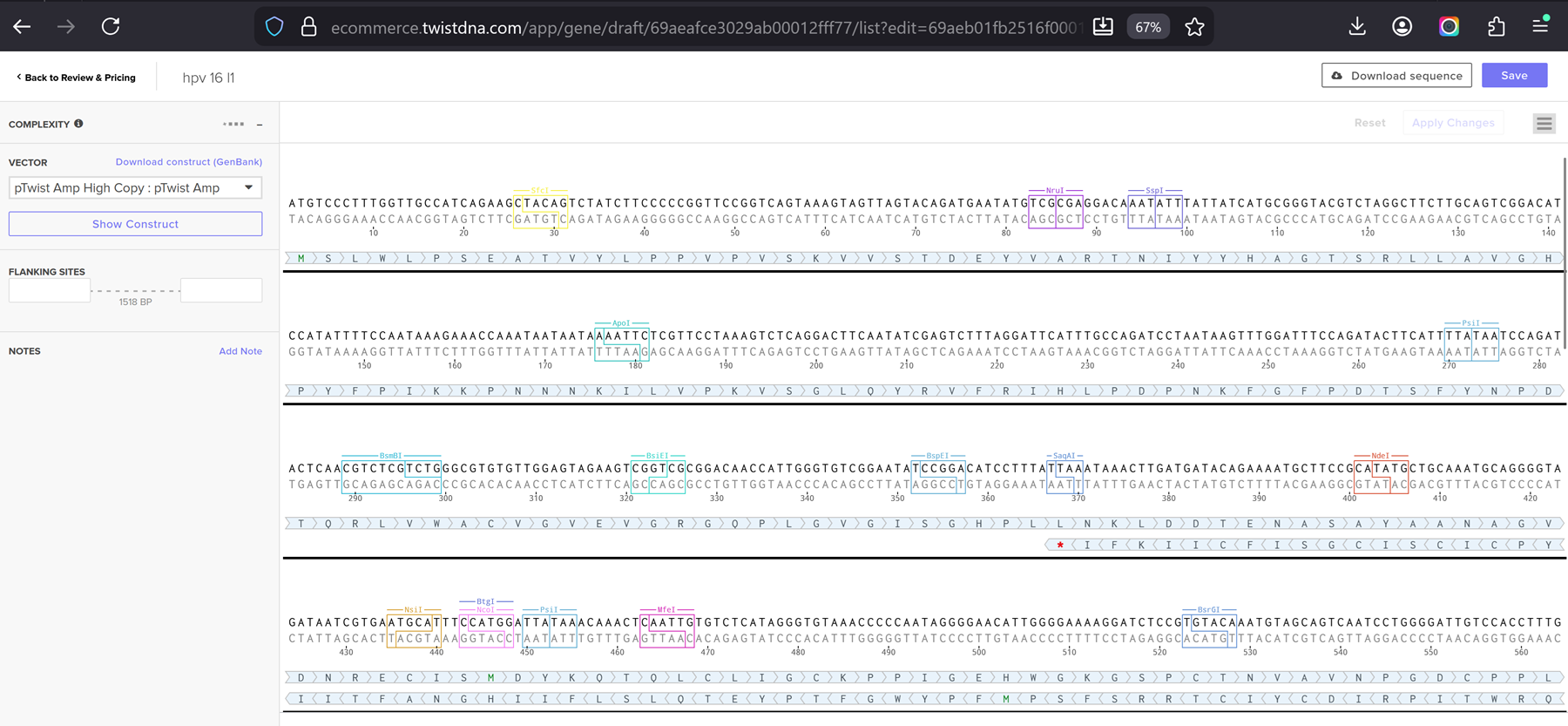

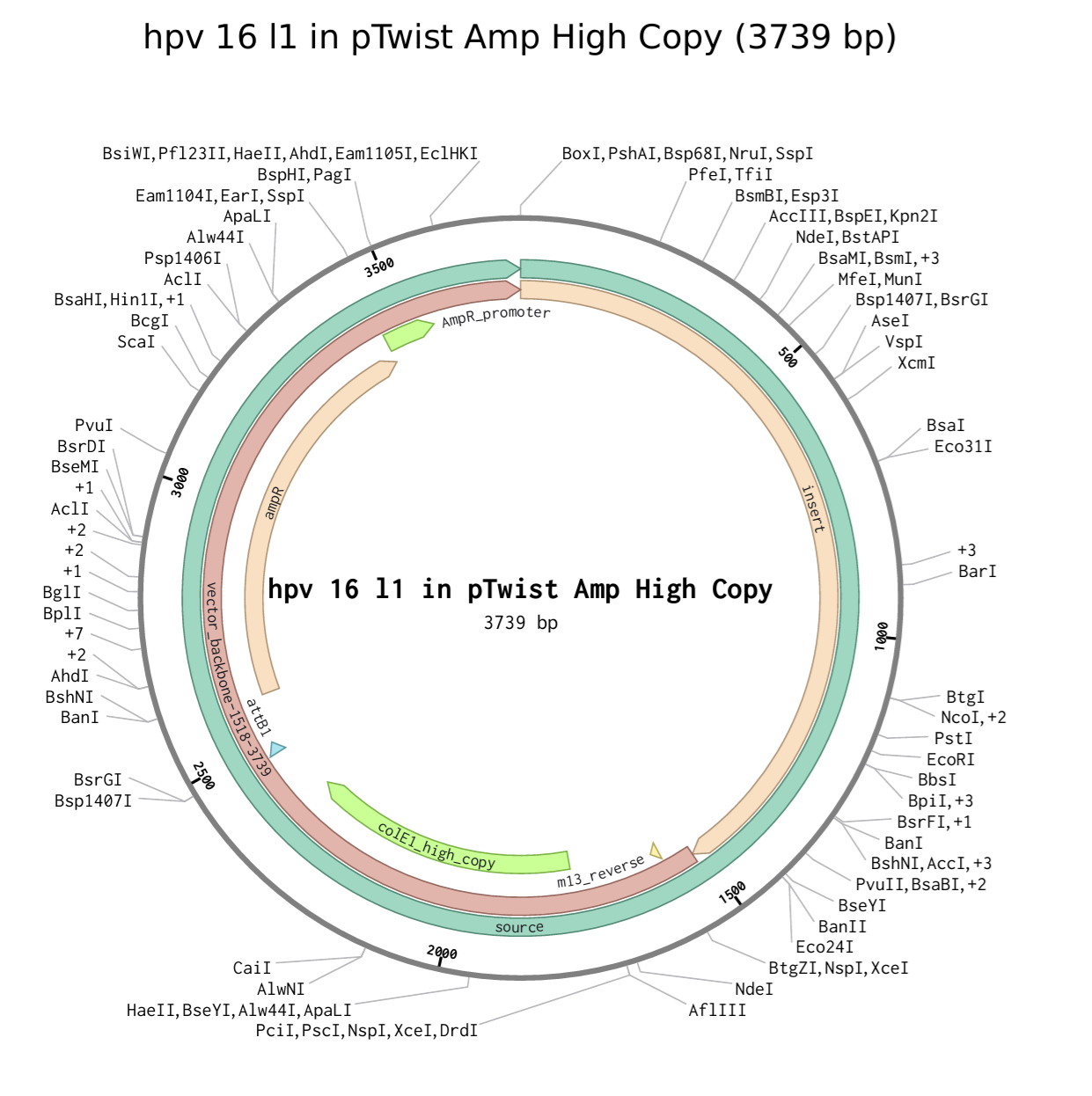
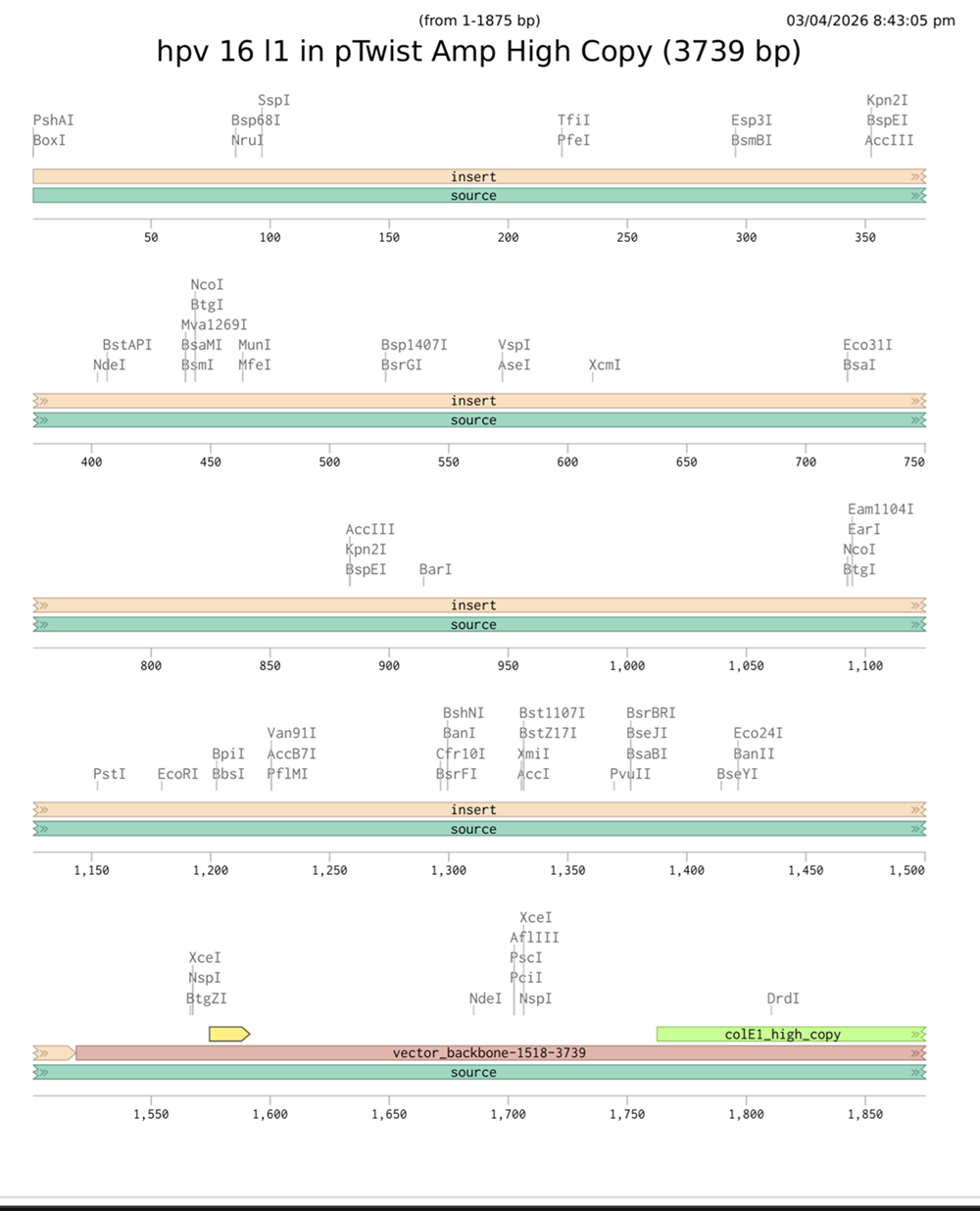
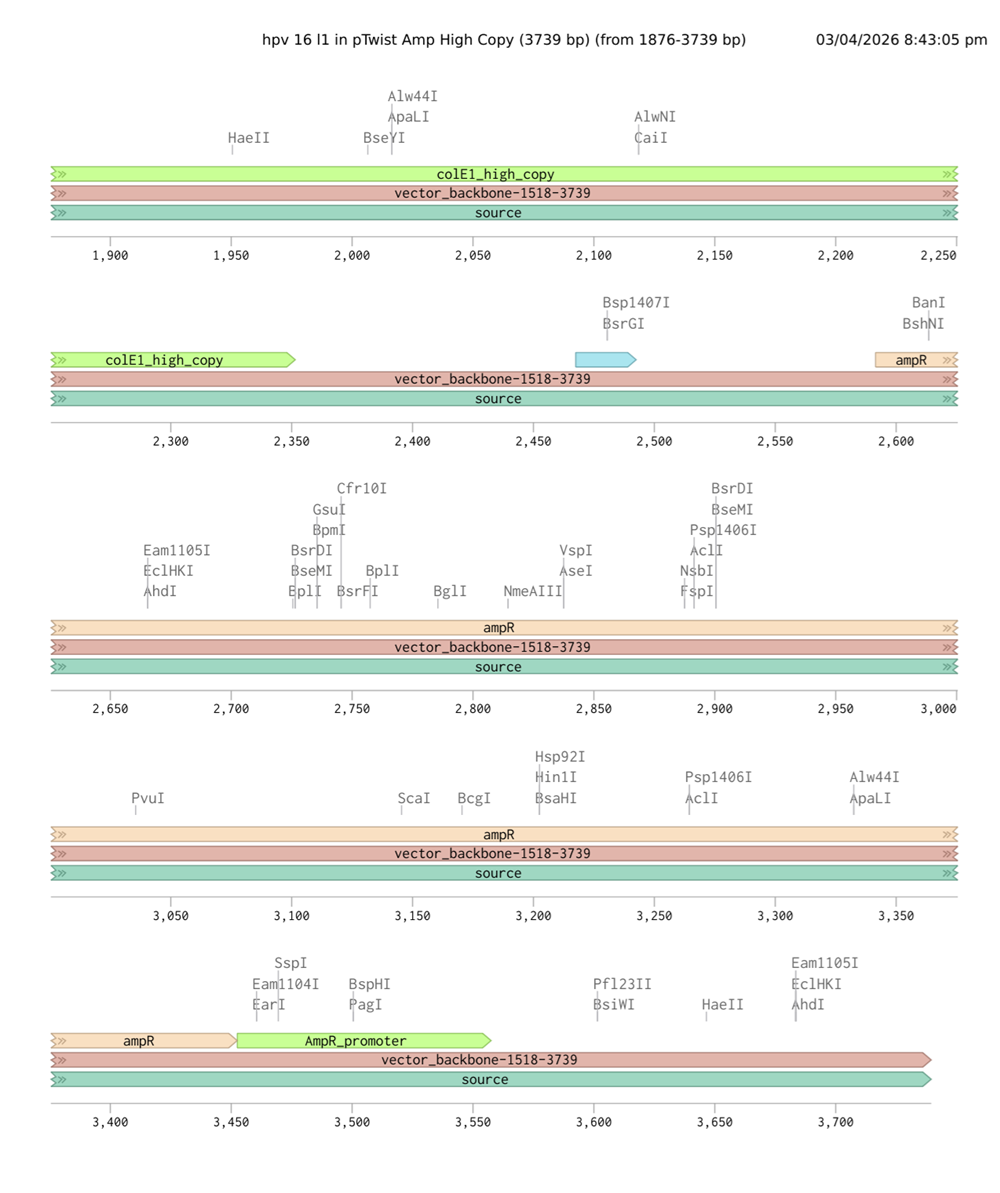
I had a look at the construct viever after I added the cloning vectors like pTwist Amp High Copy into my hpv l1 sequence.
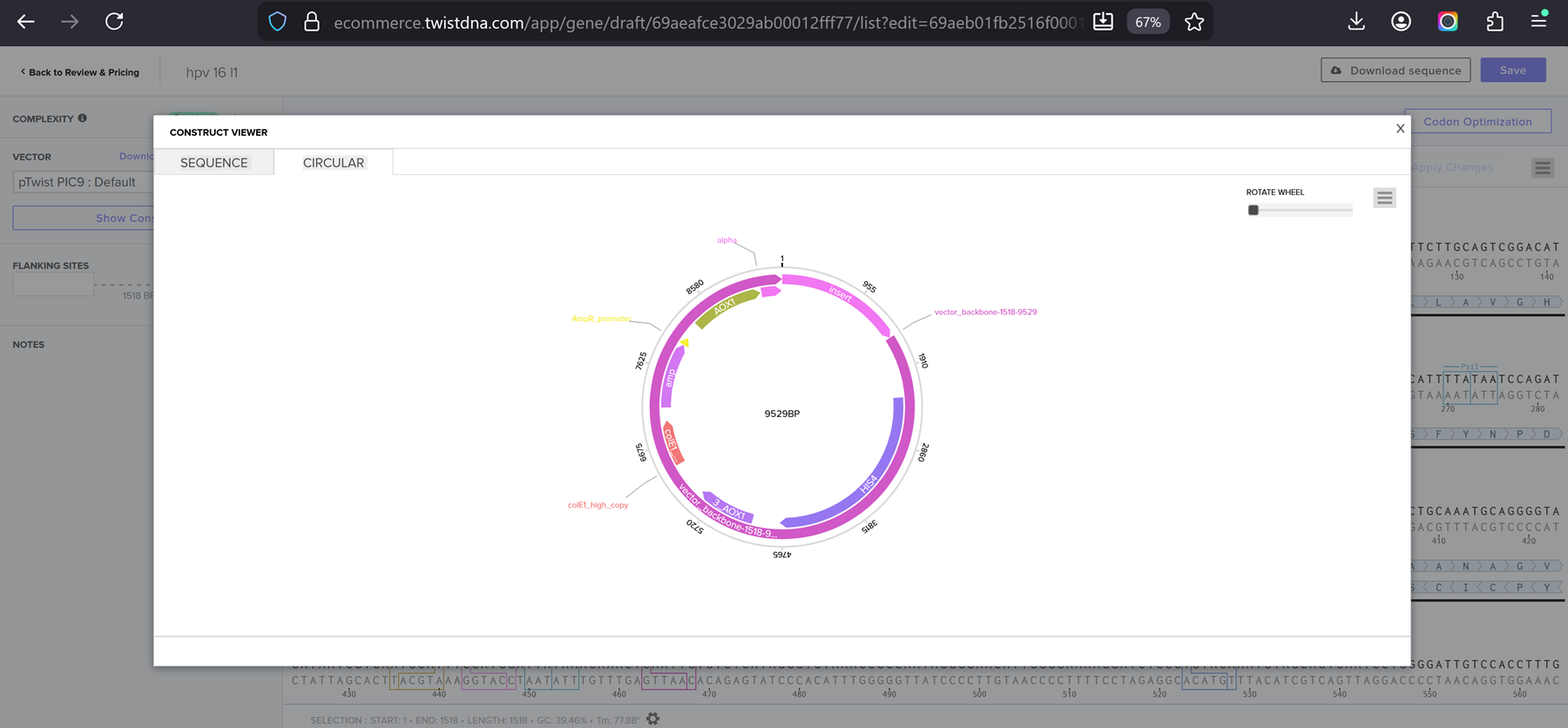


I clicked onto the cloning sequence and select download construct (GenBank) to get the full plasmid sequence.

AAAAAaaannnd back to my Benchling account. Inside of a folder, I clicked the import DNA/RNA sequence button and upload the GenBank file I just downloaded.



Part 5: Read, write, edit!🔮
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why? This could be DNA related to human health (e.g. genes related to disease research), environmental monitoring (e.g., sewage waste water, biodiversity analysis), and beyond (e.g. DNA data storage, biobank)?
I would like to read HPV16 AND/or HPV18. It is important to me because of personal reasons.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
Several advanced technologies are used to analyze HPV DNA and RNA sequences, ranging from established clinical screening methods to cutting-edge research tools for detecting viral integration. The primary techniques include Next-Generation Sequencing (NGS), PCR-based methods, and molecular hybridization.
Here is a breakdown of the technologies used for HPV DNA/RNA sequence analysis
Next-Generation Sequencing (NGS), NGS is used for high-throughput, comprehensive genomic analysis, including identifying multiple HPV subtypes, mutations, and integration sites.
+Nanopore Sequencing (Third-Generation): This technology is used for long-read sequencing, allowing for the characterization of complete HPV genomes and the identification of HPV integration into the host genome. It is particularly useful for identifying chimeric cellular–viral reads. +Illumina Sequencing: Often combined with hybrid capture for high-accuracy sequencing of full HPV genomes. +HPV-KITE: A specialized algorithm that uses k-mer data analysis for rapid HPV detection from NGS data.
Nucleic Acid Amplification & Detection (DNA/RNA)
+Real-Time PCR (qPCR): The most common method, using primers (e.g., L1, E6/E7) to amplify and quantify HPV DNA. Examples include Cobas HPV and BD Onclarity. +RT-PCR (Reverse Transcription PCR): Used specifically for detecting mRNA expression of E6 and E7 oncoproteins. +Transcription-Mediated Amplification (TMA): Used in the Aptima HPV Assay to detect E6/E7 mRNA for high-risk HPV. +Isothermal Amplification (IATs): Methods like Loop-Mediated Isothermal Amplification (LAMP) and Nucleic Acid Sequence-Based Amplification (NASBA) are used for rapid, isothermal detection without a thermocycler. +Droplet Digital PCR (ddPCR): Used for absolute quantification of HPV DNA/RNA with high sensitivity.
Signal Amplification & Hybridization
+Hybrid Capture (HC2): A signal amplification method that uses RNA probes to hybridize with HPV DNA, which is then captured and detected via chemiluminescence. +Invader Technology: A signal amplification method (used in Cervista tests) that uses special enzymes to cleave DNA, creating a fluorescent signal. +DNA Microarray/Chips: Technologies like Linear Array or PapilloCheck detect multiple HPV types by hybridizing amplified DNA to specific probes.
Summary of Technologies by Goal
Goal Technology Full Genome Integration Nanopore Sequencing, Illumina High-Risk DNA Screening Real-Time PCR (Cobas, Abbott), Hybrid Capture 2 (HC2) Active Infection (RNA) RT-PCR (Aptima, NASBA) Point-of-Care/Rapid LAMP, RPA, CRISPR-Cas12a
Also answer the following questions
+Is your method first-, second- or third-generation or other? How so?
+What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps.
+What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?
+What is the output of your chosen sequencing technology?
5.2 - DNA WRITE
(i) What DNA would you want to synthesize (e.g., write) and why? These could be individual genes, clusters of genes or genetic circuits, whole genomes, and beyond. As described in class thus far, applications could range from therapeutics and drug discovery (e.g., mRNA vaccines and therapies) to novel biomaterials (e.g. structural proteins), to sensors (e.g., genetic circuits for sensing and responding to inflammation, environmental stimuli, etc.), to art (DNA origamis). If possible, include the specific genetic sequence(s) of what you would like to synthesize! You will have the opportunity to actually have Twist synthesize these DNA constructs! :)
(ii) What technology or technologies would you use to perform this DNA synthesis and why? Also answer the following questions:
What are the essential steps of your chosen sequencing methods?
What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability?
5.3 - DNA EDIT
(i) What DNA would you want to edit and why? In class, George shared a variety of ways to edit the genes and genomes of humans and other organisms. Such DNA editing technologies have profound implications for human health, development, and even human longevity and human augmentation. DNA editing is also already commonly leveraged for flora and fauna, for example in nature conservation efforts, (animal/plant restoration, de-extinction), or in agriculture (e.g. plant breeding, nitrogen fixation). What kinds of edits might you want to make to DNA (e.g., human genomes and beyond) and why?
(ii) What technology or technologies would you use to perform these DNA edits and why?
Also answer the following questions:
- How does your technology of choice edit DNA? What are the essential steps?
- What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
- What are the limitations of your editing methods (if any) in terms of efficiency or precision?