Week 7 HW: Genetic Circuits Part 2
Assignment Part 1: Intracellular Artificial Neural Networks (IANNs)
What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions?
Traditional genetic circuits operate as Boolean logic gates: they classify inputs as either “on” (1) or “off” (0) and produce outputs that are likewise binary. While this is powerful for implementing discrete decisions — such as activating a kill switch if and only if two specific signals are simultaneously present — Boolean circuits are fundamentally limited in their ability to process the continuous, graded molecular signals that characterise real biological environments. Intracellular concentrations of transcription factors, metabolites, and signalling molecules are not naturally binary; they span continuous ranges that carry information that a simple Boolean threshold necessarily discards. IANNs overcome this limitation by implementing analog computation, in which each molecular “neuron” computes a weighted sum of its continuous-valued inputs, passes that sum through a nonlinear activation function, and produces a graded output that can itself serve as an input to the next layer. This architecture enables a single engineered cell to perform multi-threshold classification — distinguishing not just “signal present” from “signal absent” but grading responses proportionally to signal intensity, and separating input patterns that no Boolean gate could resolve without an exponentially larger circuit. For example, a cell expressing a two-input biomolecular perceptron can draw a separating hyperplane in the continuous input space of two molecular concentrations, classifying cell states that would require many cascaded Boolean gates to approximate. A second key advantage is graceful degradation under noise: because IANNs operate over a continuous input range, they can be designed with soft thresholds that smooth over stochastic fluctuations in molecule numbers — a pervasive problem in cells, where copy numbers of regulatory molecules are often in the tens to hundreds range. Boolean gates, which depend on crossing a hard threshold, are comparatively fragile to such noise. Third, IANNs are in principle extendable toward online learning, in which the synaptic weights (encoded by molecular concentrations or binding affinities) can be updated as a function of experience — an entirely alien concept to hardwired Boolean logic. Taken together, IANNs expand the computational vocabulary available to synthetic biology from a finite set of logic operations to a continuous, composable, and theoretically universal function approximation framework.Describe a useful application for an IANN; include a detailed description of input/output behavior, as well as any limitations an IANN might face to achieve your goal.
One compelling application for an IANN is the continuous intracellular monitoring and correction of iron overload in patients with hereditary hemochromatosis — a genetic disorder characterised by excessive gastrointestinal absorption of dietary iron, leading to toxic iron deposition in the liver, heart, and pancreas. Current treatment requires regular phlebotomy, which is effective but burdensome and cannot respond dynamically to real-time fluctuations in free labile iron. An IANN-based therapeutic cell (for example, an engineered hepatocyte or gut epithelial cell) could be designed as follows. Two inputs are presented to a single-layer intracellular perceptron:
X₁: the intracellular concentration of labile iron pool (LIP), sensed indirectly via an iron-responsive element (IRE)–iron regulatory protein (IRP) system, which naturally controls mRNA translation in proportion to free iron levels. A synthetic construct could link IRP binding to the transcription or translation of an intermediate regulatory RNA, converting iron concentration into a molecular signal. X₂: a constitutive bias input (a fixed-level transcript) that sets the activation threshold — encoding the notion that the circuit should only respond when iron exceeds a safe baseline, analogous to a bias unit in a standard perceptron.
The perceptron computes the weighted sum of these inputs. When the weighted iron signal exceeds the threshold set by the bias, the activation function triggers expression of the output gene: a codon-optimised ferritin heavy-chain transgene, which sequesters excess free iron into inert ferritin complexes and prevents cellular damage. The output is graded — the more the iron concentration exceeds the threshold, the more ferritin is produced — in contrast to a Boolean circuit, which would either produce a fixed amount of ferritin or none at all, regardless of the severity of iron overload. Several important limitations must be acknowledged. First, IANNs currently cannot perform online weight adjustment in living cells at the speed required for therapeutic use; weights are set at the time of circuit design and cannot recalibrate if the patient’s physiology changes. Second, the molecular components encoding the perceptron — endoribonucleases, regulatory RNA hairpins, sequestration species — impose a metabolic burden on the host cell, and this burden grows with the complexity of the network, potentially impairing normal cellular function. Third, molecular noise in cells means that the effective threshold may drift over time as component concentrations fluctuate, making it difficult to guarantee that the circuit reliably distinguishes pathological from physiological iron levels. Fourth, an in vivo implementation raises significant immunogenicity concerns: the endoribonuclease components (e.g., Csy4, which originates from Pseudomonas aeruginosa) could trigger immune responses in a human host. These limitations suggest that near-term applications of IANNs may be better suited to ex vivo cell engineering or biosensor contexts rather than direct in vivo therapeutics.
Below is a diagram depicting an intracellular single-layer perceptron where the X1 input is DNA encoding for the Csy4 endoribonuclease and the X2 input is DNA encoding for a fluorescent protein output whose mRNA is regulated by Csy4. Tx: transcription; Tl: translation.
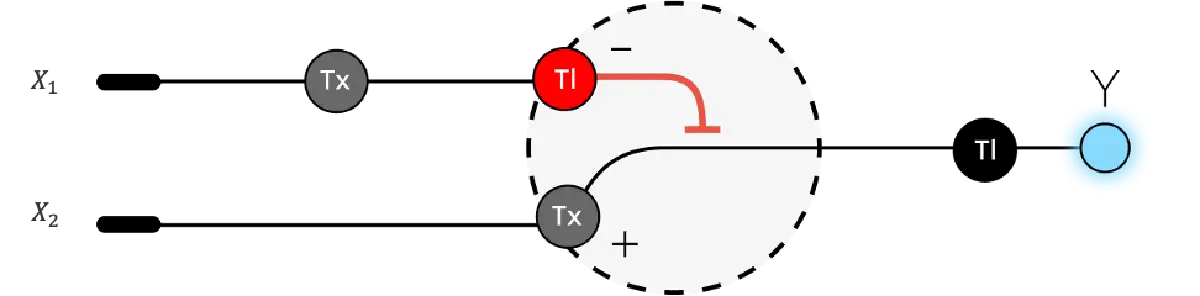

Draw a diagram for an intracellular multilayer perceptron where layer 1 outputs an endoribonuclease that regulates a fluorescent protein output in layer 2.
Assignment Part 2: Fungal Materials
- What are some examples of existing fungal materials and what are they used for? What are their advantages and disadvantages over traditional counterparts?
Fungal materials — chiefly derived from mycelium, the vegetative network of interwoven hyphae produced by filamentous fungi — have been commercialised across several industries over the past decade. The most mature application is mycelium-based composite packaging (e.g., Ecovative Design’s Mushroom® Packaging), in which agricultural waste such as corn husks or hemp hurds is inoculated with fungal spores, allowed to colonise and bind the substrate, then heat-killed and dried to produce a rigid, foam-like material used in place of expanded polystyrene for protective packaging. A second prominent category is myco-leather: pure mycelium mats produced by companies such as Bolt Threads (Mylo™) and MycoWorks (Reishi™) are processed into flexible sheets resembling animal leather and have been used in fashion accessories, including a limited-edition Hermès handbag. Third, mycoprotein — most famously Quorn, derived from Fusarium venenatum — has been on the market since the 1980s as a high-protein, meat-substitute food ingredient. More nascent applications include mycelium-based thermal insulation panels, biocement for construction, and even flexible electronic substrates exploiting the high conductivity of processed mycelium films.
The advantages of mycelium materials over their conventional counterparts are substantial. They are fully biodegradable, decomposing within weeks to months under composting conditions, in contrast to expanded polystyrene (which persists for ~50 years) or synthetic leather (derived from petroleum-based polyurethane). They can be grown on agricultural waste and byproducts — low-cost, abundant feedstocks — requiring no petroleum inputs, which reduces their carbon footprint relative to synthetic foam and plastic alternatives. They are mouldable during growth, meaning complex three-dimensional shapes can be formed without energy-intensive casting or machining. For myco-leather specifically, production avoids the toxic tanning chemicals and greenhouse gas emissions associated with conventional livestock-based leather.
The disadvantages are equally significant and should not be understated. Mycelium composites typically exhibit lower and less consistent mechanical properties than synthetic analogues: their tensile strength, compressive modulus, and moisture resistance vary substantially with fungal species, substrate composition, and growth conditions, making quality control challenging at industrial scale. High moisture absorption is a persistent problem — mycelium-based foams can absorb significant water, compromising their insulating and structural properties in humid environments. Biodegradability, while an environmental advantage, is simultaneously a durability disadvantage: myco-leather goods will degrade under prolonged exposure to moisture, UV light, or biological activity at rates that animal leather or synthetic leather would not. Finally, scaling production while maintaining consistent properties and sterility is technically demanding and costly, and life-cycle assessments suggest that the energy inputs for controlled fungal cultivation can partially offset the environmental benefits, particularly where renewable energy is not available .
- What might you want to genetically engineer fungi to do and why? What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
One application I find particularly compelling is engineering filamentous fungi for targeted heavy-metal bioremediation — specifically, the removal of cadmium, lead, and arsenic from contaminated soils and industrial wastewater. Wild-type fungi already exhibit some capacity for metal biosorption via their cell walls, but this is passive and non-selective. I would want to genetically engineer a species such as Aspergillus niger or Trichoderma reesei to overexpress metallothioneins (small cysteine-rich metal-binding proteins) and ABC-type metal transporters that actively import toxic metals into vacuoles, concentrating them intracellularly for subsequent extraction by harvesting the mycelium rather than using energy-intensive chemical treatments. A second engineering goal would be to add a biosensor output — for instance, linking metal accumulation to the expression of a fluorescent reporter — so that the mycelium itself signals when remediation capacity is saturated and biomass needs to be replaced. This is precisely the kind of continuous, graded signal-response behaviour that an IANN architecture (from Part 1) could implement.
There are several compelling advantages of performing this synthetic biology in fungi rather than bacteria. First, fungi are eukaryotes, meaning they possess the post-translational modification machinery — N-linked glycosylation, disulfide bond formation, proper protein folding in the endoplasmic reticulum — required to produce complex proteins such as metallothioneins and secreted enzymes in their active forms; many such proteins are misfolded or inactive when expressed in E. coli. Second, filamentous fungi grow as mycelial networks that can extend through soil, bridging air-liquid interfaces and penetrating into pore spaces inaccessible to bacterial biofilms — a critical advantage for in situ bioremediation, where the contaminant is spatially distributed and often in a partly air-filled matrix. Third, many filamentous fungi are GRAS-status organisms (Generally Recognised As Safe), reducing regulatory barriers for environmental release relative to engineered bacteria, some of which carry biosafety concerns. Fourth, fungi have extraordinary metabolic versatility: they can catabolise lignin, cellulose, and xenobiotic compounds via oxidative enzymes (laccases, peroxidases) that are absent from most bacteria, making them uniquely suited to environments contaminated with both heavy metals and complex organic pollutants simultaneously. Finally, the physical scaffold of mycelium itself has structural utility — a bioremediation mycelium network can be harvested as a solid biomass enriched in bound metals, rather than requiring centrifugation or filtration of a bacterial cell suspension, simplifying downstream metal recovery.
A counter-consideration is that fungal genetic engineering has historically been more technically challenging than in bacteria, due to lower rates of homologous recombination in many species and the relative scarcity of validated synthetic promoters and genetic parts. However, the development of CRISPR-Cas9 tools adapted for Aspergillus and Trichoderma, alongside growing fungal parts registries, is rapidly closing this gap.