Week 11 — Bioproduction & Cloud Labs


WEEK 11 HW
This week examines how modern bioproduction pipelines, from strain engineering to fermentation and downstream processing, are increasingly designed, executed, and optimized through cloud lab platforms and automation — enabling remote, high-throughput, and reproducible synthetic biology at industrial scale.


Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork
How it started….


How it evolved…
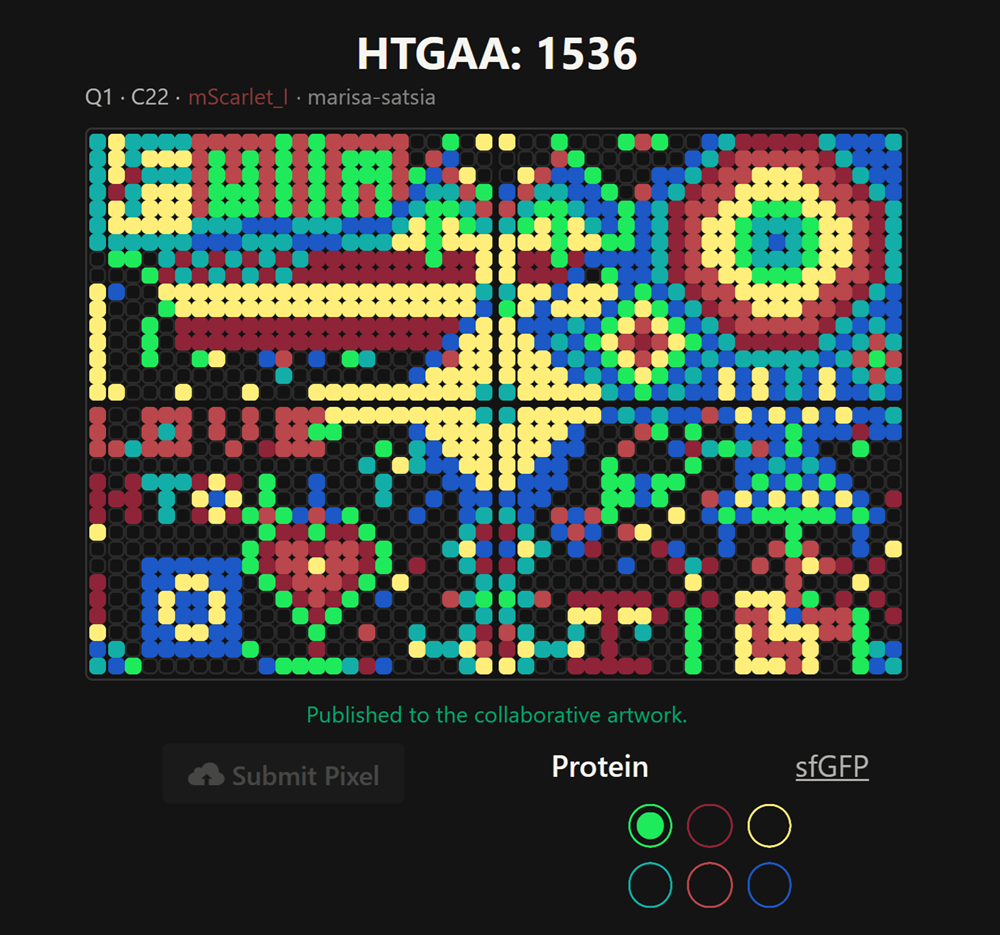


How it ended… Highjacked!
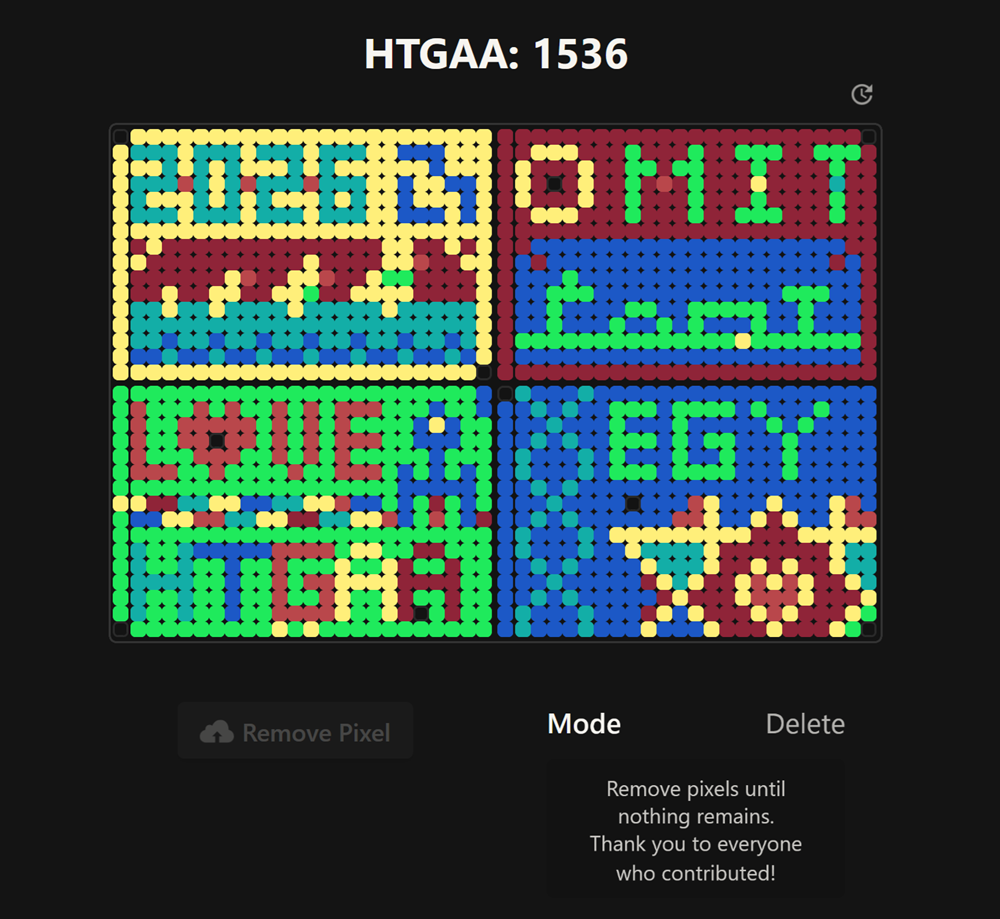

Contribute at least one pixel to this global artwork experiment before the editing ends on Sunday 4/19 at 11:59 PM EST.
+A personalized URL was sent to the email address associated with your Discourse account, and you can discuss the artwork on the Discourse. +If you did not have a chance to contribute, it’s okay, just make sure you become a TA this fall! 😉
Make a note on your HTGAA webpages including:
+what you contributed to the community bioart project (e.g., “I made part of the DNA on the bottom right plate”) +what you liked about the project, and +what about this collaborative art experiment could be made better for next year.
I made the part of the DNA in the middle of the upper 2 plates but unfortunately it was highjacked as you can see on the top! I really enjoyed this collaborative part.
Part B: Cell-Free Protein Synthesis | Cell-Free Reagents
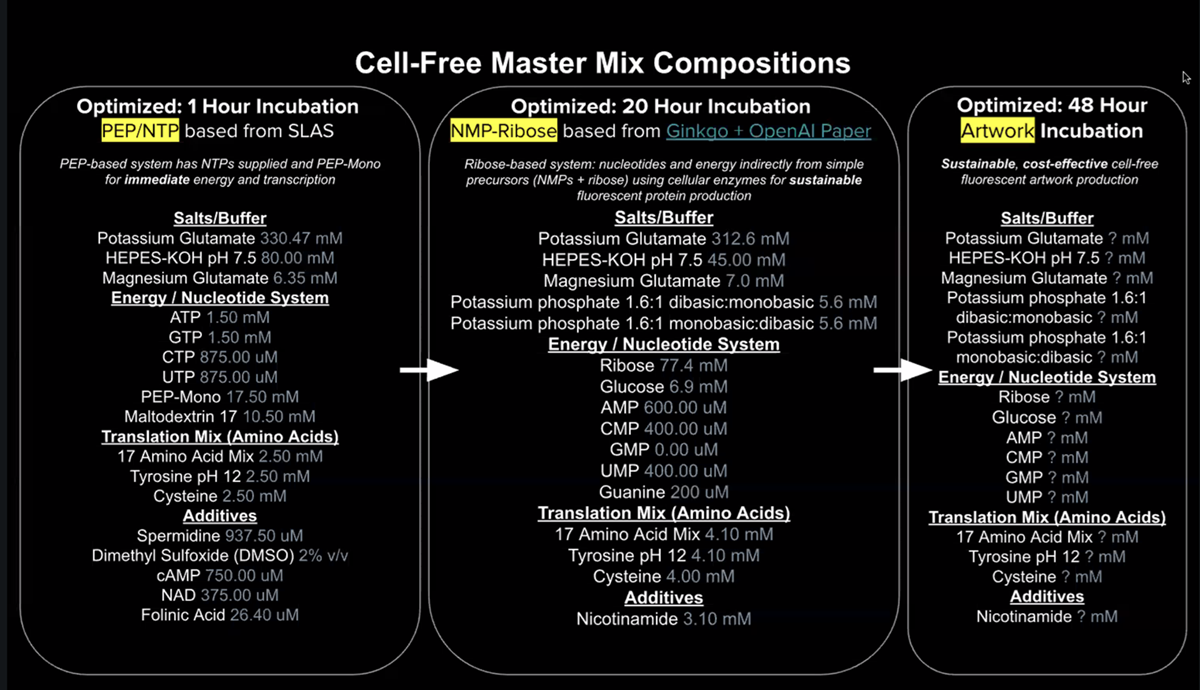

- Referencing the cell-free protein synthesis reaction composition (the middle box outlined in yellow on the image above, also listed below), provide a 1-2 sentence description of what each component’s role is in the cell-free reaction.
E. coli Lysate
- BL21 (DE3) Star Lysate (includes T7 RNA Polymerase) ->
Salts/Buffer
- Potassium Glutamate
- HEPES-KOH pH 7.5
- Magnesium Glutamate
- Potassium phosphate monobasic
- Potassium phosphate dibasic
Energy / Nucleotide System
- Ribose
- Glucose
- AMP
- CMP
- GMP
- UMP
- Guanine
Translation Mix (Amino Acids)
- 17 Amino Acid Mix
- Tyrosine
- Cysteine
Additives
- Nicotinamide->
Backfill
- Nuclease Free Water->
- Describe the main differences between the 1-hour optimized PEP-NTP master mix and the 20-hour NMP-Ribose-Glucose master mix shown in the Google Slide above. (2-3 sentences)
Main differences:
Speed: PEP-NTP is designed for fast, high-intensity, short-term production (1-hour), whereas NMP-Ribose-Glucose is designed for slow, sustained, long-term production (20-hours). Cost & Economics: The 1-hour PEP-NTP mix is more expensive due to substrates like phosphoenolpyruvate, while the 20-hour mix is more economical, using Glucose/Ribose. Metabolic Byproducts: The 20-hour mix often has better control over metabolic byproducts (such as inorganic phosphate) due to its slower, more balanced energy consumption, leading to longer reaction stability.
Bonus question: How can transcription occur if GMP is not included but Guanine is?
Part C: Planning the Global Experiment | Cell-Free Master Mix Design
- Given the 6 fluorescent proteins we used for our collaborative painting, identify and explain at least one biophysical or functional property of each protein that affects expression or readout in cell-free systems. (Hint: options include maturation time, acid sensitivity, folding, oxygen dependence, etc) (1-2 sentences each)
Superfolder GFP is a basic (constitutively fluorescent) green fluorescent protein published in 2005, derived from Aequorea victoria. It is reported to be a very rapidly-maturing weak dimer. Engineered for robust folding, even under non-optimal conditions, making it highly reliable in cell-free systems.
It has slow maturation and lower photostability, which delays fluorescence signal and reduces effective brightness in short or energy-limited cell-free reactions. Slow maturation can delay readout despite successful expression.
It is commonly used as a fusion tag for tracking protein localization and dynamics, particularly in long-term imaging due to its high photostability. mKO2 is known for relatively rapid maturation compared to other fluorescent proteins, which is critical for studies requiring fast protein turnover tracking. It is generally stable within a wide pH range, though like most fluorescent proteins, its fluorescence decreases in highly acidic environments. It is often preferred over dsRed-type proteins for studies in slightly acidic compartments, although it is not specifically designed as a strict pH sensor. The protein is highly monomeric and folds well, making it suitable for fusions to proteins where dimerization might cause dysfunction or localization issues.
From google patents
High quantum yield and fast maturation. Faster but limited still folding limited compared to GFP making its fluorescence more sensitive to translation kinetics and folding efficiency. mTurquoise2 is highly stable in acidic environments, exhibiting very low acid sensitivity.
From Systematic characterization of maturation time of fluorescent proteins in living cells.
It has moderate acid sensitivity, with a high tolerance to acidic environments. A faster-maturing variant than mScarlet (T74I mutation) matures in roughly 25–36 minutes at 37°C. mScarlet-I is generally preferred for fusions requiring fast-maturing, bright red tags.
Displays oxygen-dependent chromophore formation, so fluorescence requires sufficient oxygen availability, which can be limiting in dense or sealed reactions. Its performance is typically influenced by trade-offs between brightness, folding efficiency, and maturation kinetics, meaning signal output depends strongly on how well it folds and matures in the cell-free environment.
The amino acid sequences are shown in the HTGAA Cell-Free Benchling folder.
Create a hypothesis for how adjusting one or more reagents in the cell-free mastermix could improve a specific biophysical or functional property you identified above, in order to maximize fluorescence over a 36-hour incubation. Clearly state the protein, the reagent(s), and the expected effect.
The second phase of this lab will be to define the precise reagent concentrations for your cell-free experiment. You will be assigned artwork wells with specific fluorescent proteins and receive an email with instructions this week (by April 24). You can begin composing master mix compositions here.
The final phase of this lab will be analyzing the fluorescence data we collect to determine whether we can draw any conclusions about favorable reagent compositions for our fluorescent proteins. This will be due a week after the data is returned (date TBD!). The reaction composition for each well will be as follows:
6 μL of Lysate 10 μL of 2X Optimized Master Mix from above 2 μL of assigned fluorescent protein DNA template 2 μL of your custom reagent supplements
Total: 20 μL reaction
PART D - BONUS- BUILD A CLOUD LAB


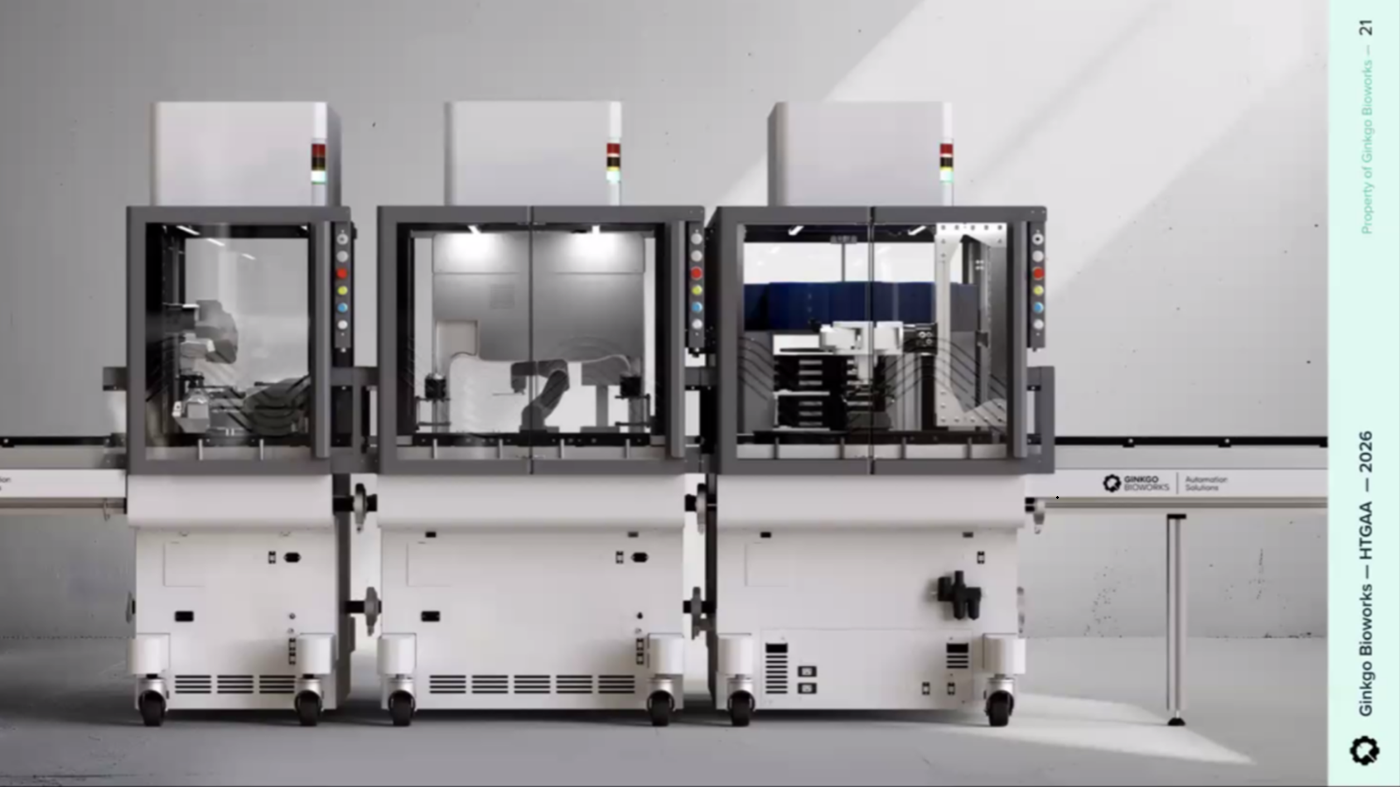
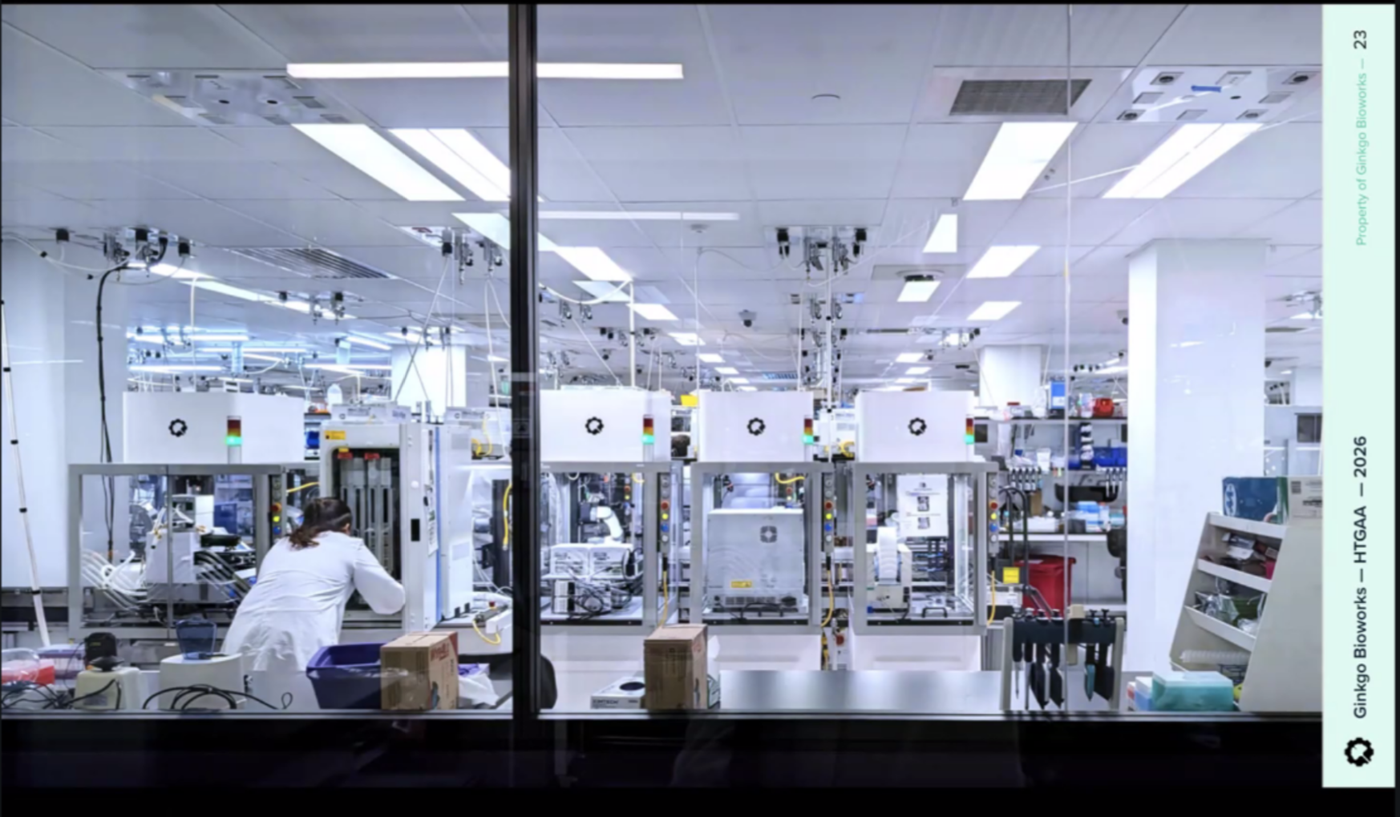
Ginkgo Nebula Cloud Laboratory Rendering, 2025
Use this simulation tool to create an interesting looking cloud lab out of the Ginkgo Reconfigurable Automation Carts. This is just a minimal implementation so far, but I would love to see some fun designs!