Week 3: Lab Automation
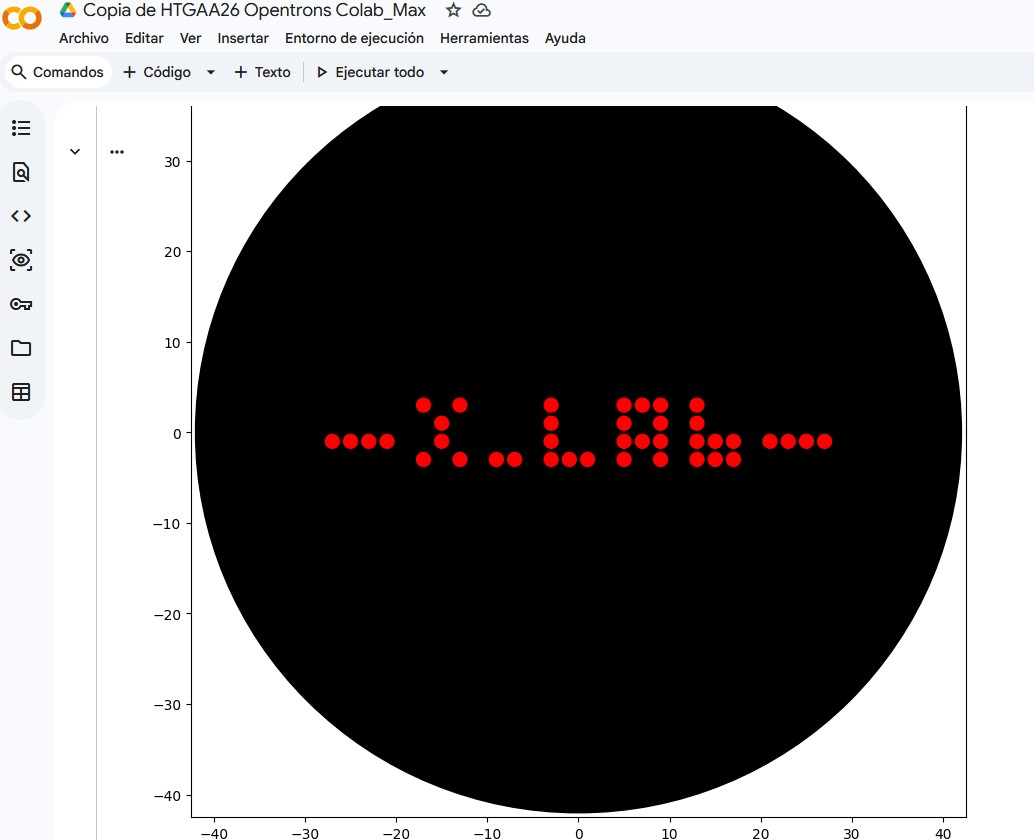

I. Assignment: Python Script for Opentrons Artwork: Max_XLAB_pattern
metadata = {# see https://docs.opentrons.com/v2/tutorial.html#tutorial-metadata }
'author': 'MaxMatus',
'protocolName': 'Max_XLab_pattern',
'description': 'XLab_pattern',
'source': 'HTGAA 2026 Opentrons Lab',
'apiLevel': '2.20'
Robot deck setup constants
TIP_RACK_DECK_SLOT = 9 COLORS_DECK_SLOT = 6 AGAR_DECK_SLOT = 5 PIPETTE_STARTING_TIP_WELL = ‘A1’
well_colors = { ‘A1’ : ‘Red’, ‘B1’ : ‘Green’, ‘C1’ : ‘Orange’ }
def run(protocol):
Load labware, modules and pipettes
tips_20ul = protocol.load_labware(‘opentrons_96_tiprack_20ul’, TIP_RACK_DECK_SLOT, ‘Opentrons 20uL Tips’)
pipette_20ul = protocol.load_instrument(“p20_single_gen2”, “right”, [tips_20ul])
temperature_module = protocol.load_module(’temperature module gen2’, COLORS_DECK_SLOT)
temperature_plate = temperature_module.load_labware(‘opentrons_96_aluminumblock_generic_pcr_strip_200ul’,
‘Cold Plate’)
color_plate = temperature_plate
agar_plate = protocol.load_labware(‘htgaa_agar_plate’, AGAR_DECK_SLOT, ‘Agar Plate’)
center_location = agar_plate[‘A1’].top()
pipette_20ul.starting_tip = tips_20ul.well(PIPETTE_STARTING_TIP_WELL)
def location_of_color(color_string): for well,color in well_colors.items(): if color.lower() == color_string.lower(): return color_plate[well] raise ValueError(f"No well found with color {color_string}")
dispense_and_detach(pipette, 1, loc) def dispense_and_detach(pipette, volume, location): """ Move laterally 5mm above the plate (to avoid smearing a drop); then drop down to the plate, dispense, move back up 5mm to detach drop, and stay high to be ready for next lateral move. 5mm because a 4uL drop is 2mm diameter; and a 2deg tilt in the agar pour is >3mm difference across a plate. """ assert(isinstance(volume, (int, float))) above_location = location.move(types.Point(z=location.point.z + 5)) # 5mm above pipette.move_to(above_location) # Go to 5mm above the dispensing location pipette.dispense(volume, location) # Go straight downwards and dispense pipette.move_to(above_location) # Go straight up to detach drop and stay high
center_location = agar_plate[‘A1’].top()
cell_well = color_plate[‘A1’] # Changed ‘F1’ to ‘A1’ because F1 was not configured to have a color.
pipette_20ul.pick_up_tip()
X_LABpattern
X_LABpattern = [ # X (0, 4), (2, 4), # Fila superior (1, 3), # Diagonal bajando (1, 2), # Centro (0, 1), (2, 1), # Fila inferior
# First dot after X - moved from (3,1) to (4,1)
(4, 1),
# Second dot for separation - moved from (4,1) to (5,1)
(5, 1),
# L (shifted by 2 units from previous adjustment: 5+2=7)
(7, 4),
(7, 3),
(7, 2),
(7, 1), (8, 1), (9, 1),
# Espacio entre letras (col 10 vacía)
# A (shifted by 2 units from previous adjustment: 9+2=11)
(11, 1), (11, 2), (11, 3), (11, 4),
(12, 4),
(13, 4), (13, 3), (13, 2), (13, 1),
(12, 2),
# Espacio entre letras (col 14 vacía)
# b (minúscula) (shifted by 2 units from previous adjustment: 13+2=15)
(15, 1), (15, 2), (15, 3), (15, 4),
(16, 2),
(16, 1),
(17, 2),
(17, 1),
]
Determine pattern dimensions for scaling and centering
x_coords = [p[0] for p in X_LABpattern] y_coords = [p[1] for p in X_LABpattern]
min_x, max_x = min(x_coords), max(x_coords) min_y, max_y = min(y_coords), max(y_coords)
Calculate the center of the pattern in its own coordinate system
center_pattern_x_unit = (min_x + max_x) / 2 center_pattern_y_unit = (min_y + max_y) / 2
Scaling factor: Adjust this to control the size of your pattern on the plate
Each unit in X_LABpattern will represent ‘scale_factor’ mm on the plate.
For a 30mm wide pattern with the current 15 unit width: 30 / (15 - 0) = 2
scale_factor = 2.0 # mm per unit
Volume to dispense per dot
dispense_volume = 1 # uL
Aspiration tracking
aspirated_volume = 0 max_aspirate_volume = 20 # uL (matching the example’s approach)
for i, (px, py) in enumerate(X_LABpattern): # If the pipette is empty or low on liquid, aspirate more # We aspirate max_aspirate_volume or whatever is needed for the remaining dots if aspirated_volume < dispense_volume: volume_to_aspirate = min(max_aspirate_volume, (len(X_LABpattern) - i) * dispense_volume - aspirated_volume) if volume_to_aspirate > 0: pipette_20ul.aspirate(volume_to_aspirate, cell_well) aspirated_volume += volume_to_aspirate
# Shift and scale the pattern coordinates
# The agar_plate['A1'].top() is the center, so we shift relative to that.
shifted_px = (px - center_pattern_x_unit) * scale_factor
shifted_py = (py - center_pattern_y_unit) * scale_factor
# Calculate the absolute position on the agar plate
adjusted_location = center_location.move(types.Point(x=shifted_px, y=shifted_py))
# Dispense the liquid using the helper function
dispense_and_detach(pipette_20ul, dispense_volume, adjusted_location)
aspirated_volume -= dispense_volume
Drop the tip at the end
pipette_20ul.drop_tip()
“”"### Protocolo Max_XLab_pattern
Este protocolo utiliza la plataforma Opentrons para patronar un diseño ‘XLab’ en una placa de agar. A continuación, se detalla el código completo:
Descripción general del funcionamiento:
- Carga de laboratorio: Se configuran las puntas, la pipeta y el módulo de temperatura, junto con las placas de colores y agar.
- Funciones auxiliares: Se definen
location_of_colorpara obtener la ubicación de un color específico ydispense_and_detachpara una dispensación limpia. - Definición del patrón
X_LABpattern: Una lista de coordenadas (x, y) que forman el diseño ‘X LAb’. Este patrón incluye la separación solicitada entre ‘X’, los puntos y ‘L’. - Cálculo de escalado y centrado: El código calcula las dimensiones del patrón para escalarlo y centrarlo en la placa de agar.
- Aspiración y dispensación: Itera sobre cada punto del patrón. La pipeta aspira líquido (‘Red’ de la posición ‘A1’) en bloques de 20 uL y dispensa 1 uL en cada coordenada ajustada en la placa de agar, utilizando la función
dispense_and_detachpara evitar manchas. - Descarte de punta: Al finalizar, la pipeta desecha la punta usada. """
from opentrons import types
metadata = { # see https://docs.opentrons.com/v2/tutorial.html#tutorial-metadata ‘author’: ‘MaxMatus’, ‘protocolName’: ‘Max_XLab_pattern’, ‘description’: ‘XLab_pattern’, ‘source’: ‘HTGAA 2026 Opentrons Lab’, ‘apiLevel’: ‘2.20’ }
Robot deck setup constants
TIP_RACK_DECK_SLOT = 9 COLORS_DECK_SLOT = 6 AGAR_DECK_SLOT = 5 PIPETTE_STARTING_TIP_WELL = ‘A1’
well_colors = { ‘A1’ : ‘Red’, ‘B1’ : ‘Green’, ‘C1’ : ‘Orange’ }
def run(protocol):
Load labware, modules and pipettes
Tips
tips_20ul = protocol.load_labware(‘opentrons_96_tiprack_20ul’, TIP_RACK_DECK_SLOT, ‘Opentrons 20uL Tips’)
Pipettes
pipette_20ul = protocol.load_instrument(“p20_single_gen2”, “right”, [tips_20ul])
Modules
temperature_module = protocol.load_module(’temperature module gen2’, COLORS_DECK_SLOT)
Temperature Module Plate
temperature_plate = temperature_module.load_labware(‘opentrons_96_aluminumblock_generic_pcr_strip_200ul’, ‘Cold Plate’)
Choose where to take the colors from
color_plate = temperature_plate
Agar Plate
agar_plate = protocol.load_labware(‘htgaa_agar_plate’, AGAR_DECK_SLOT, ‘Agar Plate’) ## TA MUST CALIBRATE EACH PLATE!
Get the top-center of the plate, make sure the plate was calibrated before running this
center_location = agar_plate[‘A1’].top()
pipette_20ul.starting_tip = tips_20ul.well(PIPETTE_STARTING_TIP_WELL)
Patterning
Helper functions for this lab
pass this e.g. ‘Red’ and get back a Location which can be passed to aspirate()
def location_of_color(color_string): for well,color in well_colors.items(): if color.lower() == color_string.lower(): return color_plate[well] raise ValueError(f"No well found with color {color_string}")
For this lab, instead of calling pipette.dispense(1, loc) use this: dispense_and_detach(pipette, 1, loc)
def dispense_and_detach(pipette, volume, location): """ Move laterally 5mm above the plate (to avoid smearing a drop); then drop down to the plate, dispense, move back up 5mm to detach drop, and stay high to be ready for next lateral move. 5mm because a 4uL drop is 2mm diameter; and a 2deg tilt in the agar pour is >3mm difference across a plate. """ assert(isinstance(volume, (int, float))) above_location = location.move(types.Point(z=location.point.z + 5)) # 5mm above pipette.move_to(above_location) # Go to 5mm above the dispensing location pipette.dispense(volume, location) # Go straight downwards and dispense pipette.move_to(above_location) # Go straight up to detach drop and stay high
YOUR CODE HERE to create your design
center_location = agar_plate[‘A1’].top()
cell_well = color_plate[‘A1’] # Changed ‘F1’ to ‘A1’ because F1 was not configured to have a color.
Aspirate
pipette_20ul.pick_up_tip()
X_LABpattern = [ # Línea horizontal izquierda (puntos en fila central, columnas negativas) (-5, 2), (-4, 2), (-3, 2), (-2, 2),
# X
(0, 4), (2, 4), # Fila superior
(1, 3), # Diagonal bajando
(1, 2), # Centro
(0, 1), (2, 1), # Fila inferior
# First dot after X - moved from (3,1) to (4,1)
(4, 1),
# Second dot for separation - moved from (4,1) to (5,1)
(5, 1),
# L (shifted by 2 units from previous adjustment: 5+2=7)
(7, 4),
(7, 3),
(7, 2),
(7, 1), (8, 1), (9, 1),
# Espacio entre letras (col 10 vacía)
# A (shifted by 2 units from previous adjustment: 9+2=11)
(11, 1), (11, 2), (11, 3), (11, 4),
(12, 4),
(13, 4), (13, 3), (13, 2), (13, 1),
(12, 2),
# Espacio entre letras (col 14 vacía)
# b (minúscula) (shifted by 2 units from previous adjustment: 13+2=15)
(15, 1), (15, 2), (15, 3), (15, 4),
(16, 2),
(16, 1),
(17, 2),
(17, 1),
# Línea horizontal derecha (puntos en fila central, columnas después de la b)
(19, 2), (20, 2), (21, 2), (22, 2),
]
Determine pattern dimensions for scaling and centering
x_coords = [p[0] for p in X_LABpattern] y_coords = [p[1] for p in X_LABpattern]
min_x, max_x = min(x_coords), max(x_coords) min_y, max_y = min(y_coords), max(y_coords)
Calculate the center of the pattern in its own coordinate system
center_pattern_x_unit = (min_x + max_x) / 2 center_pattern_y_unit = (min_y + max_y) / 2
Scaling factor: Adjust this to control the size of your pattern on the plate
Each unit in X_LABpattern will represent ‘scale_factor’ mm on the plate.
For a 30mm wide pattern with the current 15 unit width: 30 / (15 - 0) = 2
scale_factor = 2.0 # mm per unit
Volume to dispense per dot
dispense_volume = 1 # uL
Aspiration tracking
aspirated_volume = 0 max_aspirate_volume = 20 # uL (matching the example’s approach)
for i, (px, py) in enumerate(X_LABpattern): # If the pipette is empty or low on liquid, aspirate more # We aspirate max_aspirate_volume or whatever is needed for the remaining dots if aspirated_volume < dispense_volume: volume_to_aspirate = min(max_aspirate_volume, (len(X_LABpattern) - i) * dispense_volume - aspirated_volume) if volume_to_aspirate > 0: pipette_20ul.aspirate(volume_to_aspirate, cell_well) aspirated_volume += volume_to_aspirate
# Shift and scale the pattern coordinates
# The agar_plate['A1'].top() is the center, so we shift relative to that.
shifted_px = (px - center_pattern_x_unit) * scale_factor
shifted_py = (py - center_pattern_y_unit) * scale_factor
# Calculate the absolute position on the agar plate
adjusted_location = center_location.move(types.Point(x=shifted_px, y=shifted_py))
# Dispense the liquid using the helper function
dispense_and_detach(pipette_20ul, dispense_volume, adjusted_location)
aspirated_volume -= dispense_volume
Drop the tip at the end
pipette_20ul.drop_tip()
Execute Simulation / Visualization – don’t change this code block
protocol = OpentronsMock(well_colors) run(protocol) protocol.visualize()
II. Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
Article: BOTany Methods: Accessible Automation for Plant Synthetic Biology Moni Qiande, Abigail Lin, Lianna Larson, Cătălin Voiniciuc bioRxiv 2025.08.21.671538; doi: https://doi.org/10.1101/2025.08.21.671538 https://www.biorxiv.org/content/10.1101/2025.08.21.671538v1
I choose this article because is very accesible and usefull for a non specialized public, udergrads or people that is starting to learn how to use Opentrons and automation toos for biological applicatios. This is the abstract:
Most members of the synthetic biology community, particularly plant scientists, lack access to liquid handling robots to scale up experiments, enhance reproducibility, and accelerate the Design, Build, Test, Learn cycle. Biofoundries enable high throughput data acquisition to train AI models and to develop new bioproducts, but they are capital-intensive to set up and not widely distributed. Entry-level, 3D-printed robots offer more affordable alternatives, but suffer from a shortage of validated protocols that can be modified without prior coding experience. To enhance access to biological automation, we developed a collection of modular BOTany Methods using Opentrons OT-2 robots to streamline the most common methods for molecular biology research and education. Our comprehensive workflow offers automation for a variety of procedures, ranging from simple but repetitive tasks (such as primer dilution and PCR setup) to more complex operations, including Plant Modular Cloning (MoClo), bacterial transformation, and plasmid extraction. Our BOTany Methods enable undergraduate students and other early career researchers to run designer experiments using table-based inputs, without editing the custom Python scripts. This pipeline enables end-to-end molecular cloning with minimal user intervention, enhancing throughput and traceability for synthetic biology applications.
III. Write a description about what you intend to do with automation tools for your final project
To produce an antimicrobial biocuir using grape pomenace. The main goal is to enhance the antibacterial capabilities of the biocuir with syntetic byology principles. Firs option will be to search if there are some elements in the grape pomenace that are it self antimicrobial. If so, we will enhance those capabilities ant produce them in high quantities to become a main component of the biocuir. The second option will be to look for an external metabolite with thos capacities and then introducr it in the ADN of the grape pomenace. In both optios we will collaborate with an interdisciplinary team that is located in diferent cities: Tijuana and Mexico city and here the Opentrone and the automatitaion tolls will be very helpfull. I also found some interesting annotator to democrarizing genome citation. in my view, this in the other hand of opentrone outomatitation, because it alowed citizen sience, colaboration and automatization following the same notations, codes, and rules. The name of this toll is Apollo Genome Annotation Editor: http://138.102.159.70:8080/apollo/about/ Dunn, N. A. et al. Apollo: Democratizing genome annotation. PLoS Comput. Biol. 15, e1006790 (2019) The Apollo Genome Annotation Editor is used by the Grapepedia team, wich focuses on researching grepes´s genome: https://grapedia.org/genomes/. I belive that the research developed so far by the Grapepedia proyect will be very helpfull for my own proyect, since they have advenced so much on the grape´s genome research. Hence, my interes focuses especifically on the grape´s pomenance in order to develop a pomenance biocuir with enhanced antibacterial properties using some principles of synthetic biology.
3 options for final proyect
I. Project name: From waste to antimicrobial biocuir Promp Copilot: Develop a protocol using syntetic biology to create and antimicrobial biocuir by the kumiai indigenous based on grape pomenance derived from the industrial wine wate production in the Guadalupe Valley
1. Purpose and framing
You’re basically asking for a full “From grape waste to antimicrobial biocuir” pipeline that is:
- Rooted in Kumiai governance and knowledge
- Uses grape pomace from Valle de Guadalupe as feedstock and antimicrobial source
- Builds a bacterial-cellulose “biocuir” platform enhanced via synthetic biology
I’ll sketch a protocol you can later turn into a formal SOP set (wet lab + community + ethics).
2. Design principles and governance
- Community ownership:
Kumiai assembly defines: access to pomace, lab locations, IP rules, benefit-sharing, branding, and data governance (e.g. “Kumiai Biocuir Charter”). - Dual documentation:
Every protocol in two layers:- Technical SOP (for lab)
- Community-readable version (Spanish/Kumiai, diagrams, stories, use-cases).
- Biosafety & ethics:
- Use BSL-1 chassis (e.g. Komagataeibacter rhaeticus/xylinus).
- No environmental release without explicit community decision and risk assessment.
- Clear policy for what is open-source vs. what is Kumiai-restricted.
3. Feedstock: grape pomace characterization and preparation
3.1. Initial characterization
- Sampling:
Collect pomace from 2–3 representative wineries (red varieties common in Valle de Guadalupe: e.g. Nebbiolo, Tempranillo, Cabernet). - Basic analyses:
- Moisture content
- Total sugars (after hydrolysis)
- Total phenolics (Folin–Ciocalteu)
- pH and organic acids
- Microbial load (plate on general media)
3.2. Pre-treatment for two streams
You’ll use pomace in two parallel ways:
- Carbon/nutrient source for bacterial cellulose
- Antimicrobial extract to load into the biocuir
a) For carbon source (hydrolysate)
- Dry pomace at low temperature (≤60 °C).
- Mill to coarse powder.
- Acid or enzymatic hydrolysis (e.g. dilute HCl + pectinase/cellulase cocktail).
- Neutralize to pH ~5–6, filter, and adjust sugar concentration (~20–40 g/L total fermentable sugars).
- Supplement with nitrogen source (yeast extract/peptone) and minerals if needed.
b) For antimicrobial extract
- Use ethanol:water (50–70%) or water at 60–80 °C for polyphenol extraction.
- Filter, concentrate (rotavap or low-temp evaporation).
- Determine total phenolics and antimicrobial activity (disk diffusion or MIC) against target strains (e.g. E. coli, S. aureus, Candida).
4. Microbial platform and synthetic biology strategy
4.1. Base strain for biocuir
- Preferred chassis: Komagataeibacter xylinus or K. rhaeticus (robust BC producers).
- Maintain a wild-type production strain as baseline (no genetic modifications) for regulatory flexibility.
4.2. Synthetic biology modules (optional, modular)
Design two tiers:
Non-GMO functionalization
- Load grape-pomace extracts into cellulose during or after growth.
- Crosslink with natural agents (e.g. tannins, genipin, citric acid) to retain antimicrobials.
GMO strain (only if community approves)
- Engineer Komagataeibacter to express:
- Antimicrobial peptides (AMPs) under inducible promoter.
- Pigments (e.g. melanin, carotenoids) for color and UV protection.
- Use kill-switch or auxotrophy to reduce environmental risk.
- All constructs documented and co-decided with Kumiai governance body.
- Engineer Komagataeibacter to express:
5. Bacterial cellulose/biocuir production protocol
5.1. Medium formulation (pomace-based)
- Base medium:
- Grape pomace hydrolysate (adjust to desired sugar level).
- Yeast extract + peptone (or plant-based nitrogen if desired).
- Adjust pH to ~5.0–6.0.
- Sterilize (autoclave or 0.22 µm filtration depending on clarity).
5.2. Static culture for pellicle formation
- Inoculate sterile trays or shallow bioreactors with Komagataeibacter starter culture (5–10% v/v).
- Incubate static at 28–30 °C for 7–14 days until thick pellicle forms.
- For in situ loading of grape extract:
- Add defined concentration of antimicrobial extract to the medium at inoculation or after 24–48 h (to avoid growth inhibition).
5.3. Harvest and primary purification
- Harvest pellicles, rinse thoroughly with water.
- Treat with alkaline wash (e.g. NaOH at 60–80 °C) to remove cells and residual medium.
- Rinse to neutral pH.
- Optional: post-loading by soaking purified pellicles in grape-pomace extract solution, then gentle drying.
6. Biocuir finishing and functionalization
6.1. Plasticization and crosslinking
- Plasticizers:
- Glycerol, sorbitol, or other biocompatible agents to improve flexibility.
- Crosslinkers (natural focus):
- Citric acid (heat-activated), tannins from pomace, genipin, etc.
- Optimize for: tensile strength, flexibility, water resistance, and retention of antimicrobial activity.
6.2. Drying and mechanical processing
- Dry under controlled conditions (e.g. framed drying to avoid warping).
- Press/laminate to desired thickness.
- Surface treatments: natural oils, waxes, or additional tannin-rich coatings.
6.3. Coloring and patterning (Kumiai aesthetics)
- Use grape-skin pigments, other local plant dyes, or engineered pigments.
- Co-design patterns, symbols, and textures with Kumiai artisans—this is where cultural meaning enters the material.
7. Antimicrobial and material testing
7.1. Antimicrobial assays
- Disk or film diffusion tests on agar against panel of microbes.
- Time-kill or log-reduction tests in liquid contact assays.
- Compare:
- BC without extract
- BC + grape extract (non-crosslinked)
- BC + grape extract + crosslinker
- (If used) GMO-BC with AMP expression
7.2. Material properties
- Mechanical: tensile strength, elongation at break, Young’s modulus.
- Physical: water uptake, swelling, thickness, density.
- Aging: UV exposure, repeated wet–dry cycles.
All data should be returned to the Kumiai assembly in accessible formats (visuals, stories, demos), not just as technical reports.
8. Community protocol and value chain
8.1. Co-designed workflow
Map a Kumiai-centered value chain:
- Pomace sourcing agreements with wineries (fair price or co-ownership).
- Local bioprocess node (small lab/bioreactor in Kumiai territory or nearby).
- Artisan finishing workshop for cutting, sewing, and product design.
- Brand and narrative: Kumiai-led storytelling about land, water, and circularity.
8.2. Governance instruments
- Biocuir Governance Document including:
- Data sovereignty (who owns strain data, protocols, and performance data).
- Rules for collaborations with universities/companies.
- Licensing model (e.g. community license for Kumiai, open-source for others with conditions).
9. Synthetic biology roadmap (phased)
Phase 1 – Low-risk, fast start
- Wild-type Komagataeibacter
- Pomace hydrolysate medium
- Grape-pomace extract loading (non-GMO)
- Basic antimicrobial + mechanical testing
- Kumiai branding and first prototypes
Phase 2 – Process optimization
- Medium optimization for yield and thickness
- Crosslinking and finishing optimization
- LCA and socio-economic assessment with Kumiai criteria
Phase 3 – Engineered functions (only if approved)
- Introduce AMP or pigment modules
- Implement kill-switch/auxotrophy
- Regulatory and ethical review with Kumiai at every step
SOP-01 – Community governance and feedstock sourcing
Purpose:
Define how Kumiai governance, wineries, and lab teams agree on access, ownership, and use of grape pomace.
Key elements:
- Scope: All activities involving grape pomace from Valle de Guadalupe for biocuir R&D and production.
- Responsibilities:
- Kumiai Assembly: Approves project, sets benefit-sharing, branding, and data/IP rules.
- Wineries: Provide pomace under written agreement.
- Lab Team: Follows biosafety, records data, respects governance rules.
Procedure (outline):
Community agreement:
- Draft a “Kumiai Biocuir Charter” (Spanish/Kumiai) defining: ownership, data sovereignty, IP, branding, and revenue-sharing.
- Approve via Kumiai assembly; archive signed version.
Winery agreements:
- Sign MOUs specifying: volume and type of pomace, collection schedule, hygiene expectations, and compensation/benefits.
Traceability:
- Assign batch IDs to each pomace lot (winery, date, grape variety, processing conditions).
- Record in both lab log and community-accessible register.
Ethics & biosafety:
- Confirm BSL-1 scope and no environmental release without assembly approval.
- Define policy for GMO vs. non-GMO work and where each is allowed.
SOP-02 – Grape pomace reception, characterization, and storage
Purpose:
Standardize how pomace is received, characterized, and stored as a feedstock.
Materials:
Clean containers, scales, pH meter, moisture analyzer or oven, basic microbiology media, PPE.
Procedure:
Reception:
- Inspect pomace (absence of visible mold, foreign objects).
- Weigh and assign batch ID.
- Record grape variety, winery, pressing type (red/white), and date.
Basic characterization:
- Moisture: Dry subsample at ~60 °C to constant weight.
- pH: Mix pomace with distilled water (1:5 w/v), measure pH.
- Microbial load: Plate serial dilutions on general medium; incubate and record CFU.
Storage:
- Short term (≤72 h): 4 °C.
- Long term: dry at ≤60 °C, then store in sealed containers in dark, cool place.
Documentation:
- Record all values in batch sheet and community-facing summary (simple graphs/infographics).
SOP-03 – Preparation of grape pomace hydrolysate (carbon source)
Purpose:
Produce a sterile, sugar-rich hydrolysate from pomace for bacterial cellulose fermentation.
Materials:
Dried pomace, grinder, acid (e.g. HCl), enzymes (pectinase/cellulase, if used), pH meter, filters, autoclave.
Procedure:
Milling:
- Grind dried pomace to coarse powder.
Hydrolysis:
- Suspend pomace in water (e.g. 5–10% w/v).
- Option A: Acid hydrolysis (mild acid, controlled temperature).
- Option B: Enzymatic hydrolysis (add pectinase/cellulase, incubate at recommended temperature/time).
Neutralization and clarification:
- Adjust pH to ~5–6.
- Filter or decant solids to obtain clarified hydrolysate.
Adjustment:
- Measure total sugars (e.g. colorimetric assay).
- Dilute or concentrate to target sugar concentration (e.g. 20–40 g/L).
Sterilization:
- Autoclave or filter-sterilize depending on clarity and equipment.
Storage:
- Store at 4 °C; label with batch ID and sugar concentration.
SOP-04 – Preparation of grape pomace antimicrobial extract
Purpose:
Obtain a polyphenol-rich extract from pomace to confer antimicrobial activity to biocuir.
Materials:
Pomace (fresh or dried), ethanol and water, heating/stirring setup, filters, rotavap/evaporator, Folin–Ciocalteu reagents (or equivalent), microbiology media.
Procedure:
Extraction:
- Mix pomace with ethanol:water (50–70%) or hot water (60–80 °C).
- Stir for defined time (e.g. 1–3 h), then cool.
Filtration:
- Filter to remove solids; collect extract.
Concentration:
- Concentrate under reduced pressure or low heat to desired volume.
Characterization:
- Measure total phenolics (e.g. Folin–Ciocalteu).
- Perform small-scale antimicrobial screening (disk diffusion) against reference strains.
Storage:
- Store in amber bottles at 4 °C; label with batch ID, solvent, phenolic content.
SOP-05 – Bacterial cellulose pellicle production using pomace hydrolysate
Purpose:
Produce bacterial cellulose (BC) pellicles as the base “biocuir” material.
Materials:
Komagataeibacter starter culture, pomace hydrolysate (SOP-03), nitrogen source, minerals, trays/bioreactors, incubator.
Procedure:
Medium preparation:
- Combine pomace hydrolysate with nitrogen source (e.g. yeast extract/peptone) and minerals.
- Adjust pH to ~5–6 and sterilize.
Inoculum:
- Prepare active Komagataeibacter culture in standard medium.
- Inoculate production medium at 5–10% v/v.
Static culture:
- Pour into shallow sterile trays.
- Incubate static at ~28–30 °C for 7–14 days until thick pellicle forms.
Optional in situ loading:
- Add defined concentration of grape extract (SOP-04) at inoculation or after initial growth phase, based on inhibition tests.
Harvest:
- Remove pellicles aseptically; record thickness and appearance.
SOP-06 – Purification, loading, and finishing of biocuir
Purpose:
Purify BC pellicles, load antimicrobial extract, and finish as flexible “biocuir”.
Materials:
BC pellicles, NaOH (or equivalent), water, grape extract, plasticizers (e.g. glycerol), natural crosslinkers (citric acid, tannins), drying frames/press.
Procedure:
Purification:
- Wash pellicles in water to remove medium.
- Treat with warm alkaline solution to lyse cells and remove residues.
- Rinse repeatedly to neutral pH.
Post-loading (if not in situ):
- Immerse purified pellicles in grape extract solution for defined time.
- Remove and gently blot excess.
Plasticization and crosslinking:
- Soak in plasticizer solution (e.g. glycerol in water).
- Apply natural crosslinker (e.g. citric acid, tannin-rich solution) and heat-treat if required.
Drying and shaping:
- Mount pellicles on frames to avoid warping.
- Dry under controlled temperature and airflow.
- Press/laminate to desired thickness.
Aesthetic finishing:
- Apply natural dyes (grape skins, local plants) and textures co-designed with Kumiai artisans.
SOP-07 – Antimicrobial and material performance testing
Purpose:
Evaluate antimicrobial activity and material properties of biocuir batches.
Materials:
Test microbes, agar plates, incubator, tensile tester (if available), water uptake setup.
Procedure:
Antimicrobial tests:
- Cut biocuir discs/strips.
- Place on inoculated agar (disk/film diffusion) and incubate.
- Measure inhibition zones.
- Optionally perform contact-kill or log-reduction assays in liquid.
Material tests:
- Measure tensile strength and elongation at break.
- Assess water uptake/swelling after immersion.
- Perform simple aging tests (UV exposure, wet–dry cycles).
Batch release criteria:
- Define minimum antimicrobial effect and mechanical thresholds with Kumiai assembly and artisans (fit-for-purpose).
SOP-08 – Data, communication, and community reporting
Purpose:
Ensure all technical data are translated into accessible formats and returned to Kumiai governance.
Procedure:
Data recording:
- Maintain lab notebooks and digital records for all SOPs.
- Tag each dataset with batch ID and date.
Translation and visualization:
- Convert key results into bilingual (Spanish/Kumiai) summaries with simple graphs, photos, and short narratives.
Assembly reporting:
- Present results periodically to Kumiai assembly: performance, risks, opportunities, and proposed changes.
- Record feedback and integrate into protocol revisions.
Version control:
- Maintain versioned SOPs; log changes and reasons (including community decisions).
II. Name: Antiseptic and anti-inflammatory pomenace patch Copilot prompt: How to develop and Antiseptic and anti-inflammatory patch using syntetic biology metodhs in parthnership with the the kumiai indigenous based on grape pomenance derived from the industrial wine production in the Guadalupe Valley
1. The Kummiai community. Why? Governance system.
2. Characterize the grape pomace from Guadalupe Valley
2.1. Source and preprocessing
- Source definition:
- Identify 1–2 wineries willing to provide consistent pomace (same variety, same process).
- Stabilization:
- Drying: low temperature (≤50 °C) or lyophilization to preserve polyphenols.
- Milling: standardize particle size for extraction reproducibility.
2.2. Chemical and bioactivity profiling
- Polyphenol profiling:
- Bioactivity assays (in vitro):
- Antimicrobial: MIC/MBC against Staphylococcus aureus, E. coli, and skin-relevant strains. Grape pomace extracts often show stronger activity against Gram-positive bacteria. Springer Frontiers
- Anti-inflammatory: protein denaturation inhibition, NO inhibition in macrophage cell lines, etc.—grape pomace extracts have shown promising anti-inflammatory effects. Springer MDPI
- Antioxidant: DPPH, ABTS, FRAP to correlate with anti-inflammatory potential.
Outcome of this phase: a data-backed argument that “Kumiai grape pomace X from Guadalupe Valley has antiseptic and anti-inflammatory potential at Y concentration.”
3. Synthetic biology strategy: from extract to engineered system
Engineered microbes producing key actives
- Target molecules:
- Choose 1–3 “hero” compounds from your profiling (e.g., resveratrol, quercetin derivatives, gallic acid–related pathways) that correlate strongly with antimicrobial/anti-inflammatory activity. MDPI
- Host selection:
- Yeast (e.g., Saccharomyces cerevisiae): natural fit for resveratrol and other phenolics; compatible with wine narrative.
- GRAS bacteria (e.g., Lactobacillus spp.): interesting if you want a “living” probiotic patch, but more complex regulatory path.
- Pathway engineering:
- Introduce or optimize phenylpropanoid/flavonoid pathways for your target compounds.
- Use promoters responsive to simple inducers (e.g., temperature, small molecules) to control production.
- Coupling to grape pomace:
- Use pomace hydrolysates as carbon/nutrient source for the engineered strain—closing the circular bioeconomy loop and keeping the story “Kumiai + Guadalupe Valley wine waste” coherent.
Living or semi-living patch
- Concept: A patch containing immobilized or killed engineered cells that have pre-produced the actives.
- Safer intermediate: Use cell-free extracts or purified compounds from engineered microbes, not live GMOs on skin, at least initially.
4. Designing the antiseptic/anti-inflammatory patch
Think of the patch as: [biomaterial matrix] + [grape pomace actives] + (optional) [engineered bioactives].
4.1. Choose the base material
- Biocuir / bacterial cellulose:
- Produced by Komagataeibacter spp.; high water-holding capacity, biocompatible, and already used in wound dressings.
- Can be grown on grape pomace–derived media to embed the circularity narrative.
- Alternative matrices:
- Chitosan (intrinsic antimicrobial), alginate, gelatin, or blends—depending on local availability and Kumiai preferences for texture and feel.
4.2. Loading the actives
- Approach A: Soaking/adsorption:
- Grow the biomaterial (e.g., bacterial cellulose), then soak in standardized grape pomace extract; dry to desired moisture level.
- Approach B: In situ production:
- Co-culture cellulose-producing bacteria with engineered yeast/bacteria that secrete phenolics into the medium; the matrix forms already loaded with actives.
- Release profile testing:
- In vitro diffusion assays (Franz cells or simple diffusion setups) to measure release of key phenolics over time.
4.3. Functional testing
- Antimicrobial performance:
- Place patch discs on bacterial lawns (agar diffusion) to test inhibition zones.
- Anti-inflammatory performance:
- Extract from patch and test in cell-based inflammation models (e.g., LPS-stimulated macrophages).
- Biocompatibility:
- Cytotoxicity assays (e.g., keratinocytes, fibroblasts).
- Later: small, ethically approved human patch tests for irritation.
III. Proyect name: A Yumano grape variety to promote circular economy Promp Copilot: Develop a grape variety that is resistant to water stress and has organoleptic qualities similar to the pinyon nut—a fruit that is very common in the region and a traditional food staple of Yuman Indigenous groups—requires a culturally grounded and environmentally adapted approach. The grape variety will be for the exclusive use of local Yuman Indigenous communities, who will cultivate it and manage it sustainably, while also transforming the grape pomace into antibacterial biocuir. This means the entire initiative will function as a circular‑economy project from beginning to end, with a strong cultural dimension and a direct contribution to community health. For this reason, the grape’s properties must include very high antimicrobial capacity from the outset. To achieve this, synthetic biology techniques can be used.
🌱 Project goal
Develop a new grape variety designed to:
- Withstand extreme water stress and poor soils.
- Express organoleptic qualities inspired by pinyon nuts (a traditional food of Yuman peoples).
- Produce intrinsic antimicrobial compounds in its skin and pulp.
- Be cultivated and used exclusively by local Yuman Indigenous communities.
- Enable the transformation of grape pomace into antibacterial biocuir (biocuir leather).
- Operate as a fully circular, culturally grounded, community‑health–oriented system.
🍇 Organoleptic traits inspired by pinyon nuts
Pinyon nuts have a distinctive sensory profile:
- Resinous notes (terpenes such as α‑pinene and limonene).
- Buttery aromas (lactones).
- Mild sweetness and a fatty texture (specific fatty acids).
To translate this into a grape variety:
- Select parent lines rich in terpenes (e.g., Muscat, Malvasia).
- Enhance metabolic pathways that increase lactones and sesquiterpenes.
- Modulate the synthesis of volatile fatty acids that contribute to nut‑like aromas.
💧 Drought resistance
The environmental conditions of the Valle de Guadalupe require:
- Thick cuticles.
- Low stomatal density.
- Deep root systems.
- Efficient metabolism under water deficit.
Useful genetic sources include:
- Vitis berlandieri (drought and alkaline soil tolerance).
- Vitis arizonica (desert‑adapted).
- Mediterranean varieties such as Garnacha and Monastrell.
🧬 Intrinsic antimicrobial capacity
To ensure the grape naturally produces antimicrobial compounds:
- Increase stilbenes (resveratrol and derivatives).
- Enhance flavonoids with bactericidal activity.
- Boost terpenes with antifungal properties.
- Introduce or upregulate plant antimicrobial peptides (AMPs) from desert species.
These traits support:
- Reduced agricultural inputs.
- Improved vineyard health.
- Antibacterial properties in the biocuir produced from pomace.
🔬 Biotechnological strategies
Marker‑assisted classical breeding
- Controlled crosses between selected parent lines.
- Early seedling selection for drought tolerance, terpene production, and stilbene pathways.
Gene editing (CRISPR/Cas)
- Modulate terpene synthesis genes to achieve pinyon‑like aromas.
- Enhance drought‑response genes (DREB, aquaporins).
- Increase stilbene biosynthesis for antimicrobial strength.
Metabolic engineering
- Introduce or optimize biosynthetic pathways for antimicrobial compounds.
- Adjust lactone and terpene synthesis for the desired aromatic profile.
Synthetic biology for pomace transformation
- Engineer microorganisms to convert pomace into antibacterial biocuir.
- Integrate genes that produce natural polymers with antimicrobial properties.
🌀 Integration into a Yuman circular economy
The project can function as a complete system:
- Cultivation: vineyards managed by Yuman communities.
- Transformation: scientific and artisanal winemaking.
- By‑products: pomace → biocuir → crafts, clothing, tools.
- Community health: natural antimicrobial compounds.
- Cultural identity: a grape expressing the ancestral flavor of pinyon nuts.
🌍 Ethical governance and Indigenous sovereignty
This initiative can become a model of:
- Indigenous‑led innovation.
- Biotechnology with free, prior, and informed consent.
- Collective ownership of germplasm.
- Fair and culturally aligned benefit‑sharing frameworks.
Great—let’s dive into the synthetic biology design and keep it focused and actionable.
1. Overall architecture
Two coupled synthetic biology systems:
Engineered grape plant
- Drought‑resilient.
- Pinyon‑like aroma profile.
- High intrinsic antimicrobial metabolite production.
Engineered microbial platform for pomace → antibacterial biocuir
- Uses grape pomace as main carbon/nutrient source.
- Produces a polymeric matrix (biocuir) with embedded antimicrobial activity.
2. Plant synthetic biology design
2.1. Target traits and modules
Module A – Terpene & pinyon‑like aroma module
- Goal: Resinous, nutty, pine‑like notes.
- Targets:
- Upregulate monoterpene synthases (e.g., α‑pinene synthase, limonene synthase).
- Introduce/boost sesquiterpene synthases linked to nutty/resinous notes.
- Strategy:
- Use fruit/skin‑specific promoters (e.g., Vitis berry‑specific) to avoid growth penalties.
- CRISPR activation (CRISPRa) of endogenous terpene pathway genes where possible.
Module B – Antimicrobial stilbene/flavonoid module
- Goal: High antimicrobial capacity in skin and pomace.
- Targets:
- Overexpress stilbene synthase (STS) variants for resveratrol and derivatives.
- Upregulate flavonoid biosynthesis genes (e.g., CHS, CHI, F3H, FLS) with known antimicrobial activity.
- Strategy:
- Stack multiple STS genes under inducible or pathogen‑responsive promoters.
- Tune flux via overexpression of PAL (phenylalanine ammonia‑lyase) to feed phenylpropanoid pathway.
Module C – Drought‑tolerance module
- Goal: Maintain yield and metabolite production under water stress.
- Targets:
- Transcription factors: DREB/CBF, NAC, bZIP associated with drought response.
- Aquaporins and osmoprotectant pathways (e.g., proline biosynthesis).
- Strategy:
- Use stress‑inducible promoters (RD29A‑like or grape homologs) to avoid constitutive stress phenotype.
- Prefer cis‑regulatory editing (promoter tuning) of endogenous genes to reduce regulatory burden.
Module D – Safety & containment
- Goal: Align with Indigenous governance and biosafety.
- Design elements:
- Avoid antibiotic resistance markers in final lines (use marker‑free selection or excision systems).
- Prefer cisgenic/intragenic approaches when possible (grape or closely related Vitis genes).
- Genomic integration in well‑characterized “safe harbors” to ensure stability.
3. Microbial synthetic biology for antibacterial biocuir
3.1. Chassis selection
- Primary candidates:
- Komagataeibacter xylinus (Gluconacetobacter) for bacterial cellulose (excellent for “leather‑like” materials).
- Optionally co‑culture with:
- Lactic acid bacteria (for acidification, preservation).
- Yeasts (if needed for aroma or cross‑feeding).
3.2. Feedstock integration (pomace as substrate)
- Design goals:
- Use grape pomace hydrolysates (skins, seeds, stems) as carbon and nitrogen source.
- Engineering steps:
- Introduce or enhance pectinase, cellulase, tannase and polyphenol‑tolerant traits in the chassis or in a helper strain.
- Design a pre‑fermentation enzymatic cocktail (could be microbial or added enzymes) to:
- Release sugars from pomace.
- Retain and concentrate antimicrobial phenolics from the engineered grape.
3.3. Antibacterial biocuir module
- Matrix:
- High‑purity bacterial cellulose sheets grown in shallow trays with pomace‑derived medium.
- Antimicrobial enhancement:
- Two layers of design:
- Passive loading:
- The medium already contains high levels of stilbenes/flavonoids from the engineered grape.
- These bind or embed into the cellulose matrix.
- Active production:
- Engineer the cellulose‑producing bacteria to express:
- Plant‑inspired antimicrobial peptides (AMPs).
- Or biosynthetic pathways for small antimicrobial molecules (e.g., phenazine‑like, but carefully chosen for safety and regulation).
- Engineer the cellulose‑producing bacteria to express:
- Passive loading:
- Two layers of design:
- Control of expression:
- Use inducible promoters (e.g., sugar‑ or pH‑responsive) to trigger AMP production only during biocuir growth.
- Include kill‑switch or auxotrophy to prevent survival outside controlled fermentation.
4. Interface between plant and microbial systems
- Design principle: The grape is “pre‑programmed” to feed the biocuir process.
- Key couplings:
- High stilbene/flavonoid content in skins → higher antimicrobial loading in biocuir.
- Specific sugar and organic acid profile in pomace → optimized for Komagataeibacter growth.
- Potential to select for pomace‑tolerant strains co‑evolved with the specific grape variety.
5. Experimental roadmap (very high level)
In silico & in vitro pathway design
- Metabolic modeling of grape phenylpropanoid and terpene pathways.
- Selection of gene targets and promoters.
Grape transformation & CRISPR editing
- Generate multiple edited lines with different combinations of Modules A–C.
- Screen for metabolite profiles (terpenes, stilbenes, flavonoids) and drought performance.
Chassis engineering for biocuir
- Engineer Komagataeibacter for:
- Growth on pomace hydrolysate.
- Stable cellulose production.
- Optional AMP or antimicrobial metabolite production.
- Engineer Komagataeibacter for:
Integrated trials
- Pilot vinification → pomace collection → biocuir fermentation.
- Test mechanical properties and antimicrobial activity of biocuir sheets.
Co‑design with Yuman communities
- Iterative feedback on:
- Sensory profile of the grape and wine.
- Look, feel, and cultural meaning of the biocuir.
- Governance, naming, and access rules.
- Iterative feedback on:
If you’d like, next step I can:
- Go deeper into specific gene candidates and promoter choices, or
- Sketch a work package structure (WP1–WP6) for a grant or consortium proposal around this design.
Here’s a focused list of concrete gene candidates and promoter choices, mapped to the modules we sketched.
1. Antimicrobial stilbene/flavonoid module
1.1. Core stilbene synthase genes
Gene candidates (Vitis vinifera):
- VINST1 / Stilbene synthase 1 (STS1, P28343)
- Well‑characterized STS, catalyzes resveratrol formation and enhances pathogen resistance. UniProt
- VvSTS gene family members (e.g., VvSTS29, others in the cluster)
- Large multi‑gene family with distinct regulatory regions and inducible expression patterns. Springer Oxford Academic
Design idea:
- Stack 2–3 STS isoforms with complementary promoter responses (e.g., MeJA‑responsive, pathogen‑responsive) to broaden induction conditions.
1.2. Transcription factor layer
- VviMYB14, VviMYB15 (R2R3‑MYB TFs)
- Direct activators of STS promoters; strongly co‑expressed with STS genes under stress. Oxford Academic
- VviWRKY03, VviWRKY24, VviWRKY43, VviWRKY53
- WRKY TFs combinatorially regulating STS expression; VviWRKY24 strongly activates VvSTS29 promoter. Oxford Academic
Design idea:
- Create a synthetic regulatory cassette where:
- A stress‑inducible promoter drives VviMYB14.
- A pathogen‑responsive promoter drives VviWRKY24.
- Downstream, native VvSTS promoters control STS coding sequences (cisgenic feel, strong induction).
1.3. Promoter choices for stilbene module
- Native VvSTS promoters (e.g., VvSTS29 promoter)
- Show distinct responses to MeJA, cyclodextrins, and other elicitors; promoter diversity can be exploited for fine‑tuning. Springer Oxford Academic
- General phenylpropanoid/stress promoters (grape homologs of PAL, PR genes)
- For broader stress‑responsive expression of upstream enzymes like PAL.
Practical pattern:
- Promoter_STS29::STS1 (VINST1)
- Promoter_STSx::STS_isoform2
- Promoter_stress::VviMYB14
- Promoter_pathogen::VviWRKY24
2. Terpene & pinyon‑like aroma module
Here we rely more on general Vitis terpene synthase (TPS) logic than specific named genes in the sources, but the pattern is:
Gene types:
- Monoterpene synthases: α‑pinene synthase, limonene synthase (Vitis TPS family members).
- Sesquiterpene synthases: for resinous/nutty notes.
Promoter choices:
- Berry/skin‑specific promoters (to localize aroma to the fruit):
- VvUFGT promoter (UDP‑glucose:flavonoid 3‑O‑glucosyltransferase; berry‑skin specific in color development).
- VvMybA1 promoter (controls anthocyanin pathway in skins; strong berry‑specific activity).
Design pattern:
- pVvUFGT::α‑pinene synthase
- pVvUFGT::limonene synthase
- pVvMybA1::sesquiterpene synthase
This keeps terpene overproduction largely confined to the berry skin, minimizing whole‑plant metabolic burden.
3. Drought‑tolerance module
Gene candidates (conceptual, grape homologs to be identified):
- DREB/CBF transcription factors (DREB2‑like, CBF‑like)
- Master regulators of drought and cold responses.
- NAC and bZIP TFs associated with ABA and drought signaling.
- Aquaporins (PIP, TIP families) and proline biosynthesis enzymes (e.g., P5CS).
Promoter choices:
- Stress‑inducible promoters:
- Grapevine homologs of RD29A‑like promoters (dehydration‑responsive).
- Native drought‑responsive promoters identified from grape transcriptomic data (e.g., promoters of strongly drought‑induced genes).
Design pattern:
- pDrought_inducible::VvDREB_homolog
- pDrought_inducible::VvNAC_homolog
This ensures drought‑tolerance pathways are activated under stress, not constitutively.
4. Promoter logic for safety and cultural governance
To align with Yuman governance and biosafety:
- Prefer native Vitis promoters (STS, PAL, berry‑specific, stress‑responsive) over viral promoters (e.g., 35S), to keep the design cisgenic/intragenic as much as possible. Springer UniProt
- Use marker‑free systems:
- Cre/lox or recombinase‑based excision of selection markers after transformation.
- Consider Indigenous‑defined “off‑switch” promoters:
- For example, a promoter responsive to a specific agro‑ecological practice (e.g., a foliar spray allowed only within community‑controlled fields) that can down‑regulate key TFs if needed.
5. Microbial biocuir module – key elements
Even though the search was grape‑focused, we can specify a reasonable synthetic design for Komagataeibacter:
Genes:
- Cellulose synthase operon (bcsABCD) – native, but we can upregulate via stronger promoters.
- Secreted AMPs:
- Short, plant‑inspired peptides (e.g., defensin‑like) with broad‑spectrum but low‑toxicity profiles.
Promoters:
- Constitutive medium‑strength promoter for cellulose (to avoid metabolic overload).
- Inducible promoter for AMPs:
- Sugar‑inducible (e.g., arabinose‑like logic) or pH‑responsive, activated only during biocuir growth phase.
Design pattern:
- pConstitutive_medium::bcs_operon (if needed for boost)
- pInducible_pomace_sugar::AMP1‑secreted
If you’d like, next step I can:
- Turn this into a schematic genetic circuit map (module by module), or
- Draft a methods‑style section (for a paper or grant) describing how these constructs would be built and tested in phases.
Here’s a text‑based schematic genetic circuit map for the main modules. Think of each as a layer you can redraw later in vector form.
1. Plant circuit – stilbene / antimicrobial module
Output: high resveratrol + related stilbenes in berry skin and pomace.
2. Plant circuit – terpene / pinyon‑like aroma module
Output: pine‑resin, nutty, pinyon‑like volatiles localized in berry skin.
3. Plant circuit – drought‑tolerance module
Output: improved water‑stress resilience without constitutive stress phenotype.
4. Microbial circuit – antibacterial biocuir (Komagataeibacter)
Output: cellulose‑based biocuir with embedded antimicrobial phenolics + secreted AMPs.
5. Safety / governance layer (conceptual)
This layer is where Yuman governance can be encoded: a controllable “dial” on how intense the antimicrobial/secondary metabolism is, tied to practices only allowed within community‑managed systems.

