Week 9 HW — Cell-Free Systems

This week introduces synthesis of proteins using cellular machinery outside of a cell. I LOVED THE LECTURE and I loved Kate Adamalas work.
General homework questions
- Explain the main advantages of cell-free protein synthesis over traditional in vivo methods, specifically in terms of flexibility and control over experimental variables. Name at least two cases where cell free expression is more beneficial than cell production.
A cell-free system allows biological reactions to occur outside of living cells. By extracting and using cellular components like ribosomes, RNA polymerase, amino acids, and ATP, this method enables reactions in a controlled, simplified environment. Cell-free systems allow for the engineering, expression, and analysis of genetic constructs without the complexity of living cells.
Applications of Cell-Free Systems include:
+ Synthetic Biology: Designing and testing biological circuits or pathways without cellular constraints.
+ Protein Engineering: Rapid protein production and screening, especially for proteins that are toxic or hard to express in cells.
+ Metabolic Engineering: Production of high-value chemicals, biofuels, and pharmaceuticals via synthetic pathways.
+ Biosensing: Creating diagnostic tools that are portable and easy to use, like paper-based biosensors.
Gene Editing Research: Testing CRISPR-based systems or genome editing tools in a controlled environment.
Cases where cell free expression is more benedicial than cell production?
Cell free expression is often more beneficial than conventional cell-based (in vivo) production in scenarios requiring speed, high-throughput screening, or when producing proteins that are toxic or difficult to fold. By bypassing the need for living cells, cell free protein expression removes constraints associated with cell viability, growth, and genetic stability, acting as an “open” system that allows higher flexibility, speed, control over the environment and direct manipulation.
- Describe the main components of a cell-free expression system and explain the role of each component.
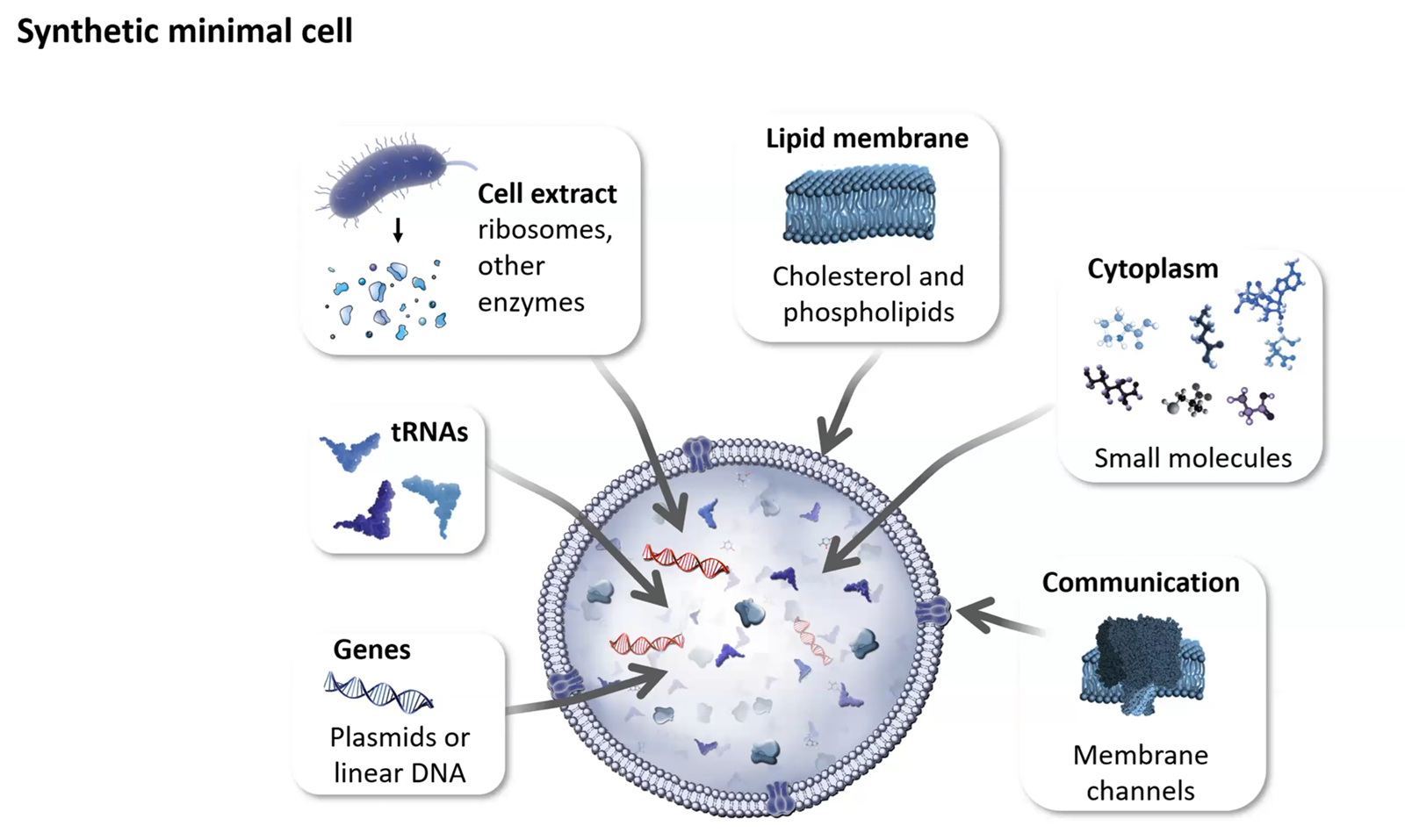

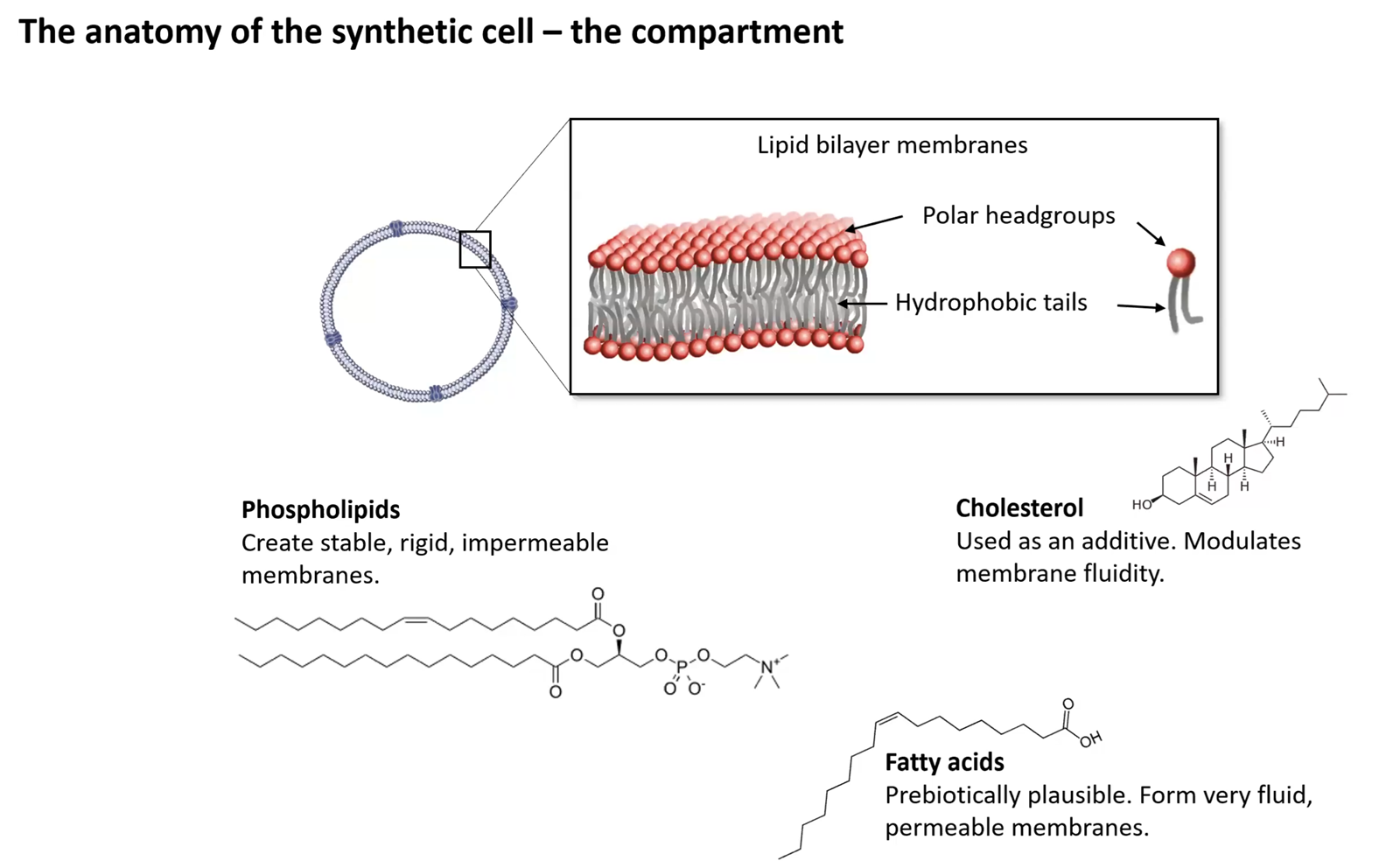

Our chassis for engineering!
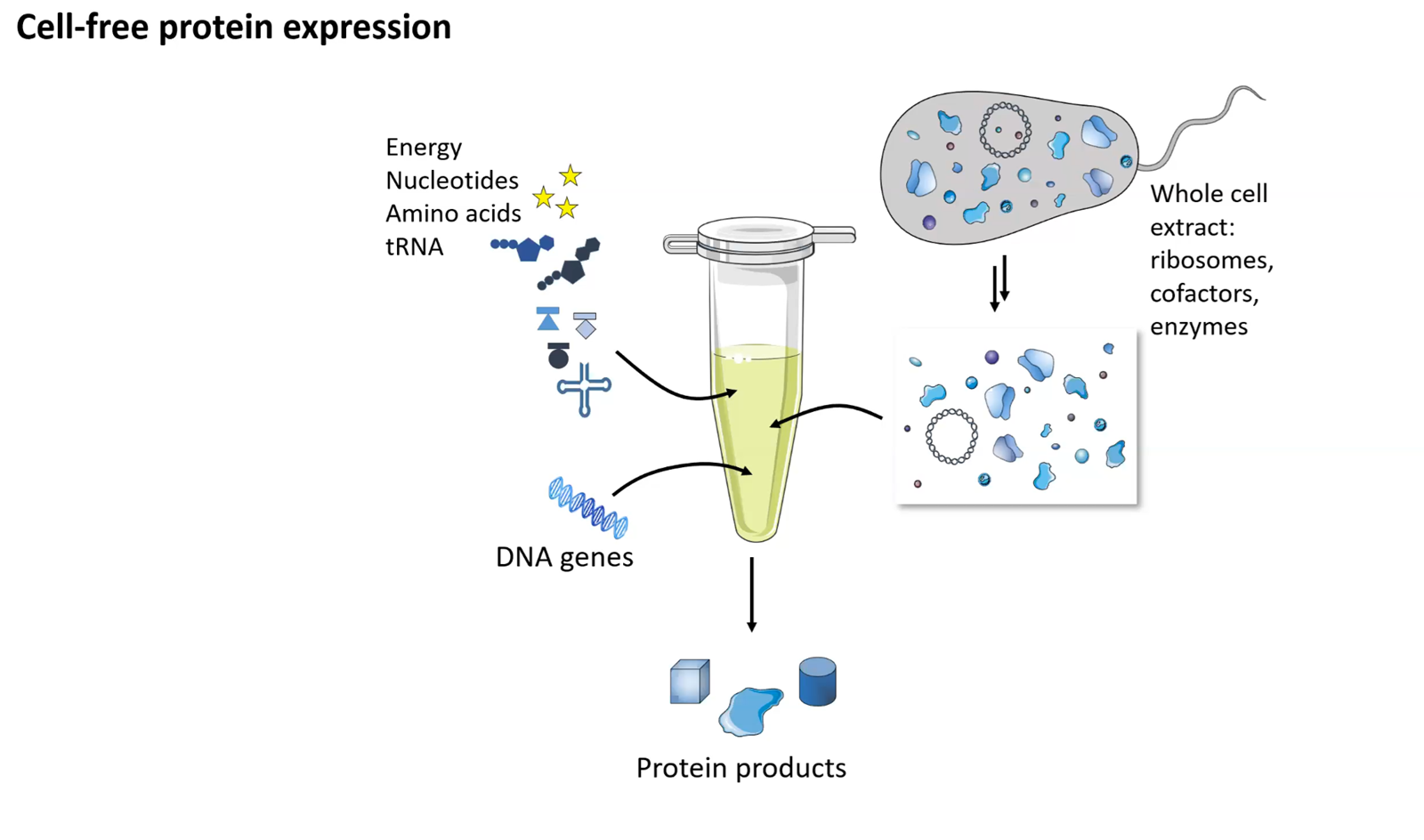

Explain the role of each component:
From the bioclub hangout-> slight confusion between cell free systems vs synthetic cells!
Cell-free systems use extracted, non-living cellular machinery (ribosomes, enzymes) in an open, in vitro environment to perform tasks like protein synthesis, offering high control and flexibility. Synthetic cells (or protocells) are bottom-up, engineered, microscopic compartments—often using lipid vesicles—that encapsulate biochemical reactions to mimic living cell structure and function, providing compartmentalization. Cell-Free Systems (CFS)
Definition: Molecular machinery removed from cells (e.g., E. coli lysates) used in an open reaction mixture.
Key Advantage: Direct access to the machinery, allowing for easy monitoring, modulation of conditions, and elimination of cell-membrane transport issues.
Applications: Rapid prototyping of genetic circuits, producing toxic proteins, and building molecular sensors.
Limitations: Lack of compartmentalization leads to degradation over time and inability to maintain complex homeostasis.
Synthetic Cells (Protocells)
Definition: Artificial structures, such as liposomes, that package synthetic biology components (like a cell-free system) inside a membrane to simulate a functional cell.
Key Advantage: Compartmentalization allows for local concentration of components, mimicry of cell architecture, and protection of the internal environment.
Applications: Studying the origin of life, creating artificial organisms for drug delivery, and modeling cellular interactions.
Limitations: High complexity in building the membrane and transporting materials in/out of the structure.
Key Comparisons
Control: Cell-free systems allow direct, open control, while synthetic cells require managing artificial membrane boundaries.
Complexity: Synthetic cells are more complex because they aim to reconstruct a minimal, living-like unit.
Stability: Cell-free reactions can be rapid but fleeting, whereas synthetic cells can be designed for higher stability.
In summary, cell-free systems are a technology for utilizing machinery, while synthetic cells are a platform for engineering life-like structures.
SOURCES
- Cell-Free PURE System: Evolution and Achievements.
- Cell-Free Synthetic Biology: Thinking Outside the Cell.
- Advancing synthetic biology through cell-free protein synthesis.
- Why is energy provision regeneration critical in cell-free systems? Describe a method you could use to ensure continuous ATP supply in your cell-free experiment.
Found in Energy Systems for ATP Regeneration in Cell-Free Protein Synthesis Reactions.
Method for Continuous ATP Supply: Phosphoenolpyruvate (PEP) Regeneration System. A highly effective method to ensure continuous ATP supply is using the PEP/pyruvate kinase (PK) system.
Component: Add Phosphoenolpyruvate (PEP) (approx. 20–30 mM) and Pyruvate Kinase enzyme to the reaction mixture. Mechanism: Pyruvate kinase catalyzes the transfer of a phosphate group from PEP to ADP, converting it back into ATP, thereby maintaining high ATP/ADP ratios. Alternative Approach: For reduced cost and inorganic phosphate buildup, using [Glucose or Glucose-6-Phosphate in combination with NAD+]((https://pmc.ncbi.nlm.nih.gov/articles/PMC4651010/) can enable sustained ATP generation through glycolysis.
- Compare prokaryotic versus eukaryotic cell-free expression systems. Choose a protein to produce in each system and explain why.
Prokaryotic cell-free protein synthesis (CFPS) systems, notably E. coli, offer high-yield, low-cost production for simple proteins, while eukaryotic systems (e.g., wheat germ, rabbit reticulocyte) excel in folding complex proteins requiring post-translational modifications. Prokaryotic systems are faster and produce higher protein concentrations (up to 2 mg/mL), whereas eukaryotic systems provide superior machinery for complex post-translational modifications
Prokaryotic vs. Eukaryotic Cell-Free Expression Systems
Prokaryotic (E. coli):
Advantages: Rapid, cost-effective, high yield of protein (>2μg/ml). Ideal for high-throughput screening.
Disadvantages: Lacks endoplasmic reticulum/Golgi, meaning it cannot perform complex PTMs (glycosylation) and often fails to fold complex eukaryotic proteins properly.
Eukaryotic (Wheat Germ, Rabbit Reticulocyte Lysate - RRL):
Advantages: Contains necessary chaperones, folding enzymes, and membranes to support disulfide bonds, glycosylation, and proper post-translational modifications.
Disadvantages: Generally lower yield, higher costs, and slower, more complex protocols compared to E. coli.
Systems and proteins chosen
Prokaryotic-> Green Fluorescent Protein (GFP) As a relatively small, fast-folding protein that does not require glycosylation, GFP is ideal for high-yield expression in E. coli extracts. The goal here is sheer quantity, speed, and cost-effectiveness for imaging or structural studies.
Eukaryotic-> Human Insulin Active insulin requires the formation of specific disulfide bonds to link the A and B chains. Eukaryotic systems (such as Wheat Germ or RRL) provide the necessary chaperone environment to fold this protein correctly, whereas E. coli would likely produce it as an inactive inclusion body.
Sources
- Cell-Free Protein Synthesis: Pros and Cons of Prokaryotic and Eukaryotic Systems.
- A Comparative Guide: Prokaryotic vs. Eukaryotic Cell-Free Expression Systems for Eukaryotic Proteins.
- How would you design a cell-free experiment to optimize the expression of a membrane protein? Discuss the challenges and how you would address them in your setup.
Designing a cell-free experiment to optimize phospholipid membrane protein expression involves using a rapid, flexible platform like E. coli-based MyTxTL or PURE system. Key steps include optimizing DNA template design (T7 promoter), supplementing with detergents (e.g., Brij-35) or lipid nanodiscs/liposomes for solubility, and using a continuous exchange system to improve yield. You can Improve Cell-Free Expression of Model Membrane Proteins by Tuning Ribosome Cotranslational Membrane Association and Nascent Chain Aggregation.
Some of the challenges and solutions
Protein Aggregation & Misfolding: MPs are hydrophobic and aggregate without a lipid bilayer.
Solution: Introduce pre-formed liposomes or nanodiscs into the reaction, or use detergents to create artificial membrane environments for direct insertion during synthesis.
Low Expression Yields: High costs and time can be wasted if expression is low.
Solution: Use automated liquid handling for small-scale (50
L) screening and optimize the ATP regeneration system.
Template Degradation: Linear templates are vulnerable to endogenous nucleases.
Solution: Use Gam protein to inhibit exonucleases or switch to circular plasmid templates.
Incorrect Redox Potential: The cytoplasm of the lysate is reducing, while many membrane proteins require an oxidizing environment for disulfide bonds.
Solution: Add glutathione (GSH/GSSG) to balance the redox state and promote correct folding.
Degradation by Proteases: Cell lysates contain proteases.
Solution: Add protease inhibitor cocktails directly to the reaction mixture.
Sources
- Combining In vitro Folding with Cell-Free Protein Synthesis for Membrane Protein Expression
- Enhancing the Cell-Free Expression of Native Membrane Proteins by In Silico Optimization of the Coding Sequence—An Experimental Study of the Human Voltage-Dependent Anion Channel
- Imagine you observe a low yield of your target protein in a cell-free system. Describe three possible reasons for this and suggest a troubleshooting strategy for each.
I imagine this has to do with rapid energy depletion. A synthetic cell also has a limited lifespan and components and does not have the ability to regenarate.
Rapid Energy Depletion: Cell-free systems rely on high-energy molecules like ATP and phosphoenolpyruvate (PEP) to fuel transcription and translation. These energy sources are quickly used up in a batch reaction, stopping protein synthesis within 1-3 hours. Inhibitory Byproduct Accumulation: The metabolic activities within the cell lysate produce byproducts, such as inorganic phosphate, that accumulate and inhibit further synthesis. Limited Lifespan of Machinery: The transcriptional and translational machinery (ribosomes, enzymes) is not being regenerated as it would be in a living cell, leading to a loss of functionality over time.
Here you can find some of the info i wrote above.
Troubleshooting!
Overcoming Low Yields:
- Continuous Exchange Cell-Free (CECF): Using dialysis systems to constantly add energy and remove byproducts can increase reaction times up to 24 hours and boost yields significantly.
- Optimized Energy Systems: Switching from PEP to more efficient systems like glucose or maltodextrin can prevent byproduct accumulation.
- GamS Supplementation: Adding GamS protein helps protect linear DNA from degradation, boosting yield.
Homework question from Kate Adamala
Design an example of a useful synthetic minimal cell as follows:


Artificial cells translate chemical signals for E. coli. (a) In the absence of artificial cells (circles), E. coli (oblong) cannot sense theophylline. (b) Artificial cells can be engineered to detect theophylline and in response release IPTG, a chemical signal that induces a response in E. coli.
- Pick a function and describe it.
a. What would your synthetic cell do? What is the input and what is the output?
My synthetic cell will produce vitamin D (cholecalciferol) on demand. Input (Precursor): 7-Dehydrocholesterol (7-DHC). This molecule, present in cell membranes, serves as the provitamin D3. The secondI input (Energy): UVB light (typically 290–315 nm). Output (Product): Previtamin D3, which subsequently rearranges into Vitamin D3 (Cholecalciferol) (see ref below for sources 1 and 2 that helped me figure this out).
Based on synthetic biology principles and the cutaneous vitamin D synthesis pathway, a synthetic minimal cell engineered to produce Vitamin D functions as a specialized, self-contained factory, transforming a specific sterol precursor into the vitamin.
b. Could this function be realized by cell-free Tx/Tl alone, without encapsulation?
Yes, this function can be realized by cell-free Transcription-Translation (Tx/Tl) systems without encapsulation, essentially treating the reaction as a “liquid catalyst” rather than a cell. In a cell-free setup, you would provide the DNA templates for the necessary enzymes (like the cytochrome P450s) directly to a lysate (such as E. coli or wheat germ).
c. Could this function be realized by genetically modified natural cell?
Yes, this function has already been successfully realized using genetically modified natural cells, primarily in yeast (Saccharomyces cerevisiae) and plants like tomatoes. Unlike minimal synthetic cells, these natural hosts provide an existing metabolic framework (the mevalonate pathway) that can be redirected to accumulate Vitamin D precursors (3).
d. Describe the desired outcome of your synthetic cell operation.
The desired outcome of this synthetic cell operation is the autonomous, high-yield production of active Vitamin D in a controlled environment. By stripping away the “noise” of a natural organism, the operation achieves three specific goals (1,2,3):
Continuous Synthesis: Unlike humans, who rely on intermittent sun exposure and complex organ signaling (skin to liver to kidney), the synthetic cell provides a one-stop shop. It takes in a simple precursor (7-DHC) and UVB light to output the final, bioactive form (Calcitriol) immediately.
Purity and Specificity: In natural systems, UVB exposure often creates inactive “side products” (like lumisterol) to prevent Vitamin D toxicity. The synthetic cell is engineered to bypass these safety brakes, ensuring that nearly 100% of the input is converted into the useful metabolite without “wasted” chemistry.
Deployment Flexibility: The ultimate goal is to create a “biomachine” that can function where natural cells cannot—such as in a wearable patch that synthesizes Vitamin D through the skin, or as a fortification factory in a bioreactor that produces vegan-sourced Vitamin D3 at an industrial scale.
Essentially, the outcome is a standardized, programmable source of Vitamin D that removes the unpredictability of diet, geography, and skin pigmentation.
Sources
Vitamin D: Newer Concepts of Its Metabolism and Function at the Basic and Clinical Level.
Engineered yeast for efficient de novo synthesis of 7-dehydrocholesterol.
Design all components that would need to be part of your synthetic cell. I used google search for this and AI.
To design a synthetic minimal cell for Vitamin D production, you must integrate four primary structural and functional “modules.” This bottom-up design ensures the cell can host the necessary chemistry while remaining lean and efficient.
1. Structural Chassis: The Boundary Layer
The cell requires a physical barrier to concentrate enzymes and protect the reaction.
- Composition: A Phospholipid Bilayer made of a mix of zwitterionic lipids like Phosphatidylethanolamine (PE) and anionic lipids like Phosphatidylglycerol (PG).
- Stabilizers: Adding Cholesterol or its analogs increases membrane toughness and prevents the leakage of small molecules.
- Encapsulation: These lipids are assembled into Giant Unilamellar Vesicles (GUVs), which are large enough (10–100 µm) to be easily monitored and handled.
2. Metabolic Machinery: Membrane-Bound Enzymes
The conversion of precursors into active Vitamin D relies on specific proteins that must be embedded in the membrane.
- Internal Organelle (Nanodiscs): Since Vitamin D enzymes are membrane-bound, the cell should contain internal lipid nanodiscs or micelles to provide docking sites for proper protein folding.
- Key Catalysts:
- 25-hydroxylase (CYP2R1): Anchored to the internal membrane to convert cholecalciferol into calcidiol.
- 1-alpha-hydroxylase (CYP27B1): The “final step” enzyme that produces the active hormone, Calcitriol.
3. Genetic Information Unit: The Software
Instead of a full genome, this cell uses a minimal genetic circuit to maintain its production line.
- Genome Chassis: A stripped-down DNA template based on the JCVI-syn3.0 model, which contains only the ~473 genes essential for basic survival.
- Custom Operon: A specialized DNA “plug-in” containing the genes for the CYP enzymes mentioned above, controlled by a light-sensitive or chemical promoter to turn production “on” or “off.”
4. Energy and Support: The “Fuel” Module
A minimal cell cannot generate all its own energy, so these components must be supplied or engineered.
- External Power: UVB Light (290–315 nm) serves as the primary energy input for the first non-enzymatic step of synthesis.
- Cofactor Regeneration: A system to recycle NADPH and provide molecular oxygen, which are required for the hydroxylation steps to continue without stalling.
- Transport Proteins: Embedded pore proteins (like alpha-hemolysin) to allow the 7-DHC precursor in and the finished Vitamin D out.
| Component Category | Specific Requirement | Justification |
|---|---|---|
| Boundary | Phospholipid Bilayer | Maintains the internal environment and chemical gradients. |
| Genotype | Minimal DNA Circuit | Provides the blueprints for enzyme production with zero “waste”. |
| Enzymes | CYP2R1 & CYP27B1 | Necessary for the multi-step hydroxylation of the sterol backbone. |
| Logistics | Membrane Pores | Facilitates the “Input-Output” flow of precursors and products. |
a. What would be the membrane made of?
To ensure the membrane can support the specific “greasy” chemistry of Vitamin D synthesis, it would be composed of a hybrid phospholipid bilayer designed for stability and enzyme integration. The recipe for the membrane-
The Base: DOPC and DOPE
DOPC (Dioleoyl-phosphatidylcholine): This provides the flexible, fluid matrix. It is the “standard” lipid that forms the stable bilayer structure. DOPE (Dioleoyl-phosphatidylserine): This adds “curvature” to the membrane, which is essential for helping the bulky CYP enzymes (like CYP2R1) wedge themselves into the surface without destabilizing the cell.
The Functional Anchor: DGS-NTA(Ni)
Since your enzymes are the stars of the show, you include a small percentage of Nickel-chelating lipids. Why? These act like “Velcro” for your proteins. If your enzymes are engineered with a Histidine-tag (His-tag), they will snap onto these lipids, ensuring they stay anchored to the membrane where the reaction happens.
The Fluidity Regulator: Cholesterol
While the cell’s goal is to make a cholesterol-like product, you must include Cholesterol (around 10–20%) in the membrane itself. Why? It acts as a “buffer” for membrane thickness and prevents the lipid wall from becoming too leaky or too stiff as temperature changes during UVB exposure.
The Protective Coating: PEG-Lipids
PEG-DSPE: A lipid with a “tail” of Polyethylene Glycol. Why? This creates a “stealth” layer on the outside of the cell, preventing it from clumping together with other cells or sticking to the walls of your container (bioreactor or patch).
Transport Pores: Alpha-Hemolysin
Strictly speaking, this is a protein, but it functions as part of the membrane. These are “holes” that allow the 7-DHC input to enter the cell and the Vitamin D output to exit, as the lipid bilayer alone is too dense for them to pass through quickly.
Summary Profile:
Phase: Liquid-disordered (fluid)
Charge: Slightly negative (to mimic natural endoplasmic reticulum membranes)
Stability: High (enhanced by cholesterol)
b. What would you encapsulate inside? Enzymes, small molecules.
To make the synthetic cell functional, the interior (the lumen) must contain the “operating system” and the raw materials needed to build and power the Vitamin D synthesis machinery.
Since this is a minimal cell, you would encapsulate:
1. The Genetic Instructions (DNA/RNA)
- DNA Plasmids: Coding for the three essential enzymes: 7-DHC reductase (to manage precursor flow), CYP2R1, and CYP27B1.
- Alternative (mRNA): If you want a faster “start-up,” you can encapsulate the mRNA directly, bypassing the need for transcription.
2. Transcription/Translation Machinery (Tx/Tl)
Since the cell has no natural nucleus or ribosomes, you must “borrow” them:
- Ribosomes: Sourced from E. coli or wheat germ lysates to build the enzymes from the DNA/RNA.
- RNA Polymerase: To read the DNA templates.
- tRNAs and Amino Acids: The 20 standard building blocks required to assemble the protein enzymes.
3. Energy and Redox Small Molecules
The Vitamin D “engine” (CYP enzymes) requires a constant flow of “fuel”:
- ATP, GTP, UTP, CTP: The chemical energy needed to power protein synthesis.
- NADPH: The critical electron donor for the hydroxylation steps.
- Creatine Phosphate & Creatine Kinase: A “battery” system to regenerate ATP as it gets used up, extending the cell’s lifespan.
4. Supporting Small Molecules & Buffer
- Magnesium and Potassium Salts: Essential ions that stabilize DNA and keep ribosomes functional.
- Reducing Agents (DTT or TCEP): To prevent the enzymes from oxidizing and clumping in the watery interior.
- Molecular Oxygen ($O_2$): While a gas, it must be dissolved in the internal fluid as a co-substrate for the hydroxylation reactions.
5. Specialized “Docking” Components
- Nanodiscs or Small Unilamellar Vesicles (SUVs): Tiny internal lipid “rafts.” Since the CYP enzymes are hydrophobic, they need these internal membranes to fold correctly before they eventually migrate to the main outer membrane.
Summary of the “Inside”: It would look like a concentrated molecular soup—highly viscous, packed with the machinery of life, and primed to start building enzymes the moment the temperature or light conditions are right.
c. Which organism your Tx/Tl system will come from? Is bacterial OK, or do you need a mammalian system for some reason? (hint: for example, if you want to use small molecule modulated promotors, like Tet-ON, you need mammalian)
A bacterial Tx/Tl system (specifically an E. coli lysate like PURE) is actually the best fit for this minimal cell, provided we address one specific “mammalian” requirement.
d. How will your synthetic cell communicate with the environment? (hint: are substrates permeable? or do you need to express the membrane channel?)
The synthetic cell will communicate with its environment through a hybrid permeability strategy. Since Vitamin D synthesis involves bulky, hydrophobic sterols, we cannot rely solely on the lipid bilayer’s natural diffusion rates. [1, 2]
1. Passive Diffusion (For Lipophilic Substrates)
The primary input, 7-Dehydrocholesterol (7-DHC), and the output, Vitamin D3 (Calcitriol), are highly lipophilic.
- The Mechanism: They can naturally partition into and move through the lipid membrane.
- The Limitation: This process is relatively slow and can cause the membrane to become “saturated” with grease, potentially destabilizing the cell or slowing down production as the product builds up inside. [3, 4]
2. Engineered Membrane Channels (For Speed and Small Molecules)
To make the cell a truly efficient “factory,” we must express or insert specific membrane pores and transporters:
- Alpha-Hemolysin ($\alpha$HL): We will express this bacterial pore to create non-selective 1.4 nm “holes.” This allows for the rapid exchange of hydrophilic small molecules like ATP, NADPH, and metal ions (Mg²⁺/K⁺) if the cell is sitting in a nutrient-rich “feed” solution.
- Cyclodextrin “Shuttles”: Since the sterols (7-DHC and Vitamin D) are so hydrophobic, we can add methyl-beta-cyclodextrin to the external environment. These act as molecular “taxis,” picking up the produced Vitamin D from the membrane surface and carrying it into the bulk liquid, maintaining the concentration gradient.
- Stimuli-Responsive Gating: If we want to control when the cell releases its cargo, we could express a mechanosensitive channel like MscL. By engineering it to open under specific triggers (like pH changes or light), we can “flush” the cell’s contents on command. [5, 6, 7, 8, 9]
3. Light Communication
Because the first step of the pathway is photochemical, the environment “communicates” with the cell via UVB light (290–315 nm). This is a wireless input that doesn’t require a physical channel; it simply passes through the membrane to trigger the isomerisation of 7-DHC. [10]
Summary: The cell uses passive diffusion for the “greasy” stuff (helped by external shuttles) and expressed $\alpha$HL pores for the “watery” energy inputs.
[1] https://pmc.ncbi.nlm.nih.gov [2] https://fiveable.me [3] https://pubs.acs.org [4] https://www.ncbi.nlm.nih.gov [5] https://pubs.acs.org [6] https://www.science.org [7] https://www.quora.com [8] https://portlandpress.com [9] https://www.nature.com [10] https://portlandpress.com
- Experimental details
a. List all lipids and genes. (bonus: find the specific genes; for example, instead of just saying “small molecule membrane channel” pick the actual gene.)
To finalize the experimental blueprint for your Vitamin D-producing synthetic cell, here are the specific lipids and genes required for assembly.
a. Lipids (The Membrane Matrix)
The lipid composition is tailored to support CYP enzyme anchoring and maintain structural integrity during UVB exposure.
| Lipid Component | Full Name | Molar Ratio | Role |
|---|---|---|---|
| DOPC | 1,2-dioleoyl-sn-glycero-3-phosphocholine | 50% | The primary fluid bilayer matrix. |
| DOPE | 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine | 20% | Promotes membrane curvature for enzyme insertion. |
| Cholesterol | (Purified) | 20% | Stabilizes the membrane against UVB-induced leakiness. |
| DGS-NTA(Ni) | 1,2-dioleoyl-sn-glycero-3-[(N-(5-amino-1-carboxypentyl)iminodiacetic acid)succinyl] (nickel salt) | 5% | Histidine-tag “docking station” for anchoring enzymes. |
| DSPE-mPEG2000 | 1,2-distearoyl-sn-glycero-3-phosphoethanolamine-N-[methoxy(polyethylene glycol)-2000] | 5% | Prevents vesicle aggregation/clumping. |
b. Genes (The Genetic Circuit)
We will use a modular DNA plasmid approach. These genes are codon-optimized for the E. coli Tx/Tl system to ensure high expression.
| Gene Symbol | Origin | Function |
|---|---|---|
| CYP2R1 | Human/Mammalian | Vitamin D 25-hydroxylase: Converts cholecalciferol into calcidiol. |
| CYP27B1 | Human/Mammalian | 1$\alpha$-hydroxylase: Converts calcidiol into bioactive Calcitriol. |
| ADR | Bovine/Human | Adrenodoxin Reductase: Essential electron transfer partner for the CYP enzymes. |
| ADX | Bovine/Human | Adrenodoxin: Small iron-sulfur protein that shuttles electrons to the CYPs. |
| hlyA | S. aureus | Alpha-Hemolysin: Pore-forming protein for nutrient/product exchange. |
| tetR | E. coli | Tet Repressor: Provides the “On/Off” logic gate for the system. |
The “Logic” Layout:
- Pores: hlyA is expressed under a constitutive (always-on) promoter to ensure the cell can “breathe” immediately.
- Synthesis: CYP2R1, CYP27B1, and the electron partners (ADR/ADX) are placed under a Tet-regulated promoter.
- Activation: The system remains dormant until anhydrotetracycline (aTC) is added to the environment, triggering the “Start” of Vitamin D production.
b. How will you measure the function of your system?
To confirm the cell is actually “breathing” and producing Vitamin D, we need to track both the disappearance of the precursor and the appearance of the active hormone. We will use a three-tier approach to measure function:
1. The Gold Standard: LC-MS/MS
Since Vitamin D metabolites (7-DHC, D3, 25(OH)D, and Calcitriol) look very similar, Liquid Chromatography-Tandem Mass Spectrometry is the only way to be 100% sure of the chemical identity.
- The Process: Take samples of the external “bath” liquid at specific time intervals.
- What we look for: A clear shift in the mass-to-charge ratio (m/z) that matches the addition of hydroxyl groups (-OH) at the 25 and 1$\alpha$ positions.
2. Real-Time Tracking: Fluorescent Sterols
We can use a 7-DHC analog tagged with a fluorescent marker (like BODIPY).
- The Process: Use a Confocal Microscope to watch the cell.
- What we look for: If the fluorescent sterols move from the outside of the cell, concentrate in the membrane, and then change their emission wavelength (if using a “smart” probe sensitive to hydroxylation), we know the transport and metabolic machinery are working in real-time.
3. Biological Activity Assay (The “Calcitirol” Test)
To prove the output is biologically active and not just a “dead” chemical isomer, we can perform a VDR (Vitamin D Receptor) Reporter Assay.
- The Process: Take the output from the synthetic cell and add it to a separate culture of natural cells (like HEK293) that have been engineered with a VDR-linked Luciferase gene.
- What we look for: If the natural cells “glow,” it confirms that the synthetic cell successfully produced 1,25-dihydroxyvitamin D3 in a form that the body can actually use.
4. Metabolic Health: Oxygen Consumption
Since the CYP enzymes consume molecular oxygen ($O_2$) to function, we can use a Clark-type electrode or oxygen-sensitive fluorescent patches in the reaction vessel.
- What we look for: A steady drop in $O_2$ levels immediately following the addition of the “On” switch (aTC), which acts as a proxy for enzymatic activity.
Homework question from Peter
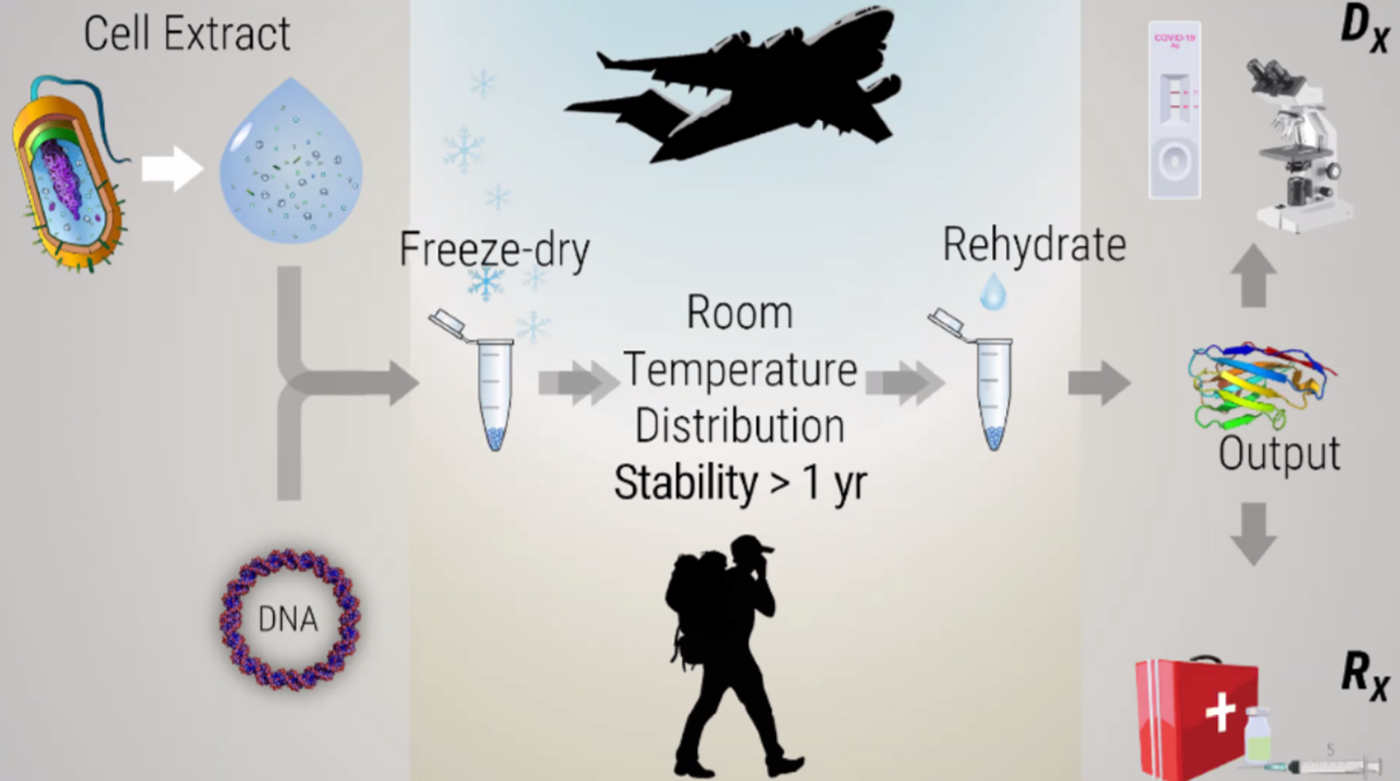

Synthetic biology without living cells! Screenshots from the presentations.
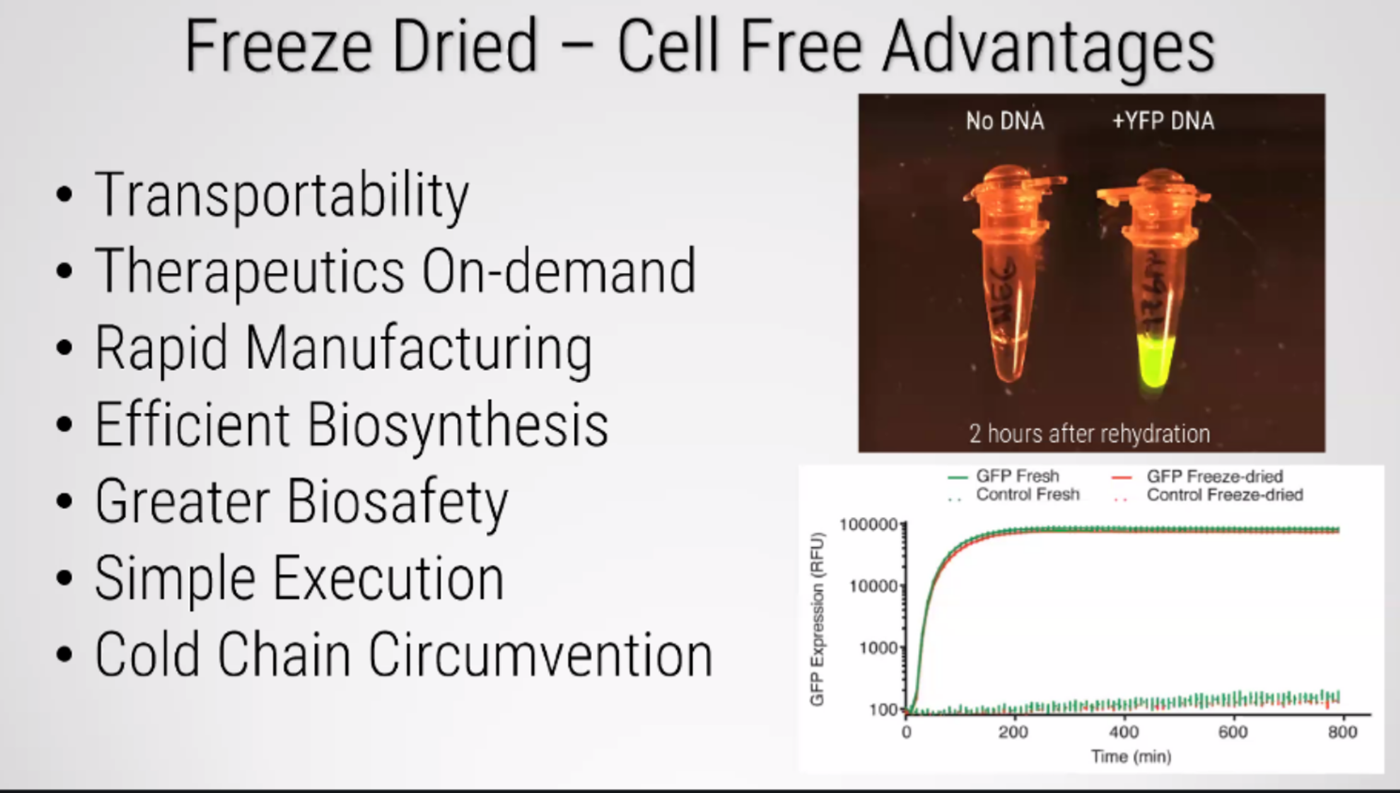

Freeze-drying technology lyophilization, originated during World War II to transport blood plasma and penicillin, later evolving into a critical method for preserving food and pharmaceuticals. It works by freezing materials and reducing pressure, allowing water to transition from ice to vapor (sublimation).
Freeze-dried cell free systems can be incorporated into all kinds of materials as biological sensors or as inducible enzymes to modify the material itself or the surrounding environment. Choose one application field - Architecture, Textiles/Fashion, or Robotics, and propose an application using cell-free systems that are functionally integrated into the material. Answer each of these key questions for your proposal pitch:
- Write a one-sentence summary pitch sentence describing your concept.
I chose to explore textiles/fashion applications of cell-free systems. Can I use this freeze fried cell free system to make a wearable that warns us against radiation exposure? I was really inspired by this project but it uses freeze fried engineered yeast (a wearable microbrewery) to save hospital lab workers from daily radiation exposure. Similarly I want to make a wearable biosensor that produces melanin when radiation is detected.
- How will the idea work, in more detail? Write 3-4 sentences or more.
The core mechanism involves using tyrosinase enzymes to convert precursor molecules (like L-DOPA or L-Tyrosine) into melanin, which is dark brown/black, creating a visible, light-absorbing color change.
The Freeze-Dried Cell-Free System
Source Fungi: Use genes from radiotrophic fungi such as Cryptococcus neoformans or Cladosporium sphaerospermum. Key Enzyme: The gene for tyrosinase (TYR) is essential, as it catalyzes the production of melanin, a pigment that changes color from colorless or light-colored to black/brown. System Formulation: Use Escherichia coli lysate-based cell-free systems, which can be lyophilized (freeze-dried) and stored without refrigeration, designed to rehydrate instantly when exposed to moisture (like sweat). Lyophilization: The reaction mixture—containing the enzyme-producing machinery, DNA blueprints (plasmid), and precursors (e.g., L-tyrosine)—is freeze-dried into a “paper-based” or powder format for integration into fabric.
Making the Wearable Device
Sensor Design: The cell-free reaction is embedded into a flexible substrate, such as a thin silicone film (PDMS) or directly into fabrics like cotton, to create a wearable “patch” or textile sensor. Activation Mechanism: To make it change color upon encountering a specific condition (e.g., radiation, or simply a passive environmental check), the lyophilized capsule/paper is placed on the fabric. When the user sweats (or a small amount of water is added), the system rehydrates, initiates protein expression, and produces melanin. Fabric Integration: The melanin-producing cell-free reaction, when dried, can be integrated into fabric microfluidic channels or porous materials that allow the reaction to proceed and display the color change on the surface.
Achieving the Color Change
Light/Dark Shift: Because melanin is highly black, the fabric will turn dark in the areas where the freeze-dried system is activated. Enhanced Functionality: The produced melanin acts as a photo-protective layer, enhancing the wearable’s ability to absorb UV or ionizing radiation.
Conceptual Steps for Assembly
Engineered Cells: Create a cell-free reaction designed to express tyrosinase from C. neoformans.
Lyophilize: Freeze-dry the lysate in a protective sugar matrix (e.g., trehalose) on a small membrane or paper patch.
Encapsulate: Place this paper patch inside a flexible, sweat-permeable, transparent, or breathable membrane patch.
Wear: Apply to the skin (e.g., as a "smart tattoo", smart clothing layer or spray a coat on a wearable 3d printed piece).
Activate: Upon moisture/sweat, the melanin produces color, turning the patch dark brown or black.
Sources
- Producing a melanin-rich topical from genes present in Cryptococcus neoformans Fungi to prevent cell irradiation in astronauts
- Radiation protection and structural stability of fungal melanin polylactic acid biocomposites in low Earth orbit -Melanin-inspired conductive thin films for multimodal-sensing wearable on-skin electronics -Visualization and documentation of capsule and melanin production in Cryptococcus neoformans
- What societal challenge or market need will this address?
In a world where we are encountering so much terrestrial pollution from nearby wars and destruction of the environment, radiation pollution and damage is becoming a real every day problem. In addition, this idea will be useful for astronauts for space exploration, nuclear plant workers and hospital radiology workers. Unfortunately, citizens encounter problems from radiation exposure on a daily basis too.
- How do you envision addressing the limitation of cell-free reactions (e.g., activation with water, stability, one-time use)?
| Challenge/limitations | Solution |
|---|---|
| High temperature denaturation | Use lyoprotectants (trehalose, maltodextrin) |
| Sensitivity issues | Use CRISPR-based tools & toehold switches |
| Rehydration failure | Integrate into hydrogels or porous substrates |
| Evaporation/Stability | Lamination and sealed reaction modules |
Homework question from Ally
Georg asked me to use my critical side to the homework question, haha. My comment was that I do not believe in space colonisation by humans.
- Provide background information that describes the space biology question or challenge you propose to address. Explain why this topic is significant for humanity, relevant for space exploration, and scientifically interesting. (Maximum 100 words)
My challenge is to find ways to warn for radiation exposure as well as ways of protection from radiation which is a terrestrial as well an extraterrestrial issue for humans and more than human ecosystems. Some ecosystems and kingdoms have developed mechanisms for protection against radiation and oxidative stress, like radiotropic fungi! Radiotropic fungi secrete melanin as a protective mechanism agains oxidative stress and radiation exposure. We can extract melanin from fungi and freeze dry it.
- Name the molecular or genetic target that you propose to study. Examples of molecular targets include individual genes and proteins, DNA and RNA sequences, or broader -omics approaches. (Maximum 30 words)
We want to target melanin producing proteins.
Describe how your molecular or genetic target relates to the space biology question or challenge your proposal addresses. (Maximum 100 words)
Clearly state your hypothesis or research goal and explain the reasoning behind it (Maximum 150 words)
Outline your experimental plan - identify the sample(s) you will test in your experiment, including any necessary controls, the type of data or measurements that will be collected, etc. (Maximum 100 words)
Homework Part B - Individual Final Project Report
Mandatory for Committed Listeners and MIT/Harvard Students. We’d like students to start exploring their final project in depth this week! The minimum requirement is filling out Aim 1.
See this link if you want examples of what you can include in your answers for each question. Note: the question numbers do not line up, and you can ignore any word requirements in this linked document.
Look here for my individual final project page!