Group Final Project
WEEK 4 PART D Group Project- Brainstorm on Bacteriophage Engineering
1. Find a group of ~3–4 students
I formed a group with the following people in our node :)
and weee have a new member!
2. Read through the Phage Reading material listed under “Reading & Resources” below.
3. Review the Bacteriophage Final Project Goals for engineering the L Protein:
-Increased stability (easiest) -> we chose this option!
-Higher titers (medium)
-Higher toxicity of lysis protein (hard)
4. Brainstorm Session
+ Choose one or two main goals from the list that you think you can address computationally (e.g., “We’ll try to stabilize the lysis protein,” or “We’ll attempt to disrupt its interaction with E. coli DnaJ.”).
+ Write a 1-page proposal (bullet points or short paragraphs) describing:
Which tools/approaches from recitation you propose using (e.g., “Use Protein Language Models to do in silico mutagenesis, then AlphaFold-Multimer to check complexes.”).
+ Why do you think those tools might help solve your chosen sub-problem?
+ Name one or two potential pitfalls (e.g., “We lack enough training data on phage–bacteria interactions.”).
+ Include a schematic of your pipeline.


From Diogos documentation
This resource may be useful: HTGAA Protein Engineering Tools.
5. Each individually put your plan on your HTGAA website
+ Include your group’s short plan for engineering a bacteriophage
From my homework on Part C of week 5 Protein Design Part 2: Final project- L-Protein Mutants
The objective of this assignment is to improve the stability and auto-folding of the lysis protein of a MS2-phage. This mechanism is key to the understanding of how phages can potentially solve antibiotic-resistance.
Ms2 Lysis Protein sequence
Note: Lysis protein contains a soluble N-terminal domain followed by a transmembrane protein (blue/last 35 residues). Transmembrane protein affects the lysis activity. The soluble domain (green) is the domain responsible for interaction with DnaJ.
After running the MS2 Lysis protein sequence on its own on Alphafold:
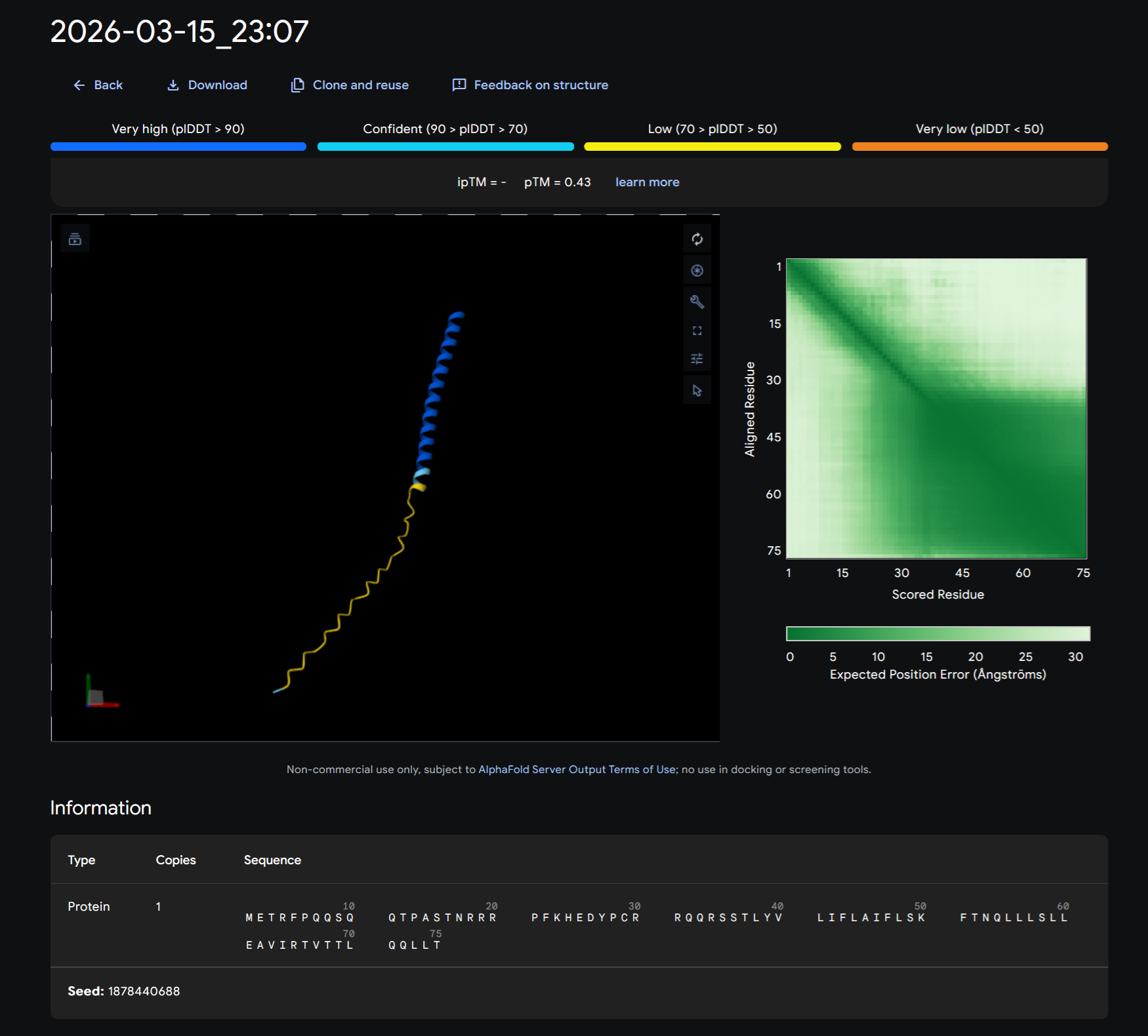

- Redered interaction of MS2 lysis protein with DnaJ
Dna J againts MS2 Lysis protein-> mutant
Dna J sequence


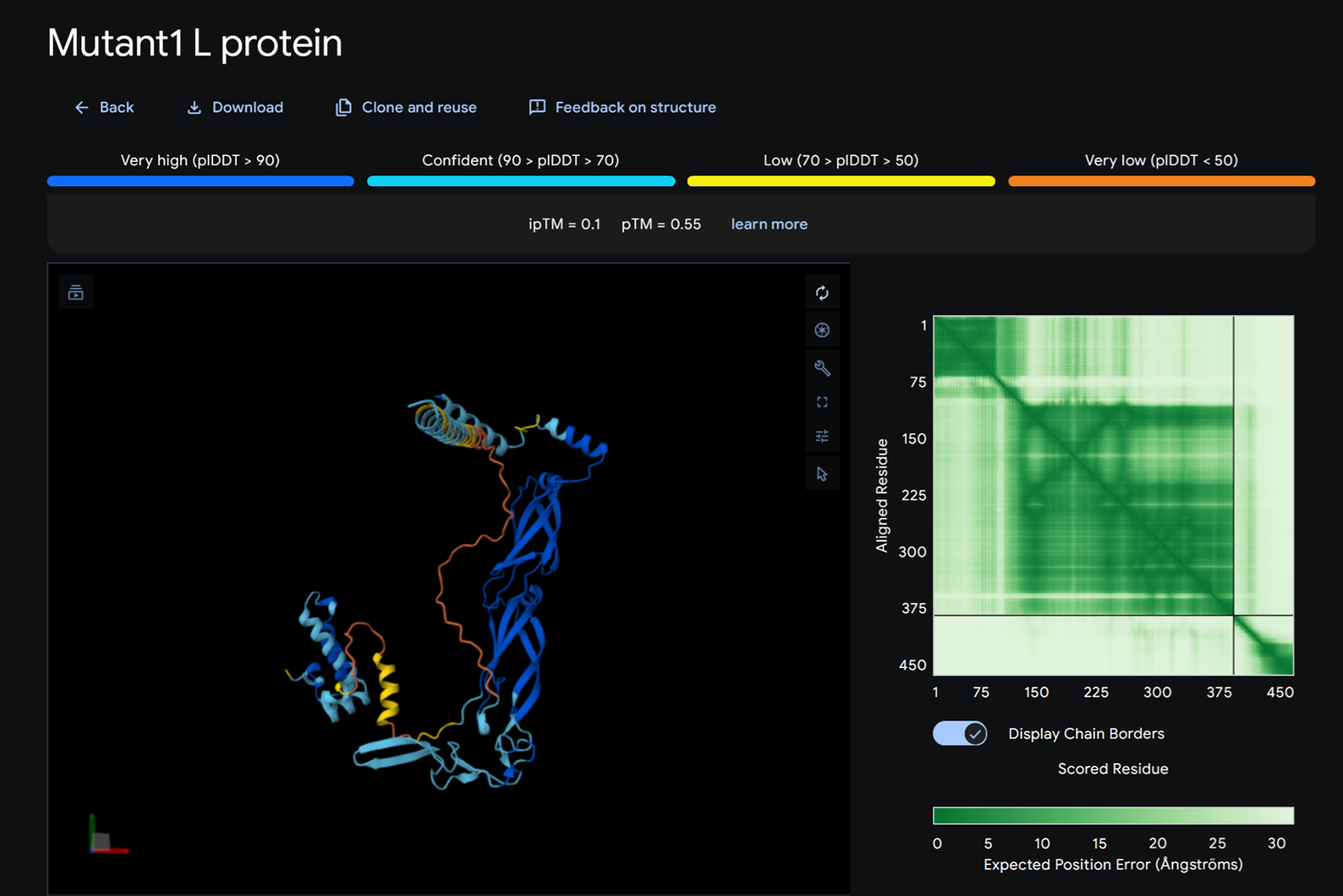

- Detail where we can see no bonds between DnaJ and L protein Mutant
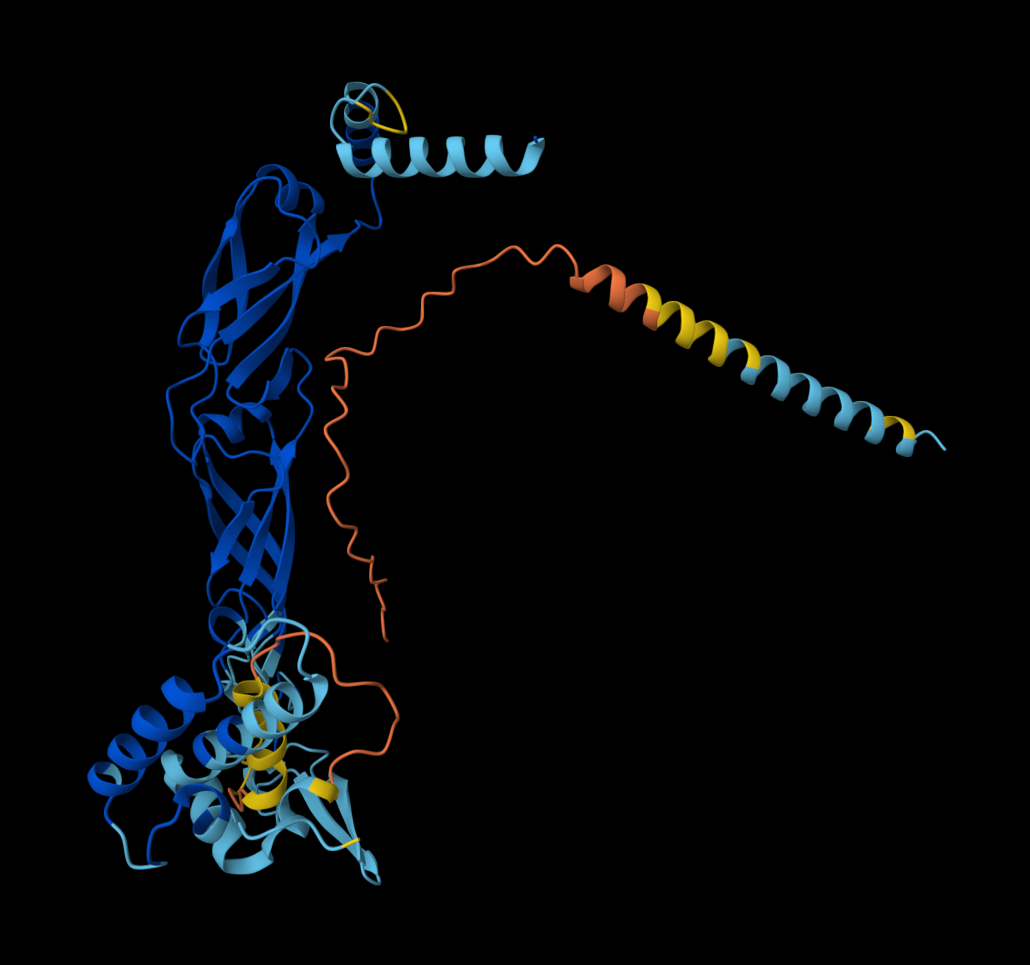

- Multimer made up of 8 Mutant L proteins. The aggregation seems close to what would be expected of wild type L Protein.