from opentrons import types
import string
metadata = {
'protocolName': '{Ashraful} - Opentrons Art - HTGAA',
'author': 'Ashraful',
'source': 'HTGAA 2026',
'apiLevel': '2.20'
}
Z_VALUE_AGAR = 2.0
POINT_SIZE = 0.75
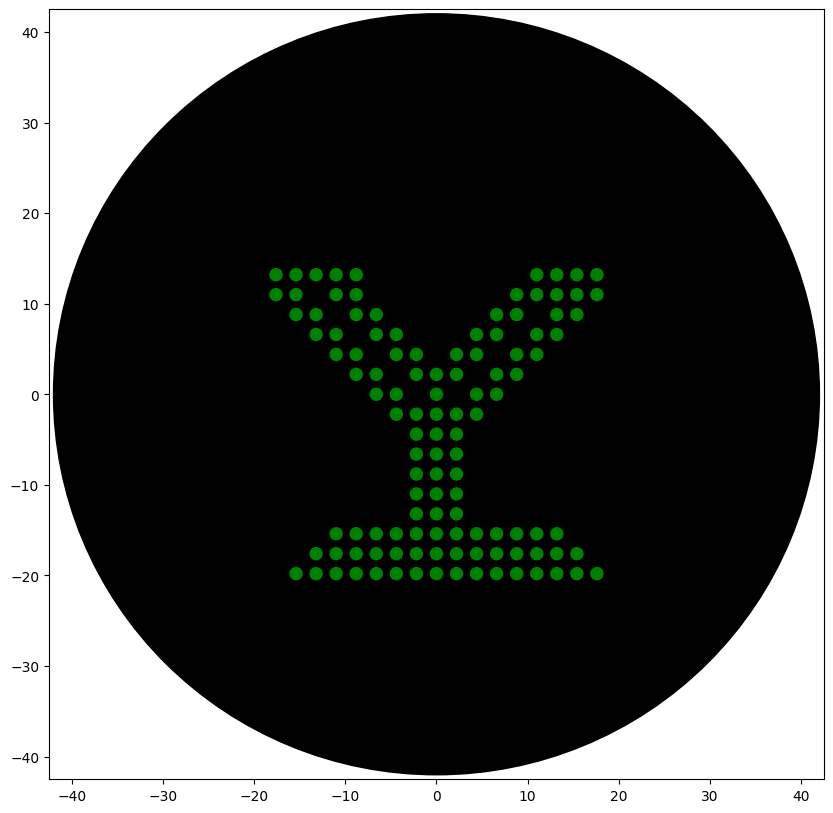
sfgfp_points = [(-17.6,13.2), (-15.4,13.2), (-13.2,13.2), (-11,13.2), (-8.8,13.2), (11,13.2), (13.2,13.2), (15.4,13.2), (17.6,13.2), (-17.6,11), (-15.4,11), (-11,11), (-8.8,11), (8.8,11), (11,11), (13.2,11), (15.4,11), (17.6,11), (-15.4,8.8), (-13.2,8.8), (-8.8,8.8), (-6.6,8.8), (6.6,8.8), (8.8,8.8), (13.2,8.8), (15.4,8.8), (-13.2,6.6), (-11,6.6), (-6.6,6.6), (-4.4,6.6), (4.4,6.6), (6.6,6.6), (11,6.6), (13.2,6.6), (-11,4.4), (-8.8,4.4), (-4.4,4.4), (-2.2,4.4), (2.2,4.4), (4.4,4.4), (8.8,4.4), (11,4.4), (-8.8,2.2), (-6.6,2.2), (-2.2,2.2), (0,2.2), (2.2,2.2), (6.6,2.2), (8.8,2.2), (-6.6,0), (-4.4,0), (0,0), (4.4,0), (6.6,0), (-4.4,-2.2), (-2.2,-2.2), (0,-2.2), (2.2,-2.2), (4.4,-2.2), (-2.2,-4.4), (0,-4.4), (2.2,-4.4), (-2.2,-6.6), (0,-6.6), (2.2,-6.6), (-2.2,-8.8), (0,-8.8), (2.2,-8.8), (-2.2,-11), (0,-11), (2.2,-11), (-2.2,-13.2), (0,-13.2), (2.2,-13.2), (-11,-15.4), (-8.8,-15.4), (-6.6,-15.4), (-4.4,-15.4), (-2.2,-15.4), (0,-15.4), (2.2,-15.4), (4.4,-15.4), (6.6,-15.4), (8.8,-15.4), (11,-15.4), (13.2,-15.4), (-13.2,-17.6), (-11,-17.6), (-8.8,-17.6), (-6.6,-17.6), (-4.4,-17.6), (-2.2,-17.6), (0,-17.6), (2.2,-17.6), (4.4,-17.6), (6.6,-17.6), (8.8,-17.6), (11,-17.6), (13.2,-17.6), (15.4,-17.6), (-15.4,-19.8), (-13.2,-19.8), (-11,-19.8), (-8.8,-19.8), (-6.6,-19.8), (-4.4,-19.8), (-2.2,-19.8), (0,-19.8), (2.2,-19.8), (4.4,-19.8), (6.6,-19.8), (8.8,-19.8), (11,-19.8), (13.2,-19.8), (15.4,-19.8), (17.6,-19.8)]
point_name_pairing = [("sfgfp", sfgfp_points)]
# Robot deck setup constants
TIP_RACK_DECK_SLOT = 9
COLORS_DECK_SLOT = 6
AGAR_DECK_SLOT = 5
PIPETTE_STARTING_TIP_WELL = 'A1'
# Place the PCR tubes in this order
well_colors = {
'A1': 'sfGFP',
'A2': 'mRFP1',
'A3': 'mKO2',
'A4': 'Venus',
'A5': 'mKate2_TF',
'A6': 'Azurite',
'A7': 'mCerulean3',
'A8': 'mClover3',
'A9': 'mJuniper',
'A10': 'mTurquoise2',
'A11': 'mBanana',
'A12': 'mPlum',
'B1': 'Electra2',
'B2': 'mWasabi',
'B3': 'mScarlet_I',
'B4': 'mPapaya',
'B5': 'eqFP578',
'B6': 'tdTomato',
'B7': 'DsRed',
'B8': 'mKate2',
'B9': 'EGFP',
'B10': 'mRuby2',
'B11': 'TagBFP',
'B12': 'mChartreuse_TF',
'C1': 'mLychee_TF',
'C2': 'mTagBFP2',
'C3': 'mEGFP',
'C4': 'mNeonGreen',
'C5': 'mAzamiGreen',
'C6': 'mWatermelon',
'C7': 'avGFP',
'C8': 'mCitrine',
'C9': 'mVenus',
'C10': 'mCherry',
'C11': 'mHoneydew',
'C12': 'TagRFP',
'D1': 'mTFP1',
'D2': 'Ultramarine',
'D3': 'ZsGreen1',
'D4': 'mMiCy',
'D5': 'mStayGold2',
'D6': 'PA_GFP'
}
# Mapping for visualization colors
VISUALIZATION_COLOR_MAP = {
'sfGFP': 'green',
'mRFP1': 'red',
'mKO2': 'orange',
'Venus': 'yellow',
'mKate2_TF': 'purple',
'Azurite': 'blue',
'mCerulean3': 'cyan',
'mClover3': 'lightgreen',
'mJuniper': 'darkgreen',
'mTurquoise2': 'teal',
'mBanana': 'gold',
'mPlum': 'plum',
'Electra2': 'navy',
'mWasabi': 'lime',
'mScarlet_I': 'darkred',
'mPapaya': 'peachpuff',
'eqFP578': 'brown',
'tdTomato': 'tomato',
'DsRed': 'indianred',
'mKate2': 'darkmagenta',
'EGFP': 'chartreuse',
'mRuby2': 'firebrick',
'TagBFP': 'slateblue',
'mChartreuse_TF': 'darkseagreen',
'mLychee_TF': 'palevioletred',
'mTagBFP2': 'darkblue',
'mEGFP': 'limegreen',
'mNeonGreen': 'lawngreen',
'mAzamiGreen': 'mediumseagreen',
'mWatermelon': 'pink',
'avGFP': 'forestgreen',
'mCitrine': 'khaki',
'mVenus': 'olivedrab',
'mCherry': 'crimson',
'mHoneydew': 'honeydew', # This will likely be too light
'TagRFP': 'rosybrown',
'mTFP1': 'dodgerblue',
'Ultramarine': 'mediumblue',
'ZsGreen1': 'springgreen',
'mMiCy': 'peru',
'mStayGold2': 'goldenrod',
'PA_GFP': 'darkgreen'
}
volume_used = {
'sfgfp': 0
}
def update_volume_remaining(current_color, quantity_to_aspirate):
rows = string.ascii_uppercase
for well, color in list(well_colors.items()):
if color == current_color:
if (volume_used[current_color] + quantity_to_aspirate) > 250:
# Move to next well horizontally by advancing row letter, keeping column number
row = well[0]
col = well[1:]
# Find next row letter
next_row = rows[rows.index(row) + 1]
next_well = f"{next_row}{col}"
del well_colors[well]
well_colors[next_well] = current_color
volume_used[current_color] = quantity_to_aspirate
else:
volume_used[current_color] += quantity_to_aspirate
break
def run(protocol):
# Load labware, modules and pipettes
protocol.home()
# Tips
tips_20ul = protocol.load_labware('opentrons_96_tiprack_20ul', TIP_RACK_DECK_SLOT, 'Opentrons 20uL Tips')
# Pipettes
pipette_20ul = protocol.load_instrument("p20_single_gen2", "right", [tips_20ul])
# Deep Well Plate
temperature_plate = protocol.load_labware('nest_96_wellplate_2ml_deep', 6, 'Deep Well Plate')
# Agar Plate
agar_plate = protocol.load_labware('htgaa_agar_plate', AGAR_DECK_SLOT, 'Agar Plate')
agar_plate.set_offset(x=0.00, y=0.00, z=Z_VALUE_AGAR)
# Get the top-center of the plate, make sure the plate was calibrated before running this
center_location = agar_plate['A1'].top()
pipette_20ul.starting_tip = tips_20ul.well(PIPETTE_STARTING_TIP_WELL)
# Helper function (dispensing)
def dispense_and_jog(pipette, volume, location):
assert(isinstance(volume, (int, float)))
# Go above the location
above_location = location.move(types.Point(z=location.point.z + 2))
pipette.move_to(above_location)
# Go downwards and dispense
pipette.dispense(volume, location)
# Go upwards to avoid smearing
pipette.move_to(above_location)
# Helper function (color location)
def location_of_color(color_string):
for well,color in well_colors.items():
if color.lower() == color_string.lower():
return temperature_plate[well]
raise ValueError(f"No well found with color {color_string}")
# Print pattern by iterating over lists
for i, (current_color, point_list) in enumerate(point_name_pairing):
# Skip the rest of the loop if the list is empty
if not point_list:
continue
# Get the tip for this run, set the bacteria color, and the aspirate bacteria of choice
pipette_20ul.pick_up_tip()
max_aspirate = int(18 // POINT_SIZE) * POINT_SIZE
quantity_to_aspirate = min(len(point_list)*POINT_SIZE, max_aspirate)
update_volume_remaining(current_color, quantity_to_aspirate)
pipette_20ul.aspirate(quantity_to_aspirate, location_of_color(current_color))
# Iterate over the current points list and dispense them, refilling along the way
for i in range(len(point_list)):
x, y = point_list[i]
adjusted_location = center_location.move(types.Point(x, y))
dispense_and_jog(pipette_20ul, POINT_SIZE, adjusted_location)
if pipette_20ul.current_volume == 0 and len(point_list[i+1:]) > 0:
quantity_to_aspirate = min(len(point_list[i:])*POINT_SIZE, max_aspirate)
update_volume_remaining(current_color, quantity_to_aspirate)
pipette_20ul.aspirate(quantity_to_aspirate, location_of_color(current_color))
# Drop tip between each color
pipette_20ul.drop_tip()