Week 2 HW: DNA READ, WRITW AND EDIT
Part 1: Benchling & In-silico Gel Art
Make a free account at benchling.com
Import the Lambda DNA.
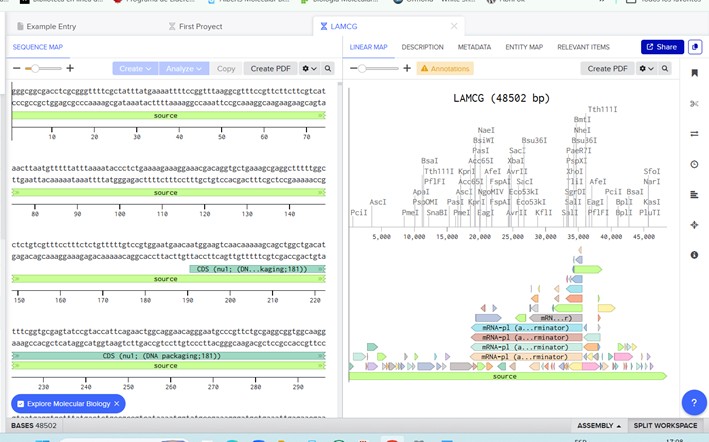

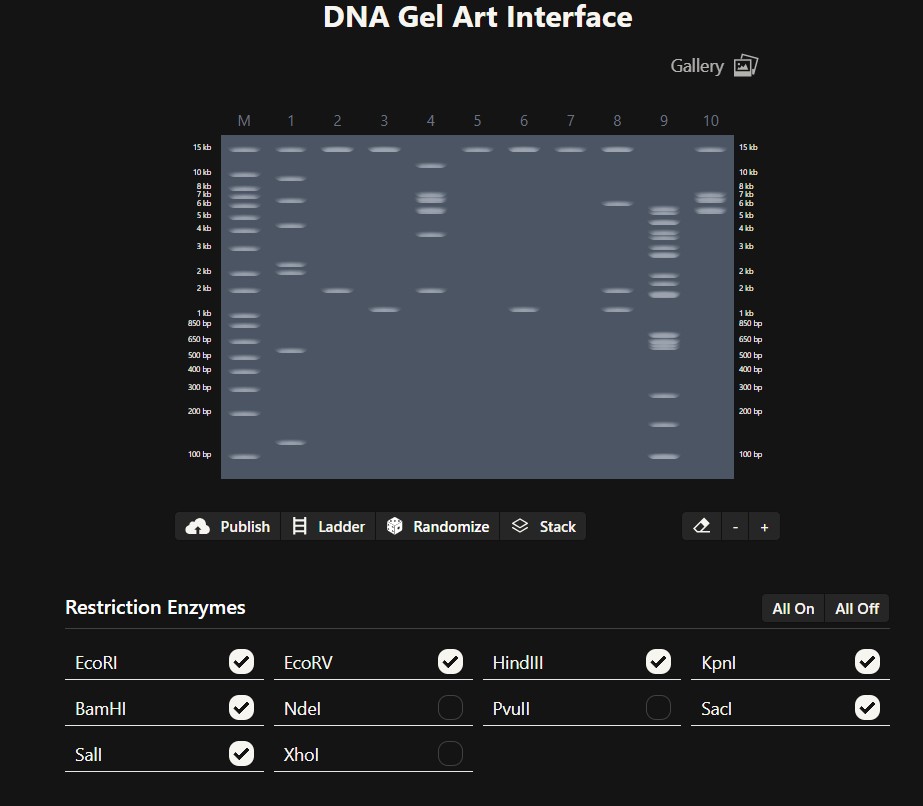
Simulate Restriction Enzyme Digestion with the following Enzymes:
EcoRI
HindIII
BamHI
KpnI
EcoRV
SacI
SalI
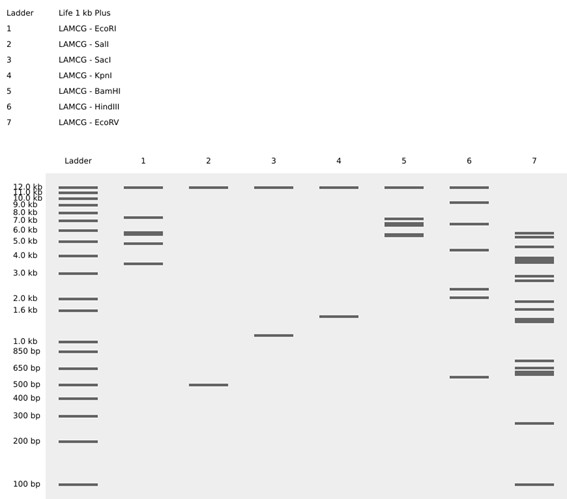

Create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks.
You might find Ronan’s website a helpful tool for quickly iterating on designs!
Part 3: DNA Design Challenge
3.1 Choose your protein
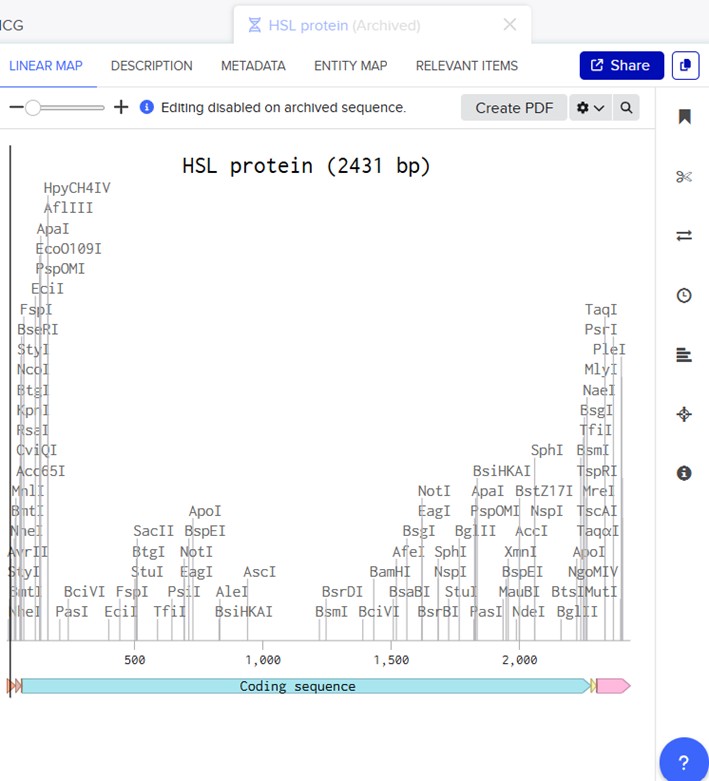
Which protein have you chosen and why? I have choose the Hormone-sensitive lipase. I chose because It is an important enzyme whose deficiency causes health problems, mainly in the liver (Xia et al. 2017).
Xia B, Cai GH, Yang H, Wang SP, Mitchell GA, Wu JW. Adipose tissue deficiency of hormone-sensitive lipase causes fatty liver in mice. PLoS Genet. 2017 Dec 12;13(12):e1007110. doi: 10.1371/journal.pgen.1007110.
sp|P54310.2|LIPS_MOUSE RecName: Full=Hormone-sensitive lipase; Short=HSL; AltName: Full=Monoacylglycerol lipase LIPE; AltName: Full=Retinyl ester hydrolase; Short=REH MDLRTMTQSLVTLAEDNMAFFSSQGPGETARRLSNVFAGVREQALGLEPTLGQLLGVAHHFDLDTETPAN GYRSLVHTARCCLAHLLHKSRYVASNRKSIFFRASHNLAELEAYLAALTQLRAMAYYAQRLLTINRPGVL FFEGDEGLTADFLQEYVTLHKGCFYGRCLGFQFTPAIRPFLQTLSIGLVSFGEHYKRNETGLSVTASSLF TGGRFAIDPELRGAEFERIIQNLDVHFWKAFWNITEIEVLSSLANMASTTVRVSRLLSLPPEAFEMPLTS DPRLTVTISPPLAHTGPAPVLARLISYDLREGQDSKVLNSLAKSEGPRLELRPRPHQAPRSRALVVHIHG GGFVAQTSKSHEPYLKNWAQELGVPIFSIDYSLAPEAPFPRALEECFFAYCWAVKHCDLLGSTGERICLA GDSAGGNLCITVSLRAAAYGVRVPDGIMAAYPVTTLQSSASPSRLLSLMDPLLPLSVLSKCVSAYSGTEA EDHFDSDQKALGVMGLVQRDTSLFLRDLRLGASSWLNSFLELSGRKPQKTTSPTAESVRPTESMRRSVSE AALAQPEGLLGTDTLKKLTIKDLSNSEPSDSPEMSQSMETLGPSTPSDVNFFLRPGNSQEEAEAKDEVRP MDGVPRVRAAFPEGFHPRRSSQGVLHMPLYTSPIVKNPFMSPLLAPDSMLKTLPPVHLVACALDPMLDDS VMFARRLRDLGQPVTLKVVEDLPHGFLSLAALCRETRQATEFCVQRIRLILTPPAAPLN
From NCBI: https://www.ncbi.nlm.nih.gov/protein/P54310.2?report=fasta
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence
Determine the nucleotide sequence that corresponds to the protein sequence you chose above
reverse translation of sp|P54310.2|LIPS_MOUSE RecName: Full=Hormone-sensitive lipase; Short=HSL; AltName: Full=Monoacylglycerol lipase LIPE; AltName: Full=Retinyl ester hydrolase; Short=REH to a 2277 base sequence of most likely codons. atggatctgcgcaccatgacccagagcctggtgaccctggcggaagataacatggcgttt tttagcagccagggcccgggcgaaaccgcgcgccgcctgagcaacgtgtttgcgggcgtg cgcgaacaggcgctgggcctggaaccgaccctgggccagctgctgggcgtggcgcatcat tttgatctggataccgaaaccccggcgaacggctatcgcagcctggtgcataccgcgcgc tgctgcctggcgcatctgctgcataaaagccgctatgtggcgagcaaccgcaaaagcatt ttttttcgcgcgagccataacctggcggaactggaagcgtatctggcggcgctgacccag ctgcgcgcgatggcgtattatgcgcagcgcctgctgaccattaaccgcccgggcgtgctg ttttttgaaggcgatgaaggcctgaccgcggattttctgcaggaatatgtgaccctgcat aaaggctgcttttatggccgctgcctgggctttcagtttaccccggcgattcgcccgttt ctgcagaccctgagcattggcctggtgagctttggcgaacattataaacgcaacgaaacc ggcctgagcgtgaccgcgagcagcctgtttaccggcggccgctttgcgattgatccggaa ctgcgcggcgcggaatttgaacgcattattcagaacctggatgtgcatttttggaaagcg ttttggaacattaccgaaattgaagtgctgagcagcctggcgaacatggcgagcaccacc gtgcgcgtgagccgcctgctgagcctgccgccggaagcgtttgaaatgccgctgaccagc gatccgcgcctgaccgtgaccattagcccgccgctggcgcataccggcccggcgccggtg ctggcgcgcctgattagctatgatctgcgcgaaggccaggatagcaaagtgctgaacagc ctggcgaaaagcgaaggcccgcgcctggaactgcgcccgcgcccgcatcaggcgccgcgc agccgcgcgctggtggtgcatattcatggcggcggctttgtggcgcagaccagcaaaagc catgaaccgtatctgaaaaactgggcgcaggaactgggcgtgccgatttttagcattgat tatagcctggcgccggaagcgccgtttccgcgcgcgctggaagaatgcttttttgcgtat tgctgggcggtgaaacattgcgatctgctgggcagcaccggcgaacgcatttgcctggcg ggcgatagcgcgggcggcaacctgtgcattaccgtgagcctgcgcgcggcggcgtatggc gtgcgcgtgccggatggcattatggcggcgtatccggtgaccaccctgcagagcagcgcg agcccgagccgcctgctgagcctgatggatccgctgctgccgctgagcgtgctgagcaaa tgcgtgagcgcgtatagcggcaccgaagcggaagatcattttgatagcgatcagaaagcg ctgggcgtgatgggcctggtgcagcgcgataccagcctgtttctgcgcgatctgcgcctg ggcgcgagcagctggctgaacagctttctggaactgagcggccgcaaaccgcagaaaacc accagcccgaccgcggaaagcgtgcgcccgaccgaaagcatgcgccgcagcgtgagcgaa gcggcgctggcgcagccggaaggcctgctgggcaccgataccctgaaaaaactgaccatt aaagatctgagcaacagcgaaccgagcgatagcccggaaatgagccagagcatggaaacc ctgggcccgagcaccccgagcgatgtgaacttttttctgcgcccgggcaacagccaggaa gaagcggaagcgaaagatgaagtgcgcccgatggatggcgtgccgcgcgtgcgcgcggcg tttccggaaggctttcatccgcgccgcagcagccagggcgtgctgcatatgccgctgtat accagcccgattgtgaaaaacccgtttatgagcccgctgctggcgccggatagcatgctg aaaaccctgccgccggtgcatctggtggcgtgcgcgctggatccgatgctggatgatagc gtgatgtttgcgcgccgcctgcgcgatctgggccagccggtgaccctgaaagtggtggaa gatctgccgcatggctttctgagcctggcggcgctgtgccgcgaaacccgccaggcgacc gaattttgcgtgcagcgcattcgcctgattctgaccccgccggcggcgccgctgaac
I used the reverse translation tool of the website bioinformatics.org.
3.3. Codon optimization
Once a nucleotide sequence of your protein is determined, you need to codon optimize your sequence. You may, once again, utilize google for a “codon optimization tool”. In your own words, describe why you need to optimize codon usage.
To increase the expression of the protein of interest
Which organism have you chosen to optimize the codon sequence for and why?
I chose the mouse Mus musculus because they have a high degree of similarity to the human genome. The articles I consulted describe tests performed on mice.
A T G G A T C T G C G C A C C A T G A C C C A G A G C C T G G T G A C C C T G G C G G A A G A T A A C A T G G C G T T T T T T A G C A G C C A G G G C C C G G G C G A A A C C G C G C G C C G C C T G A G C A A C G T G T T T G C G G G C G T G C G C G A A C A G G C G C T G G G C C T G G A A C C G A C C C T G G G C C A G C T G C T G G G C G T G G C G C A T C A T T T T G A T C T G G A T A C C G A A A C C C C G G C G A A C G G C TA T C G C A G C C T G G T G C A T A C C G C G C G C T G C T G C C T G G C G C A T C T G C T G C A T A A A A G C C G C T A T G T G G C G A G C A A C C G C A A A A G C A T T T T T T T T C G C G C G A G C C A T A A C C T G G C G G A A C T G G A A G C G T A T C T G G C G G C G C T G A C C C A G C T G C G C G C G A T G G C G T A T T A T G C G C A G C G C C T G C T G A C C A T T A A C C G C C C G G G C G T G C T G T T T T T T T G AA G G C G A T G A A G G C C T G A C C G C G G A T T T T C T G C A G G A A T A T G T G A C C C T G C A T A A A G G C T G C T T T T A T G G C C G C T G C C T G G G C T T T C A G T T T A C C C C G G C G A T T C G C C C G T T T C T G C A G A C C C T G A G C A T T G G C C T G G T G A G C T T T G G C G A A C A T T A T A A A C G C A A C G A A A C C G G C C T G A G C G T G A C C G C A G C C T G T T T A C C G G C G G C C G CT T T G C G A T T G A T C C G G A A C T G C G C G G C G A A T T T G A A C G C A T T A T T C A G A A C C T G G A T G T G C A T T T T T G G A A A G C G T T T T G G A A C A T T A C C G A A A T T G A A G T G C T G A G C A G C C T G G C G A A C A T G G C G A G C A C C A C C G T G C G C G T G A G C C G C C T G C T G A G C C T G C C G C C G G A A G C G T T T G A A A T G C C G C T G A C C GT G A C C A T T A G C C C G C C G C T G G C G C A T A C G G C C C G G C G C G G T G C T G G C G C G C C T G A T A G C T A T G A T C T G C G C G A A G G C C A G G A T A G C A A A G T G C T G A A C A G C C T G G C G A A A A G C G A A G G C C C G C G C C T G G A A C T G C G C C C G C C C G C C G C A T C A G G C G C C G C A G C C G C G C G C T G G T G G T G C A T A T T C A T G G C G G C G G C T T T G T G G C G C A G A CC A G C A A A A G C C A T G A A C C G T A T C T G A A A A A C T G G G C G C A G G A A C T G G G C G T G C C G A T T T T T A G C A T T G A T T A T A G C C T G G C G C C G A A G C G C C G T T T C C G C G C G C T G G A A G A A T G C T T T T T T G C G T A T T G C T G G G C G G T G A A A C A T T G C G A T C T G C T G G G C A G C A C C G G C G A A C G C A T T T G C C T G G C G G G C G A T A G C G C G G G C G A A C C T GT G C A T T A C C G T G A G C C T G C G C G G C G G C G T A T G G C G T G C G C G T G C C G G A T G G C A T T A T G G C G G C G T A T C C G G T G A C C A C C C T G C A G A G C G C G A G C C C G A G C C G C C T G C T G A G C C T G A T G G A T C C G C T G C C G C T G A G C G T G C T G A G C A A A T G C G T G A G C G C G T A T A G C G G C A C C G A A G C G A A G A T C A T T T T G A T A G C G A T C A G A A A GC G C T G G G C G T G A T G G G C C T G G T G C A G C G C G A T A C C A G C C T G T T T C T G C G C G A T C T G C G C C T G G G C G C G A G C A G C T G G C T G A A C A G C T T T C T G G A A C T G A G C G G C C G C A A A C C G C A G A A A A C C A C C A G C C C G A C C G C G A A A G C G T G C G C C C G A C C G A A A A G C A T G C G C C G C A G C G T G A G C G A A G C G G C G C T G G C G C A G C C G G A A G G C C T G C T G G GC A C C G A T A C C C T G A A A A A A A C T G A C C A T T A A A G A T C T G A G C A A C A G C G A A C C G A G C G A T A G C C C G G A A A T G A G C C A G A G C A T G G A A A C C C T G G G C C C G A G C A C C C C G A G C G A T G T G A A C T T T T T T C T G C G C C C G G G C A A C A G C C A G G A A G A A A G C G G A A G C G A A A G A T G A A G T G C G C C C G A T G G A T G G C G T G C C G C G C G T G C G C G C G C G G C G T T T C C GG A A G G C T T T C A T C C G C G C C G C A G C A G G G C G T G C T G C A T A T G C C G C T G T A T A C C A G C C C G A T T G T G A A A A A A C C C G T T T A T G A G C C C G C T G C T G G C G C C G G A T A G C A T G C T G A A A A A C C C T G C C G C C G G T G C A T C C G A T G C T G G A T G A T A G C G T G A T G T T T G C G C G C C G C C T G C G C G A T C T G G G C C A G C C G GT G A C C C T G A A A G T G G T G G A A G A T C T G C C G C A T G G C T T T C T G A G C C T G G C G G C G C T G T G C C G C G A A A C C C G C C A G G C G A C C G A A T T T T G C G T G C A G C G C A T T C G C C T G A T T C T G A C C C C G C C G G C G G C G C C G C T G A A C
I used the VectorBuilder website optimization tool, avoiding the Bbs I, Bbv I, and Bsa I recognition sites.
3.4. You have a sequence! Now what?
What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both.
I’m going to design the protein and find out if I can order the plasmid with my design from a company in my country; I might have to import it, which requires several permits. Then I’m going to transform bacteria with this DNA in the lab, to clone it and have bacterial cultures that can produce this protein.
3.5. How does it work in nature/biological systems?
- Describe how a single gene codes for multiple proteins at the transcriptional level.
Because after transcription, splicing causes several isoforms to be generated from the same genetic code.
- Try aligning the DNA sequence, the transcribed RNA, and also the resulting translated Protein!!! See example below.
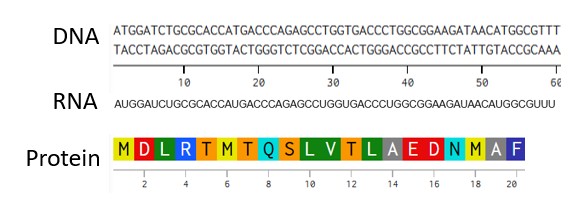

Part 4: Prepare a Twist DNA Synthesis Order
4.1. Create a Twist account and a Benchling account
I was able to create an account on Benchling but not on Twist. I live in Ecuador and it seems to be blocked in my country.
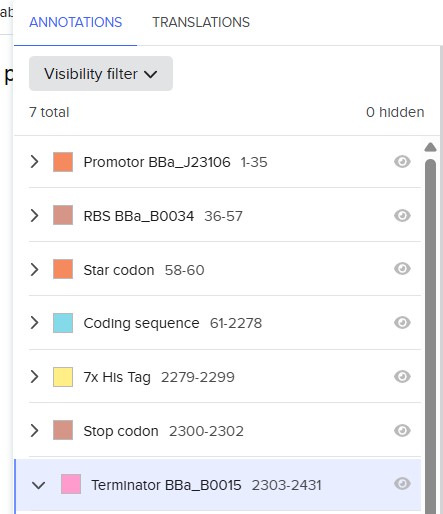
4.2. Build Your DNA Insert Sequenc
https://benchling.com/s/seq-VM2nGvwYWi5G4MEMBUeg?m=slm-1mugup1rWutlh5MYbTTq
From 4.3 to 4.6 I couldn’t make them because I can’t use Twist

Part 5: DNA Read/Write/Edit
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why?
The DNA of the venomous snake Lachesis acrochorda to learn all about its genome and conduct research on its evolutionary history and the genes for venom production
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
I would use Illumina
Also answer the following questions:
- Is your method first-, second- or third-generation or other? How so?
It is a more modern technology and is faster and cheaper than first generation techniques.
- What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps.
For Illumina, short fragments are needed.
First, the DNA is fragmented using restriction enzymes.
Second, the ends are repaired and adenylated.
Third, ligation is performed by adding adapters.
Fourth, PCR is used to amplify the fragments.
- What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?
Steps
First: fragmentation and ligation.
Second: clusters are generated by anchoring the DNA to a flow cell.
Third: sequencing begins by adding Illumina polymerases and fluorescent nucleotides.
Depending on the nucleotide that passes through the cell, it glows a different color; photos are taken and processed in the software.
- What is the output of your chosen sequencing technology?
The sequence of the fragments is obtained, which then need to be mapped.
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why?
Create antibodies that neutralize the myotoxins and neurotoxins of the snake Lachesis acrochorda
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
I would use Microarray synthesis. it is cheap and allows the synthesis of thousands of sequences.
Also answer the following questions:
- What are the essential steps of your chosen sequencing methods?
Steps
First: Design in the software.
Second: Place chemical linkers on the chips.
Third: Load the reservoirs with nucleotides.
Fourth: When the nucleotide binds to the forming chain, wash away the excess reagents.
Fifth: Remove the protecting group to add the next nucleotide and repeat until the required length is reached.
Sixth: Cut and collect the sequences.
- What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability?
It requires further amplification due to the small amount produced and cannot synthesize very long sequences.
5.3 DNA Edit
(i) What DNA would you want to edit and why?
I would like to create potatoes that contain anthocyanins. Potatoes are a widely consumed food, and this antioxidant could be added to supplement the diet.
(ii) What technology or technologies would you use to perform these DNA edits and why?
The potato is a dicotyledonous plant, so I would use Agrobacterium tumefaciens to introduce the gene. It’s a proven and widely used method.
Also answer the following questions:
- How does your technology of choice edit DNA? What are the essential steps?
First: Introduce the gene into the vector, which would be the Ti plasmid of the bacterium. The plasmid will contain genes for anthocyanin production, as many as possible, and an antibiotic resistance gene for subsequent selection.
Second: Transform the bacterium with the plasmid.
Third: Expose the bacteria to plant cells in vitro to infect them with the plasmid.
Fourth: Select the transformed cells, which are those that grow in a medium containing antibiotics.
- What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
The Ti plasmid is designed by introducing the gene of interest, in software such as Benchling.
3.What are the limitations of your editing methods (if any) in terms of efficiency or precision?
It’s very imprecise. it’s impossible to pinpoint exactly where in the genome the gene will be inserted. Consequently, the gene might not be expressed or it could affect other genes.