Week 3 HW: LAB AUTOMATION


[E-INK] MICROFLUIDICS <3


I have actually been interested in microfluidics in a while because I am into inflatables and soft robotics since 2020. I started working with bodily fluids and liquids in 2023. I love this little sweat collection and analysis wearable microfluidic system device.
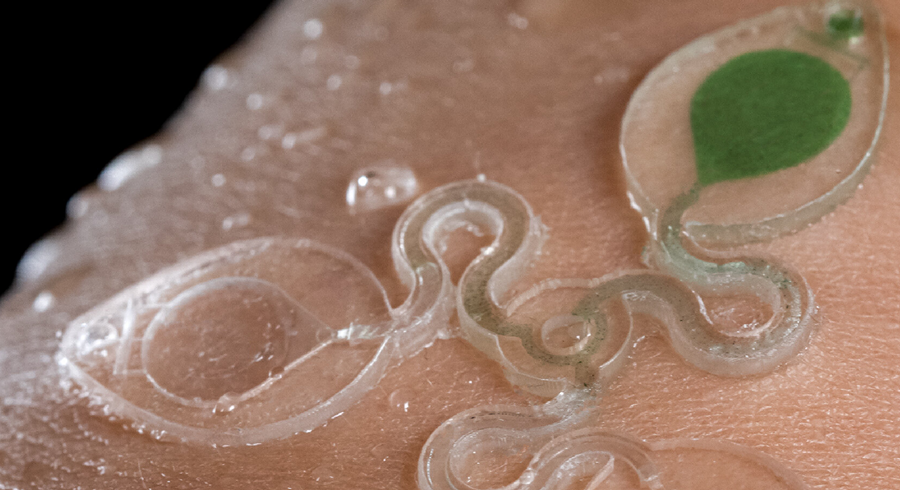

You can find another example here and the paper.
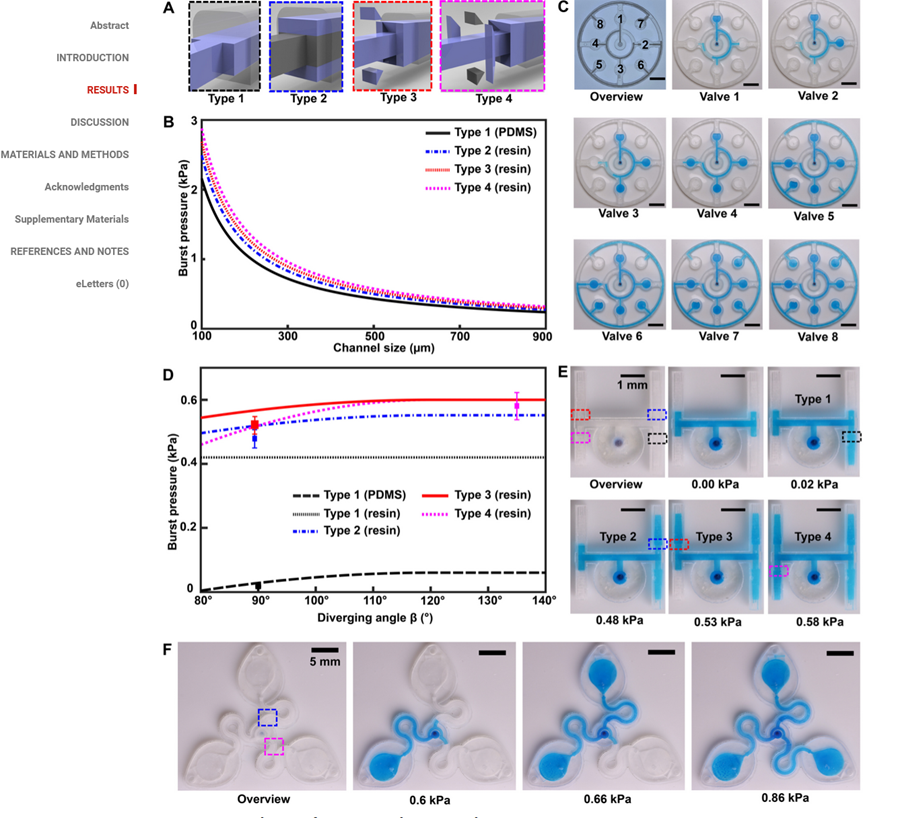

And another paper about a similar device.
Toehold switch biosensor
Apart from the cool robots for liquid handling in the lab I also really enjoyed the presentation on toehold switches/biosensors. I read some papers about the detection of HPV. I read some papers, have a look below!

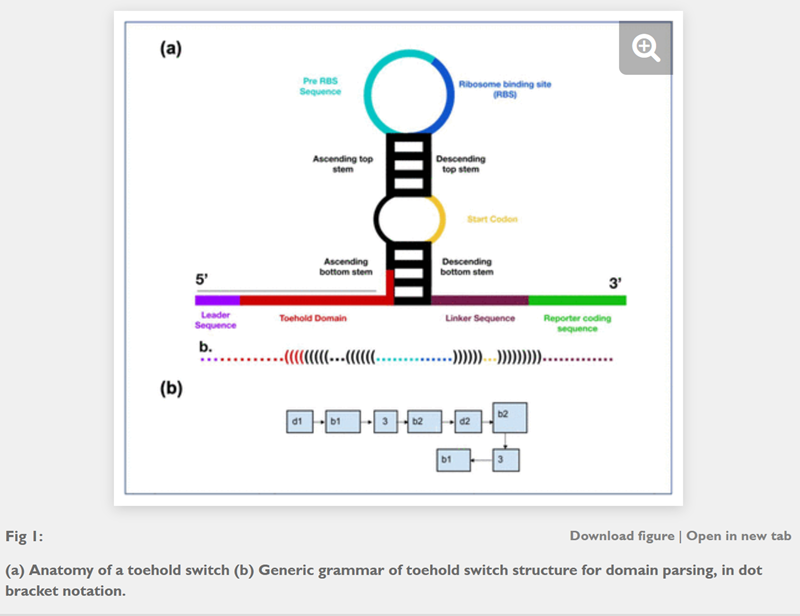

Rational design of toehold sequences


Another paper was about High-sensitivity electrochemical detection of HPV DNA via enzyme-amplified target-induced hairpin opening on a thermally controlled paper-based digital microfluidic platform. You can find it here. I have been interested in bioelectronics for a while. Furthermore, the developed platform was successfully evaluated for HPV16 DNA detection from clinical cervical swab samples without requiring direct target amplification.
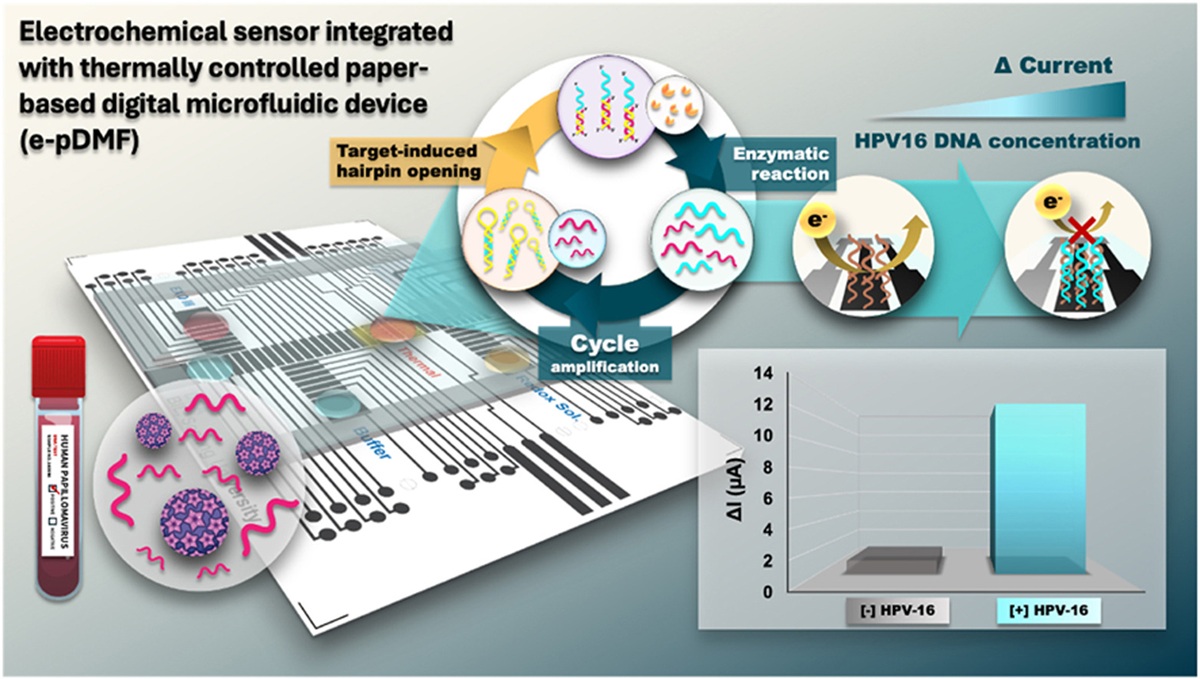

An electrochemical sensor integrated with a thermally controllable paper-based DMF (e-pDMF) device for target-induced hairpin opening with an enzyme-assisted signal amplification strategy (Scheme 1). This sensing platform is applied to detect HPV type 16 DNA (HPV16 DNA), a high-risk strain known to be a significant cause of cervical cancer. The e-pDMF device is designed to operate both transport and thermal features for precise droplet delivery and temperature control. The delivery mode enables efficient droplet manipulation and mixing, while the thermal zone on the device generates precise temperatures for optimal enzyme activity. Combining these functionalities allows for seamless operation, covering all steps from sample loading and mixing to signal amplification and electrochemical measurement. The HPV16 DNA opens the stem-loop structure of hairpin DNA (HP DNA) to form a duplex. Then, Exo III catalyzes the degradation of the duplex, releasing cleaved DNA and target DNA parts. The released target DNA part continues in cyclic enzymatic amplification, producing a large amount of cleaved DNA. This cleaved DNA is captured by a probe immobilized on the electrode surface. The decrease in current caused by electron transfer at the interface is measured using differential pulse voltammetry (DPV).
Week 03 HW
Assignment: Python Script for Opentrons Artwork — DUE BY YOUR LAB TIME!
- Your task this week is to Create a Python file to run on an Opentrons liquid handling robot. Review this week’s recitation and this week’s lab for details on the Opentrons and programming it.
- Generate an artistic design using the GUI at opentrons-art.rcdonovan.com.
- Using the coordinates from the GUI, follow the instructions in the HTGAA26 Opentrons Colab to write your own Python script which draws your design using the Opentrons.

You may use AI assistance for this coding — Google Gemini is integrated into Colab (see the stylized star bottom center); it will do a good job writing functional Python, while you probably need to take charge of the art concept.
If you’re a proficient programmer and you’d rather code something mathematical or algorithmic instead of using your GUI coordinates, you may do that instead. If you use AI to help complete this homework or lab, document how you used AI and which models made contributions.
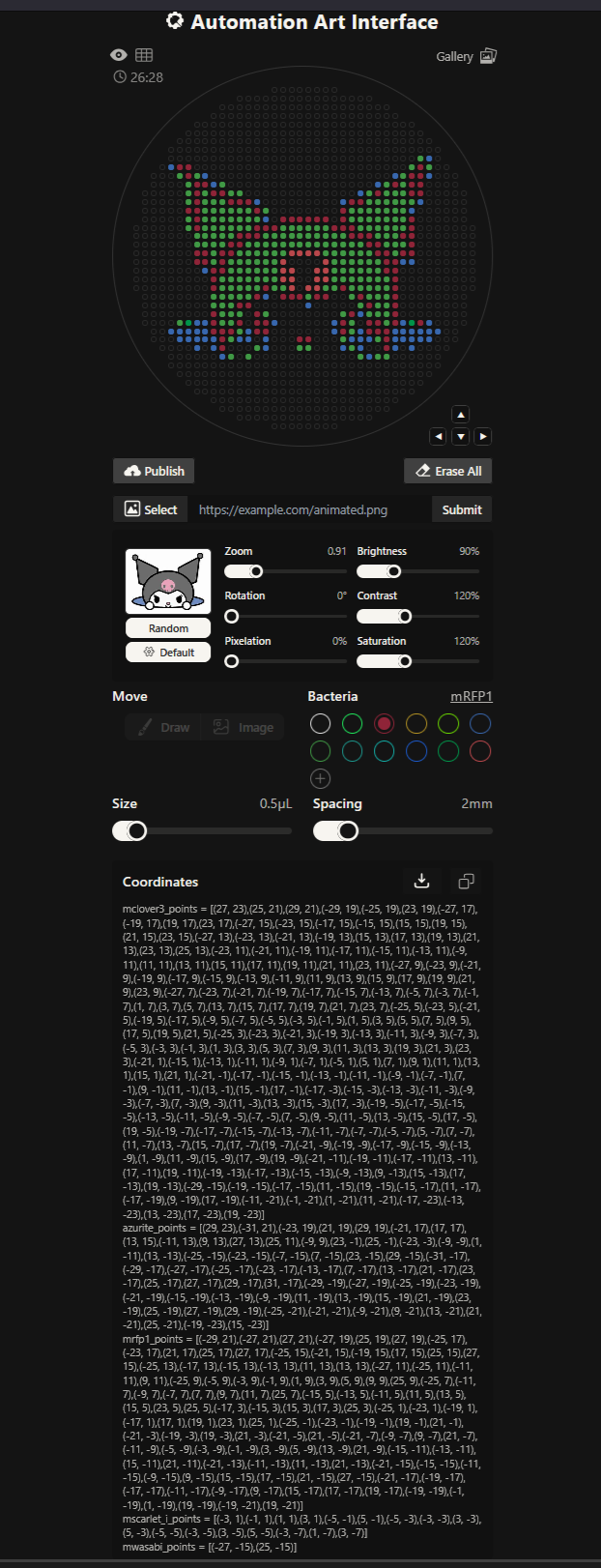

Here you can see my Kuromi design!
These are the coordinates Ronans website created!
mrfp1_points = [(-29, 21),(-27, 21),(27, 21),(-27, 19),(25, 19),(27, 19),(-25, 17),(-23, 17),(21, 17),(25, 17),(27, 17),(-25, 15),(-21, 15),(-19, 15),(17, 15),(25, 15),(27, 15),(-25, 13),(-17, 13),(-15, 13),(-13, 13),(11, 13),(13, 13),(-27, 11),(-25, 11),(-11, 11),(9, 11),(-25, 9),(-5, 9),(-3, 9),(-1, 9),(1, 9),(3, 9),(5, 9),(9, 9),(25, 9),(-25, 7),(-11, 7),(-9, 7),(-7, 7),(7, 7),(9, 7),(11, 7),(25, 7),(-15, 5),(-13, 5),(-11, 5),(11, 5),(13, 5),(15, 5),(23, 5),(25, 5),(-17, 3),(-15, 3),(15, 3),(17, 3),(25, 3),(-25, 1),(-23, 1),(-19, 1),(-17, 1),(17, 1),(19, 1),(23, 1),(25, 1),(-25, -1),(-23, -1),(-19, -1),(19, -1),(21, -1),(-21, -3),(-19, -3),(19, -3),(21, -3),(-21, -5),(21, -5),(-21, -7),(-9, -7),(9, -7),(21, -7),(-11, -9),(-5, -9),(-3, -9),(-1, -9),(3, -9),(5, -9),(13, -9),(21, -9),(-15, -11),(-13, -11),(15, -11),(21, -11),(-21, -13),(-11, -13),(11, -13),(21, -13),(-21, -15),(-15, -15),(-11, -15),(-9, -15),(9, -15),(15, -15),(17, -15),(21, -15),(27, -15),(-21, -17),(-19, -17),(-17, -17),(-11, -17),(-9, -17),(9, -17),(15, -17),(17, -17),(19, -17),(-19, -19),(-1, -19),(1, -19),(19, -19),(-19, -21),(19, -21)] mclover3_points = [(27, 23),(25, 21),(29, 21),(-29, 19),(-25, 19),(23, 19),(-27, 17),(-19, 17),(19, 17),(23, 17),(-27, 15),(-23, 15),(-17, 15),(-15, 15),(15, 15),(19, 15),(21, 15),(23, 15),(-27, 13),(-23, 13),(-21, 13),(-19, 13),(15, 13),(17, 13),(19, 13),(21, 13),(23, 13),(25, 13),(-23, 11),(-21, 11),(-19, 11),(-17, 11),(-15, 11),(-13, 11),(-9, 11),(11, 11),(13, 11),(15, 11),(17, 11),(19, 11),(21, 11),(23, 11),(-27, 9),(-23, 9),(-21, 9),(-19, 9),(-17, 9),(-15, 9),(-13, 9),(-11, 9),(11, 9),(13, 9),(15, 9),(17, 9),(19, 9),(21, 9),(23, 9),(-27, 7),(-23, 7),(-21, 7),(-19, 7),(-17, 7),(-15, 7),(-13, 7),(-5, 7),(-3, 7),(-1, 7),(1, 7),(3, 7),(5, 7),(13, 7),(15, 7),(17, 7),(19, 7),(21, 7),(23, 7),(-25, 5),(-23, 5),(-21, 5),(-19, 5),(-17, 5),(-9, 5),(-7, 5),(-5, 5),(-3, 5),(-1, 5),(1, 5),(3, 5),(5, 5),(7, 5),(9, 5),(17, 5),(19, 5),(21, 5),(-25, 3),(-23, 3),(-21, 3),(-19, 3),(-13, 3),(-11, 3),(-9, 3),(-7, 3),(-5, 3),(-3, 3),(-1, 3),(1, 3),(3, 3),(5, 3),(7, 3),(9, 3),(11, 3),(13, 3),(19, 3),(21, 3),(23, 3),(-21, 1),(-15, 1),(-13, 1),(-11, 1),(-9, 1),(-7, 1),(-5, 1),(5, 1),(7, 1),(9, 1),(11, 1),(13, 1),(15, 1),(21, 1),(-21, -1),(-17, -1),(-15, -1),(-13, -1),(-11, -1),(-9, -1),(-7, -1),(7, -1),(9, -1),(11, -1),(13, -1),(15, -1),(17, -1),(-17, -3),(-15, -3),(-13, -3),(-11, -3),(-9, -3),(-7, -3),(7, -3),(9, -3),(11, -3),(13, -3),(15, -3),(17, -3),(-19, -5),(-17, -5),(-15, -5),(-13, -5),(-11, -5),(-9, -5),(-7, -5),(7, -5),(9, -5),(11, -5),(13, -5),(15, -5),(17, -5),(19, -5),(-19, -7),(-17, -7),(-15, -7),(-13, -7),(-11, -7),(-7, -7),(-5, -7),(5, -7),(7, -7),(11, -7),(13, -7),(15, -7),(17, -7),(19, -7),(-21, -9),(-19, -9),(-17, -9),(-15, -9),(-13, -9),(1, -9),(11, -9),(15, -9),(17, -9),(19, -9),(-21, -11),(-19, -11),(-17, -11),(13, -11),(17, -11),(19, -11),(-19, -13),(-17, -13),(-15, -13),(-9, -13),(9, -13),(15, -13),(17, -13),(19, -13),(-29, -15),(-19, -15),(-17, -15),(11, -15),(19, -15),(-15, -17),(11, -17),(-17, -19),(9, -19),(17, -19),(-11, -21),(-1, -21),(1, -21),(11, -21),(-17, -23),(-13, -23),(13, -23),(17, -23),(19, -23)] mscarlet_i_points = [(-3, 1),(-1, 1),(1, 1),(3, 1),(-5, -1),(5, -1),(-5, -3),(-3, -3),(3, -3),(5, -3),(-5, -5),(-3, -5),(3, -5),(5, -5),(-3, -7),(1, -7),(3, -7)] azurite_points = [(29, 23),(-31, 21),(-23, 19),(21, 19),(29, 19),(-21, 17),(17, 17),(13, 15),(-11, 13),(9, 13),(27, 13),(25, 11),(-9, 9),(23, -1),(25, -1),(-23, -3),(-9, -9),(1, -11),(13, -13),(-25, -15),(-23, -15),(-7, -15),(7, -15),(23, -15),(29, -15),(-31, -17),(-29, -17),(-27, -17),(-25, -17),(-23, -17),(-13, -17),(7, -17),(13, -17),(21, -17),(23, -17),(25, -17),(27, -17),(29, -17),(31, -17),(-29, -19),(-27, -19),(-25, -19),(-23, -19),(-21, -19),(-15, -19),(-13, -19),(-9, -19),(11, -19),(13, -19),(15, -19),(21, -19),(23, -19),(25, -19),(27, -19),(29, -19),(-25, -21),(-21, -21),(-9, -21),(9, -21),(13, -21),(21, -21),(25, -21),(-19, -23),(15, -23)] mwasabi_points = [(-27, -15),(25, -15)]
I also downloaded the 96 well plate python code from the website and here is a screenshot.
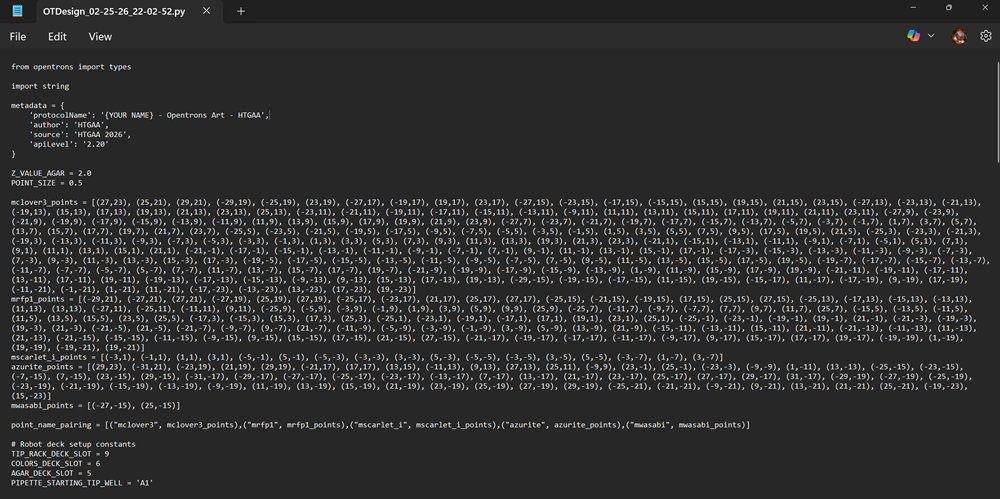

If you want copy and paste my code!
>from opentrons import types
import string
metadata = {
'protocolName': '{YOUR NAME} - Opentrons Art - HTGAA',
'author': 'HTGAA',
'source': 'HTGAA 2026',
'apiLevel': '2.20'
}
Z_VALUE_AGAR = 2.0
POINT_SIZE = 0.5
mclover3_points = [(27,23), (25,21), (29,21), (-29,19), (-25,19), (23,19), (-27,17), (-19,17), (19,17), (23,17), (-27,15), (-23,15), (-17,15), (-15,15), (15,15), (19,15), (21,15), (23,15), (-27,13), (-23,13), (-21,13), (-19,13), (15,13), (17,13), (19,13), (21,13), (23,13), (25,13), (-23,11), (-21,11), (-19,11), (-17,11), (-15,11), (-13,11), (-9,11), (11,11), (13,11), (15,11), (17,11), (19,11), (21,11), (23,11), (-27,9), (-23,9), (-21,9), (-19,9), (-17,9), (-15,9), (-13,9), (-11,9), (11,9), (13,9), (15,9), (17,9), (19,9), (21,9), (23,9), (-27,7), (-23,7), (-21,7), (-19,7), (-17,7), (-15,7), (-13,7), (-5,7), (-3,7), (-1,7), (1,7), (3,7), (5,7), (13,7), (15,7), (17,7), (19,7), (21,7), (23,7), (-25,5), (-23,5), (-21,5), (-19,5), (-17,5), (-9,5), (-7,5), (-5,5), (-3,5), (-1,5), (1,5), (3,5), (5,5), (7,5), (9,5), (17,5), (19,5), (21,5), (-25,3), (-23,3), (-21,3), (-19,3), (-13,3), (-11,3), (-9,3), (-7,3), (-5,3), (-3,3), (-1,3), (1,3), (3,3), (5,3), (7,3), (9,3), (11,3), (13,3), (19,3), (21,3), (23,3), (-21,1), (-15,1), (-13,1), (-11,1), (-9,1), (-7,1), (-5,1), (5,1), (7,1), (9,1), (11,1), (13,1), (15,1), (21,1), (-21,-1), (-17,-1), (-15,-1), (-13,-1), (-11,-1), (-9,-1), (-7,-1), (7,-1), (9,-1), (11,-1), (13,-1), (15,-1), (17,-1), (-17,-3), (-15,-3), (-13,-3), (-11,-3), (-9,-3), (-7,-3), (7,-3), (9,-3), (11,-3), (13,-3), (15,-3), (17,-3), (-19,-5), (-17,-5), (-15,-5), (-13,-5), (-11,-5), (-9,-5), (-7,-5), (7,-5), (9,-5), (11,-5), (13,-5), (15,-5), (17,-5), (19,-5), (-19,-7), (-17,-7), (-15,-7), (-13,-7), (-11,-7), (-7,-7), (-5,-7), (5,-7), (7,-7), (11,-7), (13,-7), (15,-7), (17,-7), (19,-7), (-21,-9), (-19,-9), (-17,-9), (-15,-9), (-13,-9), (1,-9), (11,-9), (15,-9), (17,-9), (19,-9), (-21,-11), (-19,-11), (-17,-11), (13,-11), (17,-11), (19,-11), (-19,-13), (-17,-13), (-15,-13), (-9,-13), (9,-13), (15,-13), (17,-13), (19,-13), (-29,-15), (-19,-15), (-17,-15), (11,-15), (19,-15), (-15,-17), (11,-17), (-17,-19), (9,-19), (17,-19), (-11,-21), (-1,-21), (1,-21), (11,-21), (-17,-23), (-13,-23), (13,-23), (17,-23), (19,-23)]
mrfp1_points = [(-29,21), (-27,21), (27,21), (-27,19), (25,19), (27,19), (-25,17), (-23,17), (21,17), (25,17), (27,17), (-25,15), (-21,15), (-19,15), (17,15), (25,15), (27,15), (-25,13), (-17,13), (-15,13), (-13,13), (11,13), (13,13), (-27,11), (-25,11), (-11,11), (9,11), (-25,9), (-5,9), (-3,9), (-1,9), (1,9), (3,9), (5,9), (9,9), (25,9), (-25,7), (-11,7), (-9,7), (-7,7), (7,7), (9,7), (11,7), (25,7), (-15,5), (-13,5), (-11,5), (11,5), (13,5), (15,5), (23,5), (25,5), (-17,3), (-15,3), (15,3), (17,3), (25,3), (-25,1), (-23,1), (-19,1), (-17,1), (17,1), (19,1), (23,1), (25,1), (-25,-1), (-23,-1), (-19,-1), (19,-1), (21,-1), (-21,-3), (-19,-3), (19,-3), (21,-3), (-21,-5), (21,-5), (-21,-7), (-9,-7), (9,-7), (21,-7), (-11,-9), (-5,-9), (-3,-9), (-1,-9), (3,-9), (5,-9), (13,-9), (21,-9), (-15,-11), (-13,-11), (15,-11), (21,-11), (-21,-13), (-11,-13), (11,-13), (21,-13), (-21,-15), (-15,-15), (-11,-15), (-9,-15), (9,-15), (15,-15), (17,-15), (21,-15), (27,-15), (-21,-17), (-19,-17), (-17,-17), (-11,-17), (-9,-17), (9,-17), (15,-17), (17,-17), (19,-17), (-19,-19), (1,-19), (19,-19), (-19,-21), (19,-21)]
mscarlet_i_points = [(-3,1), (-1,1), (1,1), (3,1), (-5,-1), (5,-1), (-5,-3), (-3,-3), (3,-3), (5,-3), (-5,-5), (-3,-5), (3,-5), (5,-5), (-3,-7), (1,-7), (3,-7)]
azurite_points = [(29,23), (-31,21), (-23,19), (21,19), (29,19), (-21,17), (17,17), (13,15), (-11,13), (9,13), (27,13), (25,11), (-9,9), (23,-1), (25,-1), (-23,-3), (-9,-9), (1,-11), (13,-13), (-25,-15), (-23,-15), (-7,-15), (7,-15), (23,-15), (29,-15), (-31,-17), (-29,-17), (-27,-17), (-25,-17), (-23,-17), (-13,-17), (7,-17), (13,-17), (21,-17), (23,-17), (25,-17), (27,-17), (29,-17), (31,-17), (-29,-19), (-27,-19), (-25,-19), (-23,-19), (-21,-19), (-15,-19), (-13,-19), (-9,-19), (11,-19), (13,-19), (15,-19), (21,-19), (23,-19), (25,-19), (27,-19), (29,-19), (-25,-21), (-21,-21), (-9,-21), (9,-21), (13,-21), (21,-21), (25,-21), (-19,-23), (15,-23)]
mwasabi_points = [(-27,-15), (25,-15)]
point_name_pairing = [("mclover3", mclover3_points),("mrfp1", mrfp1_points),("mscarlet_i", mscarlet_i_points),("azurite", azurite_points),("mwasabi", mwasabi_points)]
# Robot deck setup constants
TIP_RACK_DECK_SLOT = 9
COLORS_DECK_SLOT = 6
AGAR_DECK_SLOT = 5
PIPETTE_STARTING_TIP_WELL = 'A1'
# Place the PCR tubes in this order
well_colors = {
'A1': 'sfGFP',
'A2': 'mRFP1',
'A3': 'mKO2',
'A4': 'Venus',
'A5': 'mKate2_TF',
'A6': 'Azurite',
'A7': 'mCerulean3',
'A8': 'mClover3',
'A9': 'mJuniper',
'A10': 'mTurquoise2',
'A11': 'mBanana',
'A12': 'mPlum',
'B1': 'Electra2',
'B2': 'mWasabi',
'B3': 'mScarlet_I',
'B4': 'mPapaya',
'B5': 'eqFP578',
'B6': 'tdTomato',
'B7': 'DsRed',
'B8': 'mKate2',
'B9': 'EGFP',
'B10': 'mRuby2',
'B11': 'TagBFP',
'B12': 'mChartreuse_TF',
'C1': 'mLychee_TF',
'C2': 'mTagBFP2',
'C3': 'mEGFP',
'C4': 'mNeonGreen',
'C5': 'mAzamiGreen',
'C6': 'mWatermelon',
'C7': 'avGFP',
'C8': 'mCitrine',
'C9': 'mVenus',
'C10': 'mCherry',
'C11': 'mHoneydew',
'C12': 'TagRFP',
'D1': 'mTFP1',
'D2': 'Ultramarine',
'D3': 'ZsGreen1',
'D4': 'mMiCy',
'D5': 'mStayGold2',
'D6': 'PA_GFP'
}
volume_used = {
'mclover3': 0,
'mrfp1': 0,
'mscarlet_i': 0,
'azurite': 0,
'mwasabi': 0
}
def update_volume_remaining(current_color, quantity_to_aspirate):
rows = string.ascii_uppercase
for well, color in list(well_colors.items()):
if color == current_color:
if (volume_used[current_color] + quantity_to_aspirate) > 250:
# Move to next well horizontally by advancing row letter, keeping column number
row = well[0]
col = well[1:]
# Find next row letter
next_row = rows[rows.index(row) + 1]
next_well = f"{next_row}{col}"
del well_colors[well]
well_colors[next_well] = current_color
volume_used[current_color] = quantity_to_aspirate
else:
volume_used[current_color] += quantity_to_aspirate
break
def run(protocol):
# Load labware, modules and pipettes
protocol.home()
# Tips
tips_20ul = protocol.load_labware('opentrons_96_tiprack_20ul', TIP_RACK_DECK_SLOT, 'Opentrons 20uL Tips')
# Pipettes
pipette_20ul = protocol.load_instrument("p20_single_gen2", "right", [tips_20ul])
# Deep Well Plate
temperature_plate = protocol.load_labware('nest_96_wellplate_2ml_deep', 6)
# Agar Plate
agar_plate = protocol.load_labware('htgaa_agar_plate', AGAR_DECK_SLOT, 'Agar Plate')
agar_plate.set_offset(x=0.00, y=0.00, z=Z_VALUE_AGAR)
# Get the top-center of the plate, make sure the plate was calibrated before running this
center_location = agar_plate['A1'].top()
pipette_20ul.starting_tip = tips_20ul.well(PIPETTE_STARTING_TIP_WELL)
# Helper function (dispensing)
def dispense_and_jog(pipette, volume, location):
assert(isinstance(volume, (int, float)))
# Go above the location
above_location = location.move(types.Point(z=location.point.z + 2))
pipette.move_to(above_location)
# Go downwards and dispense
pipette.dispense(volume, location)
# Go upwards to avoid smearing
pipette.move_to(above_location)
# Helper function (color location)
def location_of_color(color_string):
for well,color in well_colors.items():
if color.lower() == color_string.lower():
return temperature_plate[well]
raise ValueError(f"No well found with color {color_string}")
# Print pattern by iterating over lists
for i, (current_color, point_list) in enumerate(point_name_pairing):
# Skip the rest of the loop if the list is empty
if not point_list:
continue
# Get the tip for this run, set the bacteria color, and the aspirate bacteria of choice
pipette_20ul.pick_up_tip()
max_aspirate = int(18 // POINT_SIZE) * POINT_SIZE
quantity_to_aspirate = min(len(point_list)*POINT_SIZE, max_aspirate)
update_volume_remaining(current_color, quantity_to_aspirate)
pipette_20ul.aspirate(quantity_to_aspirate, location_of_color(current_color))
# Iterate over the current points list and dispense them, refilling along the way
for i in range(len(point_list)):
x, y = point_list[i]
adjusted_location = center_location.move(types.Point(x, y))
dispense_and_jog(pipette_20ul, POINT_SIZE, adjusted_location)
if pipette_20ul.current_volume == 0 and len(point_list[i+1:]) > 0:
quantity_to_aspirate = min(len(point_list[i:])*POINT_SIZE, max_aspirate)
update_volume_remaining(current_color, quantity_to_aspirate)
pipette_20ul.aspirate(quantity_to_aspirate, location_of_color(current_color))
# Drop tip between each color
pipette_20ul.drop_tip()
Unfortunately I am having issues on google colab and cannot run the simulation. Find my colab here. I will try again. I keep following the errors but i am a bit lost.
More designs I made!


our grid plate looks amazing on ronans website!
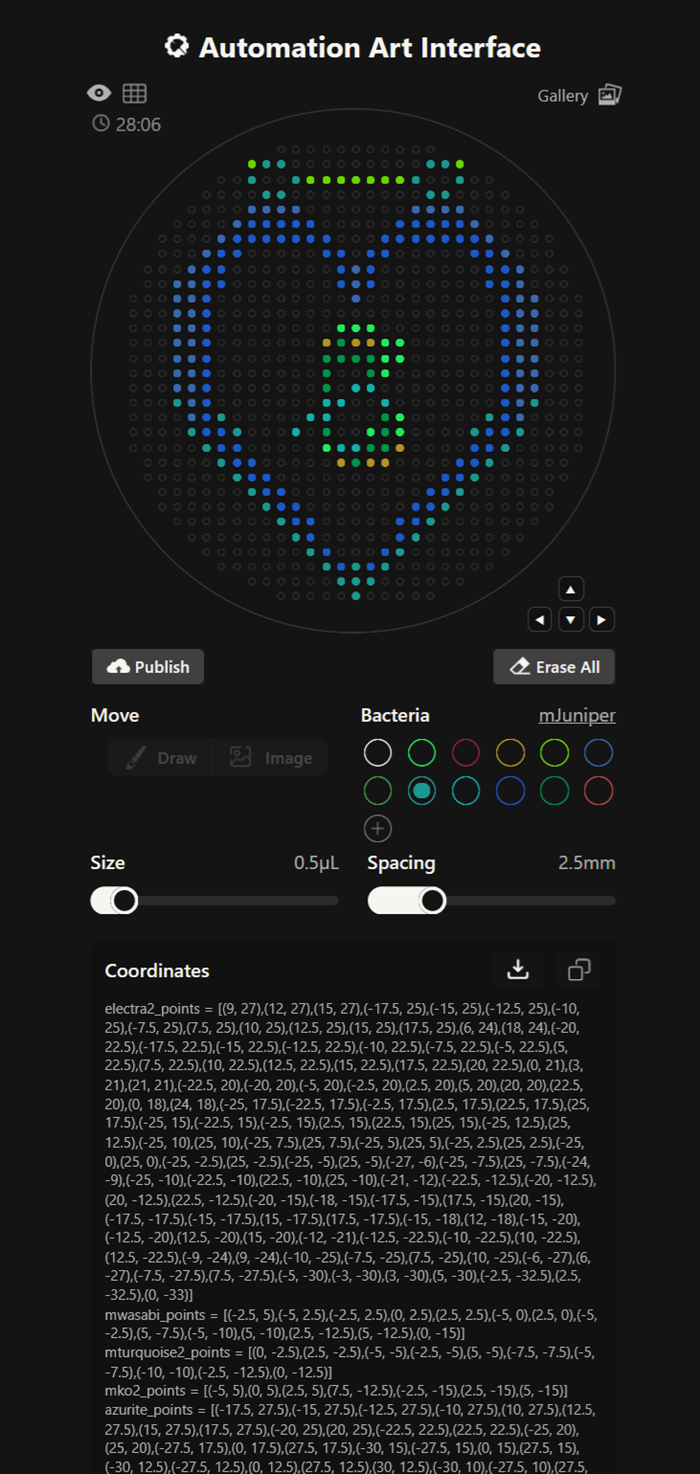

I made this for a dear friend <3 You can find it here.


The grid looks amazing!!!! I also imported a png picture of a cute rainbow i found online and then added more points or edited out existing ones on ronans website!
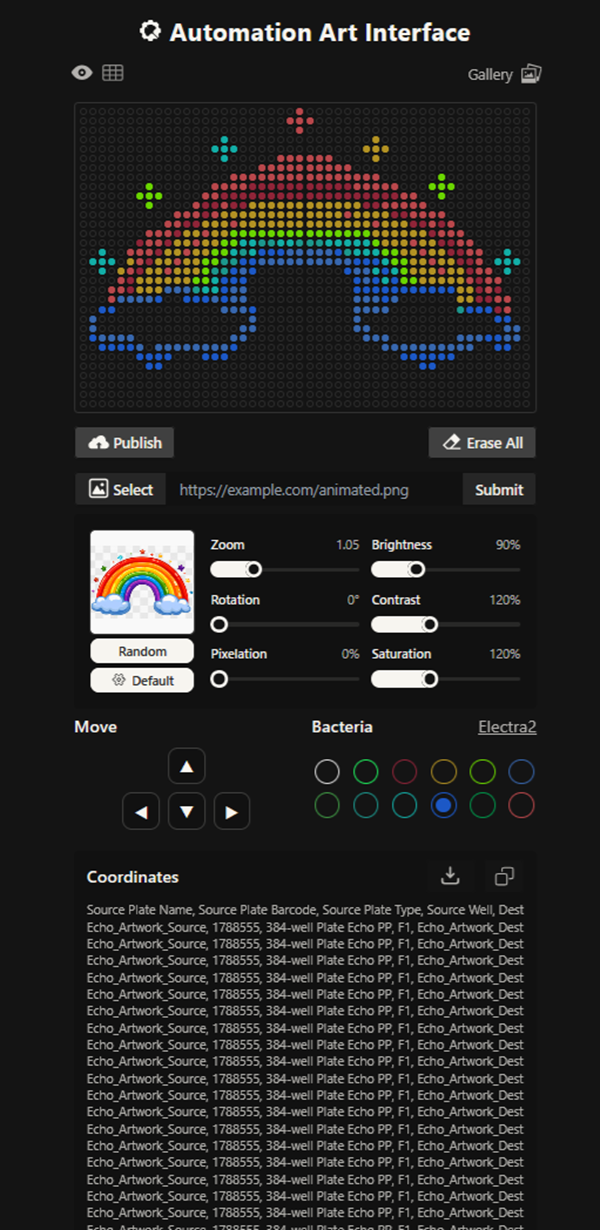

Here you can find my design!
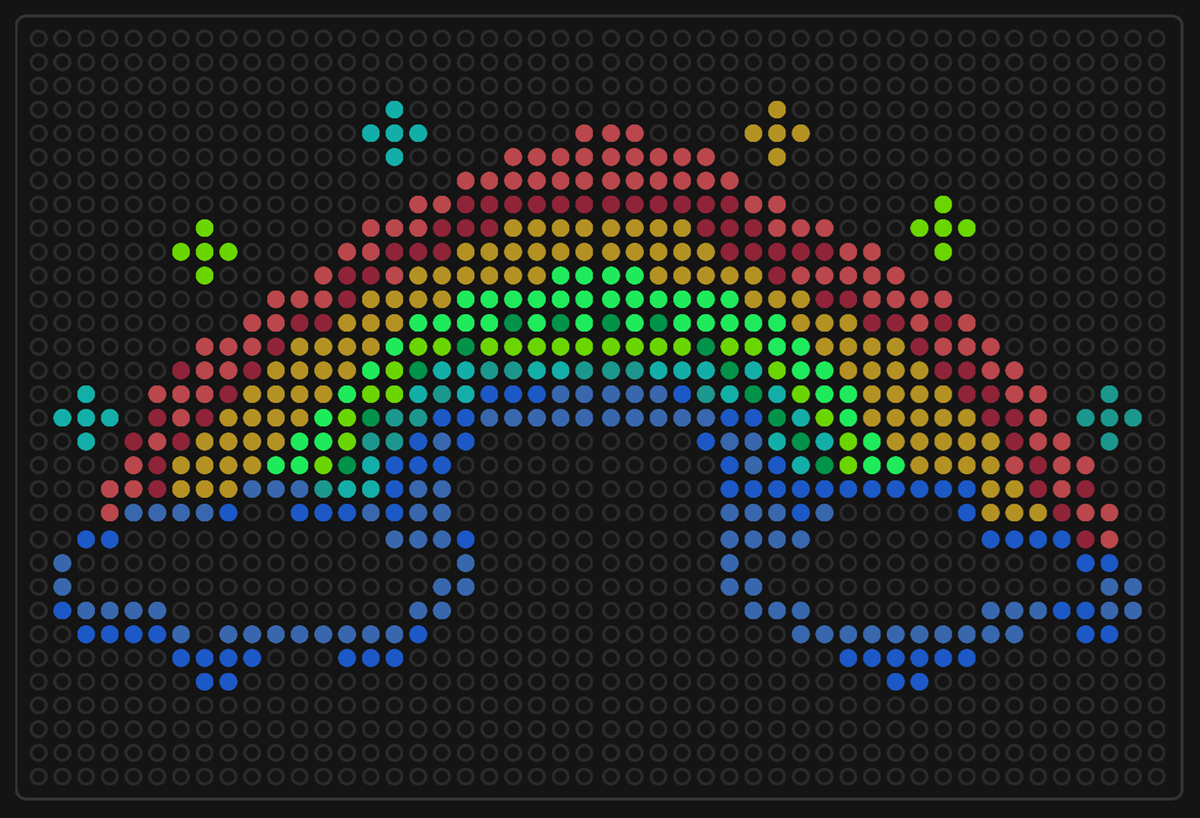


This design had waaaay to many points and the code was 30 pages long. It has 7 colours or something like this!
Post-Lab Questions — DUE BY START OF FEB 24 LECTURE
One of the great parts about having an automated robot is being able to precisely mix, deposit, and run reactions without much intervention, and design and deploy experiments remotely.
For this week, we’d like for you to do the following:
- Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
I found 2 papers but none of them uses opentrons particularly. I am interested in bacterial and textile dyes.


is this amazing paper on DIY liquid handling robots for integrated STEM education and life science research.
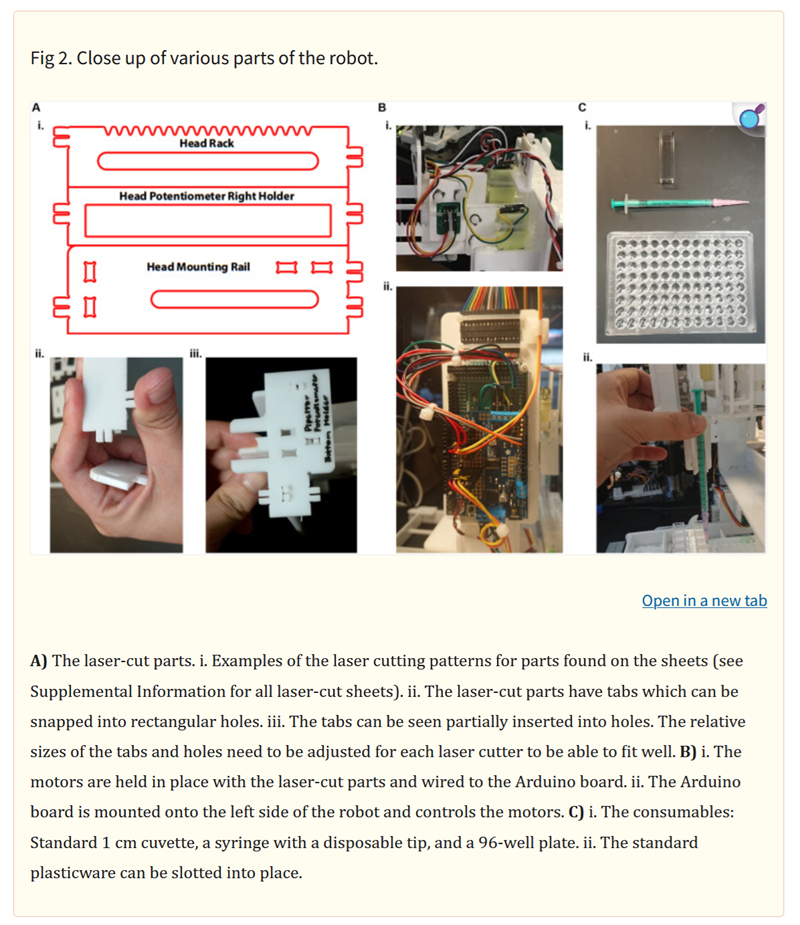



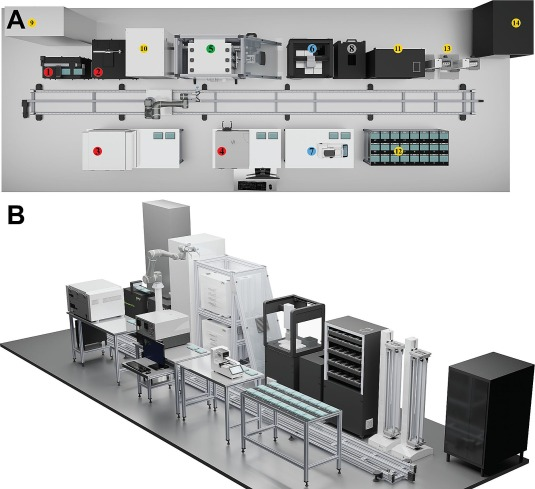
Here you can find the second paper on Automated phenotyping of microalgae: scalable solution for high-throughput analysis.



- Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more. You may reference this week’s recitation slide deck for lab automation details.
While your description/project idea doesn’t need to be set in stone, we would like to see core details of what you would automate. This is due at the start of lecture and does not need to be tested on the Opentrons yet.
Example 1: You are creating a custom fabric, and want to deposit art onto specific parts that need to be intertwined in odd ways. You can design a 3D printed holder to attach this fabric to it, and be able to deposit bio art on top. Check out the Opentrons 3D Printing Directory.
Example 2: You are using the cloud laboratory to screen an array of biosensor constructs that you design, synthesize, and express using cell-free protein synthesis.
- Echo transfer biosensor constructs and any required cofactors into specified wells.
- Bravo stamp in CPFS reagent master mix into all wells of a 96-well / 384-well plate.
- Multiflo dispense the CFPS lysate to all wells to start protein expression.
- PlateLoc seal the plate.
- Inheco incubate the plate at 37°C while the biosensor proteins are synthesized.
- XPeel remove the seal.
- PHERAstar measure fluorescence to compare biosensor responses.
For my final project I would be interested in making a printer or a diy dye handling machine that works with natural or bacterial dyes and prints directly on fabric or maybe make an open source one. I think I need a bit more time to find more opentrons examples in textile dyeing!
Final Project Ideas — DUE BY START OF FEB 24 LECTURE
To be completely honest I am more interested in natural and bacterial dyes, food and crystallisation more than ever. My personal work reflects this. I have a lot of ideas but I will try stick to these even though I want to focus more on edible delights, food, bacterial and natural dyes and pigments.
We have to add 3 final ideas here.

My final project idea number one explores the mining of colour inducing bacteria from the human microbiome for food, cosmetics, the production of sustainable textile dyes and other natural fibres such as human hair, natural weaving material such as straw, palm tree leaves etc.


here is me using my DIY inoculation loop that I designed and printed to inoculate some Janthinobacterium lividum bacteria that produces the Violacein pigment and give off a Purple colour
According to this paper these are the pigment producing bacteria.


Color-producing bacteria exist in various environments, including on human skin, in water, and in soil, where they produce pigments as a survival mechanism against UV radiation, oxidative stress, or to compete with other microbes. These bacteria, often found in the human skin microbiome, produce natural, biodegradable, and often non-toxic pigments such as carotenoids (yellow/red/orange), violacein (purple), prodigiosin (red), and melanin (black/brown).
Key Color-Producing Bacteria on Human Skin
- Staphylococcus aureus (Golden Yellow): Produces staphyloxanthin, a carotenoid pigment that gives it a golden color. This pigment acts as an antioxidant, helping the bacterium withstand oxidative bursts from human immune cells.
- Micrococcus luteus (Yellow): Frequently found on human skin, this bacterium produces yellow carotenoid pigments that can absorb UV radiation.
- Pseudomonas aeruginosa (Blue-Green): Often found in infections (e.g., burns, wounds), it produces pyocyanin (blue-green) and pyoverdine (yellow-green).
- Corynebacterium species (Various/Creamish): Some species in the skin microbiome produce pigments like indogoidine (blue).
- Streptococcus agalactiae (Orange-Red): Known to produce a pigment called granadaene, which is linked to its virulence.
Common Pigments and Their Sources
- Prodigiosin (Red): Produced by Serratia marcescens, a bacterium that can be found on skin or in the environment.
- Violacein (Purple): Produced by Chromobacterium violaceum and Janthinobacterium lividum, these are found in water and soil, but sometimes on skin.
- Melanin (Black/Brown): Produced by various bacteria, including Pseudomonas and Bacillus species, providing photoprotection.
Significance to Humans
- Clinical Diagnosis: The distinct colors of these bacteria on agar plates are used in clinical labs for rapid identification (e.g., the “golden” S. aureus).
- Skin Health/Pathogenesis: Pigments like staphyloxanthin help pathogens evade the immune system, acting as virulence factors.
- Industrial/Medical Applications: i. Textiles: Bacteria like Janthinobacterium lividum are used to dye fabrics (silk, cotton, wool) with natural purple colors. ii. Cosmetics/Medicine: Bacterial pigments are being researched as natural, UV-protective ingredients for sunscreens and as anti-cancer agents. iii. Food: Some, like prodigiosin, are explored for use as natural food colorants, though many are still under study for safety.
These pigments are not just for color; they are essential for bacterial survival under stress.
Here are some resource from my fabricademy page on how to do bacterial dyeing step by step. Here is a steo by step guide. This is another great resource from open cell with the protocol for bacterial dyeing.
I have a few ideas on how I can work with this concept based on the significance to humans section above. Just like natural pigments I suspect that you can also create a fully circular system with utilising bacterial pigments too.
Here you can find my own open source resource that I created for my students for a zero waste circular journey in natural dyes! I am interested in this model for my project idea too.
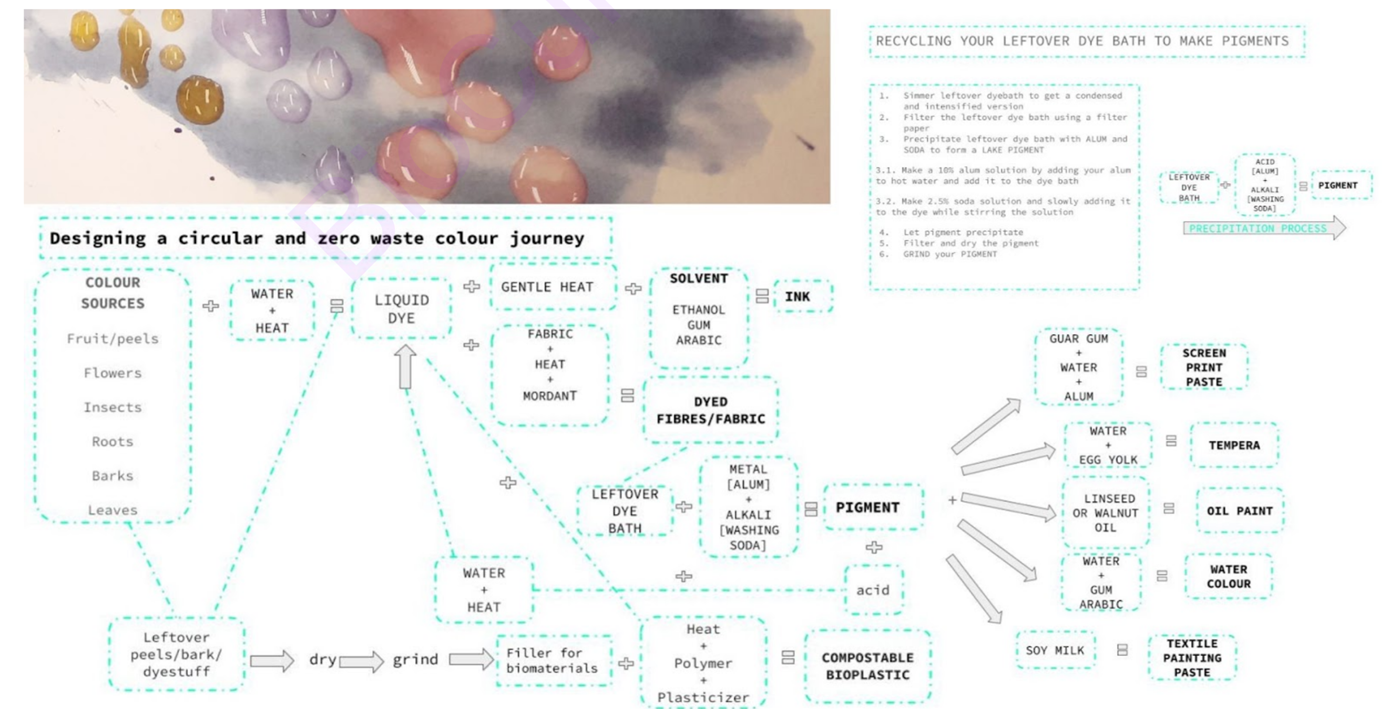

Here is a screenshot of the circular system design. I did not design it. Its Cecilia Raspanti of textile lab in Waag academy that did for fabricademy.
If you want to get into BIOCHROMES (natural sources of colour) and want to try it at home using foodscraps (avocado pits, promegranate peels, pecan or wallnut shells etc) use my diy guide below, enjoy <3




My second idea that is again based on circularity is A domestic DIY mycelium lab for breaking down household single use plastics. How to train your mycelium…to eat plastic
I asked chatGPT to make me a picture but this is not my vision to be honest. I just appreciate the humour of chatGPT, haha. I did not imagine my domestic diy mycelium lab like this.


Inspired by this project called Fungi mutarium (2011) by Katharina Unger, recycles plastic while growing edible treats. It is a prototype system that uses fungi to grow edible biomass (mycelium) on plastic waste. The process involves placing plastic in agar cups (“FU”) filled with fungi. The aim of Livin Studio’s project is to use commonly uneaten parts of fungi to break down plastic while simultaneously producing a novelty food product.
She began working with two widely consumed types of fungus: Pleurotus Ostreatus, more commonly known as Oyster Mushroom and found on Western supermarket shelves, and Schizophyllum Commune, colloquially named Split Gill that is eaten in Asia, Africa and Mexico.


Producing edible treats from this process adds more dimensions to the project and creates a zero waste circular journey adding to the circularity of the system explored <3 I LOVE THIS PROJECT AND I HAD THIS IDEA OF MAKING EDIBLE FUNGI SCAFFOLDS MYSELF. MORE LINKS COMING SOON!


Single-use plastics are goods that are made primarily from fossil fuel–based chemicals (petrochemicals) and are meant to be disposed of right after use—often, in mere minutes. Single-use plastics are most commonly used for packaging and serviceware, such as bags, bottles, wrappers, and straws.
Single use plastics PET (Polyethylene Terephthalate): Used for drink bottles, water bottles, and food containers. HDPE (High-Density Polyethylene): Found in milk jugs, shampoo bottles, and sturdy, often reusable shopping bags. LDPE (Low-Density Polyethylene): Used for flexible plastics like cling wrap, bread bags, and grocery bags. PP (Polypropylene): Common in microwaveable food containers, yogurt tubs, potato chip bags, and bottle caps. PS (Polystyrene) & EPS (Expanded Polystyrene): Used for disposable cutlery, plates, cups, and foam food packaging.
As well as recycling nutrients and helping plants and crop grow efficiently, fungi provide us with compounds that produce antibiotics, statins for treating cholesterol and immunosuppressants. Fungarium projects like at Kew Gardens, focused on breaking down plastic, often termed mycoremediation, involve using specialized fungi to degrade synthetic polymers into organic matter. Research from institutions like Kew Gardens and various university teams has identified fungi capable of breaking down plastics—specifically polyurethane and polypropylene—in a matter of weeks, rather than centuries.
Kew Gardens Research: Scientists are mapping the “terrestrial plastisphere” to identify how fungal enzymes can degrade common, hard-to-recycle plastics.
Dr Irina Druzhinina has been studying hundreds of fungal species, as well as bacteria, that make their home on the surface of plants like Welwitschia and certain palms. What makes these plants interesting is their thick, waxy leaf cuticles made of polymers with remarkably similar traits to plastic. To avoid being swept away from their leaf surface home by the elements, fungi secrete enzymes that digest waxy leaf polymers, allowing for better grip. If they can easily digest plant polymers, it stands to reason they may have some ability to digest plastic too. Already, Irina and her international collaborators have identified more than 180 species whose enzymes could digest basic plastics in a lab setting. Identifying the genes associated with this ability and making use of a huge new fungal DNA dataset, could accelerate the finding of other fungi with plastic-eating potential far more quickly than we can now. A fungus-based solution to the enormous issue of plastic pollution could be just years away.


here is Kew gardens fungarium collection <3
and a beautiful video by Katharina Unger on her process for Fungi mutarium


My third idea is a wearable microfluidic sweat collector and biosensor that enables the detection of hormones and endocrine-disrupting chemicals (EDC), including xenoestrogens.
For non-invasive, personalized reproductive hormone monitoring. Apart ftom PFAS sweat collectors can also monitor reproductive health in the paper - A wearable aptamer nanobiosensor for non-invasive female hormone monitoring.
This 2026 paper on Sweat-wearable biosensors for real-time monitoring of endocrine-disrupting chemicals: Materials, analytics, and public-health integration states that researchers have developed flexible, skin-interfaced biosensors that utilize sweat-based monitoring to provide real-time, non-invasive alternatives to traditional blood tests for tracking hormonal health.
Another paper and project this time from caltech is the Wearable Patch Wirelessly Monitors Estrogen in Sweat.
Here is a video from Gao Research Group on their Wearable Estrogen Sensor
In addition, there is a portable sensor that could detect PFAS on site. Led by PhD student Henry Bellette and Dr Saimon Moraes Silva, Director of La Trobe’s Biomedical and Environmental Sensor Technology (BEST) Research Centre, the research has been published in the journal ACS Sensors.
“Most PFAS testing relies on expensive laboratory equipment and specialist analysis, which makes regular monitoring difficult,” he said.
“This biosensor could be used on site and provides a simple yes or no result, allowing water to be screened quickly and easily.”
As you can see on the top section of the page i added some more microfluidics research based on sweat collection <3


Paper microfluidics are also very fun to do at a DIY level. I read this paper!
Another amazing example on paper microfluidics is the 3D Paper Mmicrofluidic Device Fabricated by Embossing fabricated on two layers of omniphobic paper containing different microchannels. Liquids flowing on the upper layer get transfer to the layer underneath to avoid mixing.