Week 10 HW: Advanced Imaging and Measurement Tech
Part 1: Molecular Weight
eGFP Sequence:
Based only on the predicted amino acid sequence of eGFP (see below), what is the calculated molecular weight? You can use an online calculator like the one here: https://web.expasy.org/compute_pi/
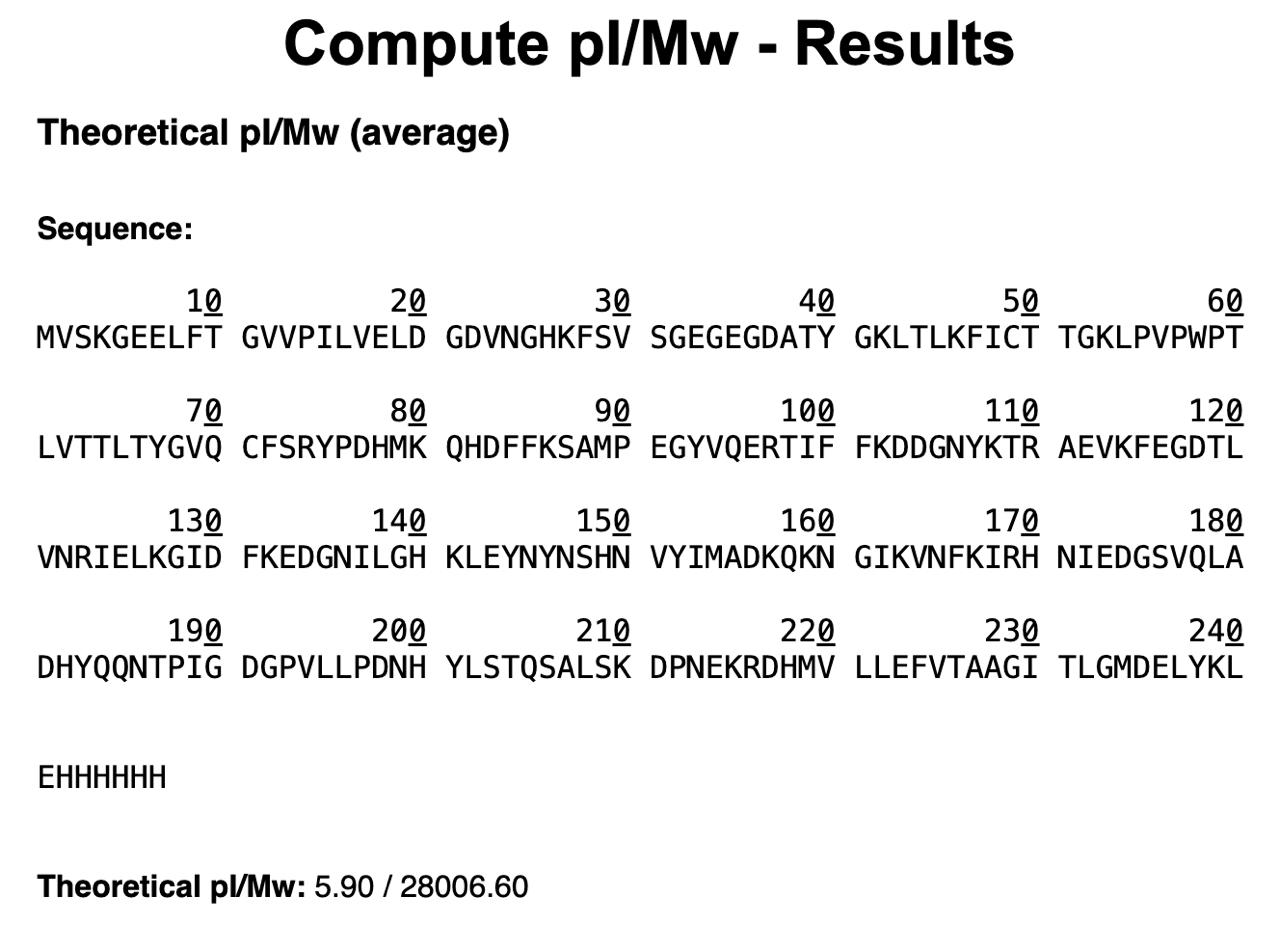

Calculate the molecular weight of the eGFP using the adjacent charge state approach described in the recitation. Select two charge states from the BioAccord data and:
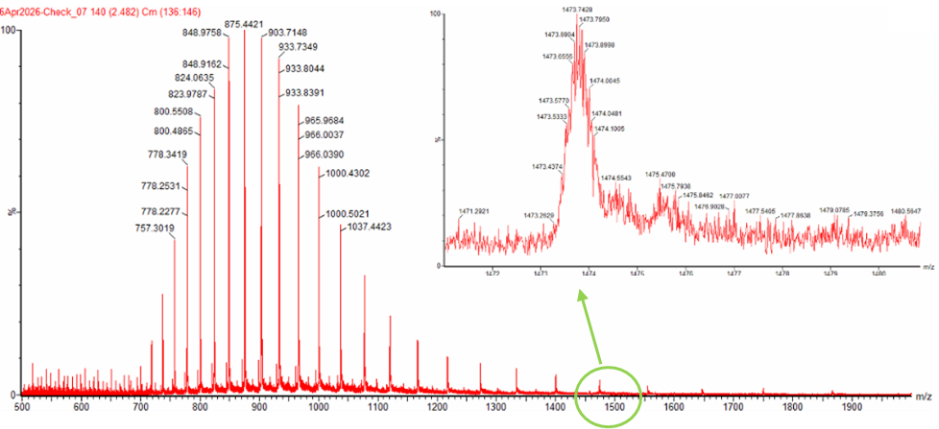
 Figure 1. Mass Spectrum of intact eGFP protein from the Waters Xevo G3 LC-MS (a mass spectrometer with 30,000 resolution) with individual charge state peaks labeled with values.
Figure 1. Mass Spectrum of intact eGFP protein from the Waters Xevo G3 LC-MS (a mass spectrometer with 30,000 resolution) with individual charge state peaks labeled with values.Determine z for each (n, n+1)
Peak 1: (m/z_n)= 903.7148 Peak 2: (m/z_n+1)= 875.4421
Calculating z:
$$n=\left(\frac{m}{z_{n+1}}\right) / \left(\frac{m}{z_n} - \frac{m}{z_{n+1}}\right)$$
$$n=(875.4421) / (903.7148-875.4421)$$
$$n= (875.4421)/ (28.2727)$$
$$n=30.963$$
n= 31
n+1= 32
- Determine the MW of the protein using the relationship between m/z, MW and z
$$M = z \times (m/z - H)$$
Peak 1= 903.7148
M= 31 x (903.7148 - 1.007)
M= 31 x (902.7078)
M= 27983.9418 Da
Peak 2= 875.4421
M= 32 x (875.4421 - 1.007)
M= 32 x (874.4351)
M= 27981.9232 Da
$$M_{avg}= (27983.9418 + 27981.9232) / 2$$ $$= 27982.9325 \text{ Da}$$
- Calculate the mass accuracy of the measurement using the deconvoluted MW from 2.2) and the predicted weight of the protein from 2.1).
M theoretical= 27875.41 Da
M estimated = 27982.9325 Da
Absolute error= (27982.9325 - 27875.41) = 107.5225
$$Accuracy = \frac{|MW_{experiment} - MW_{theo}|}{MW_{theo}} = \frac{107.5225}{27875.41} = 0.003858$$
Homework: Waters Part III — Peptide Mapping - primary structure
We will digest the eGFP protein standard into peptides using trypsin (an enzyme that selectively cleaves the peptide bond after Lysine (K) and Arginine (R) residues. The resulting peptides will be analyzed on the Waters BioAccord LC-MS to measure their molecular weights and fragmented to confirm the amino acid sequence within each peptide – generating a “peptide map”. This process is used to confirm the primary structure of the protein.
There are a variety of tools available online to calculate protein molecular weight and predict a list of peptides generated from a tryptic digest. We will be using tools within the online resource Expasy (the bioinformatics resource portal of the Swiss Institute of Bioinformatics (SIB)) to predict a list of tryptic peptides from eGFP.
How many Lysines (K) and Arginines (R) are in eGFP? Please circle or highlight them in the eGFP sequence given in Waters Part I question 1 above. (Note: adding the sequence to Benchling as an amino acid file and clicking biochemical properties tab will show you a count for each amino acid).
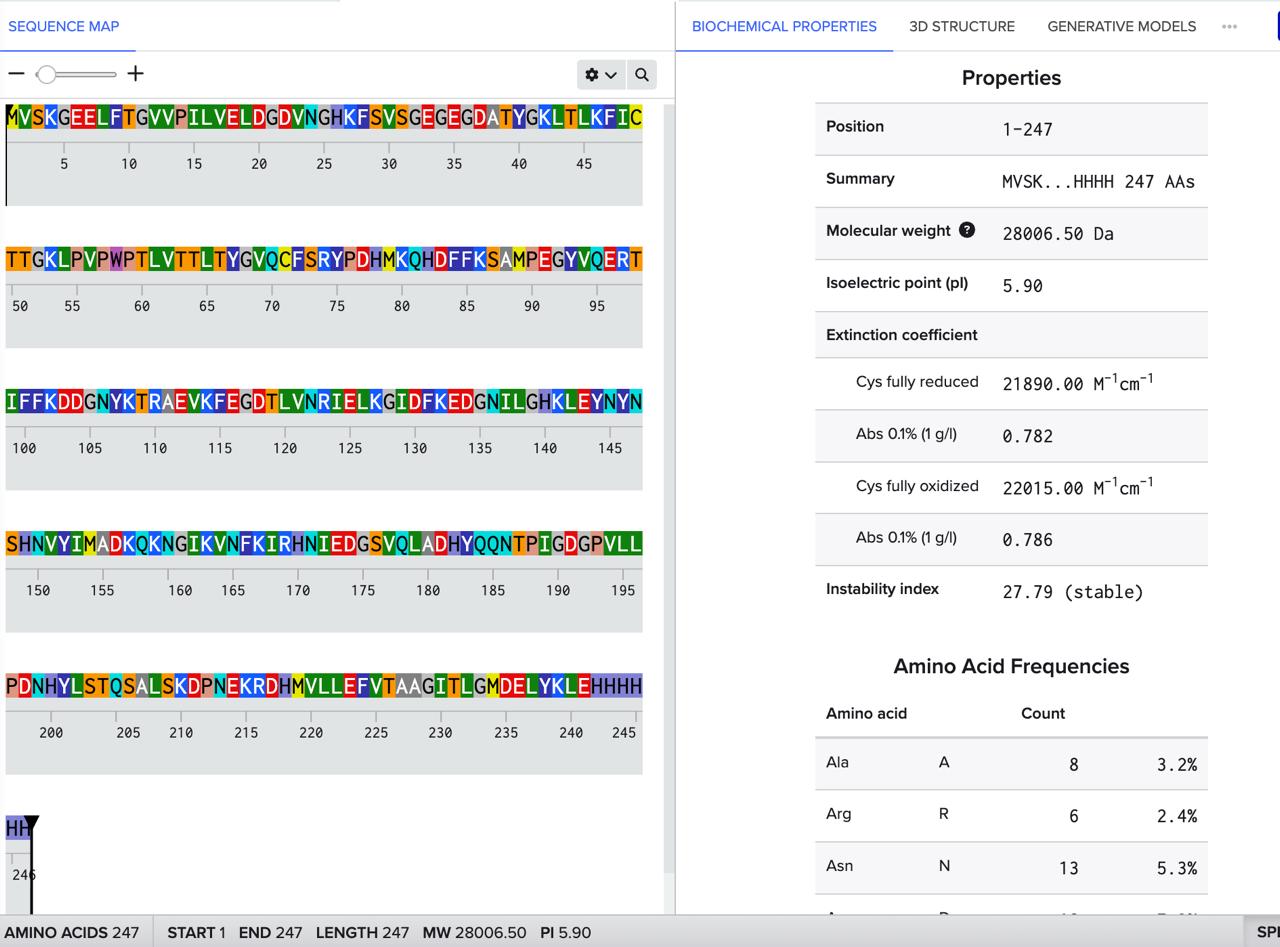

- There are 20 Lysines (K) which constitute 8.1% of the eGFP.
- There are 6 Arginines (R) which constitute 2.4% of the eGFP.
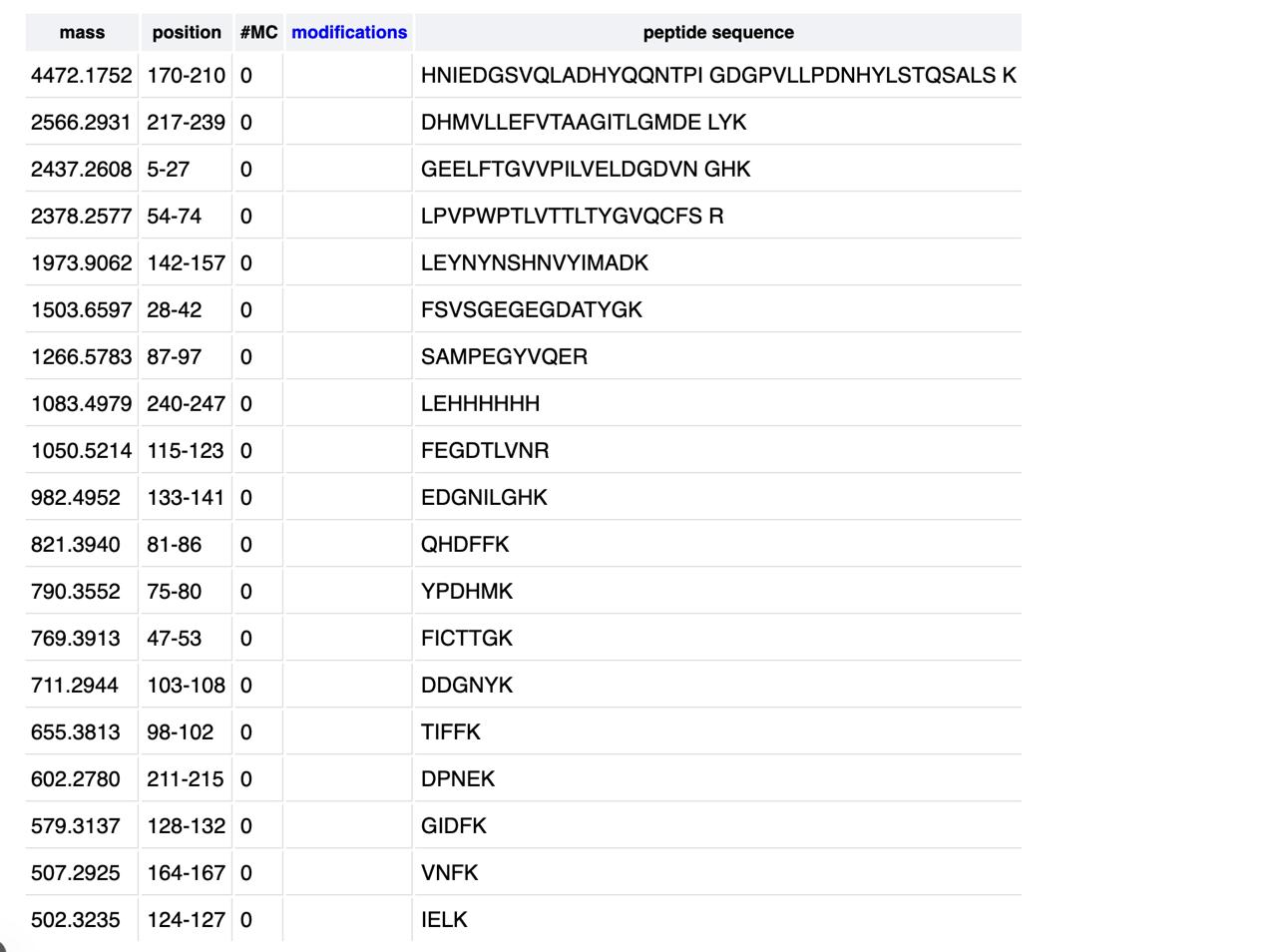
How many peptides will be generated from tryptic digestion of eGFP? a. Navigate to https://web.expasy.org/peptide_mass/ b. Copy/paste the sequence above into the input box in the PeptideMass tool to generate expected list of peptides. c. Use Figure 4 below as a guide for the relevant parameters to predict peptides from eGFP.
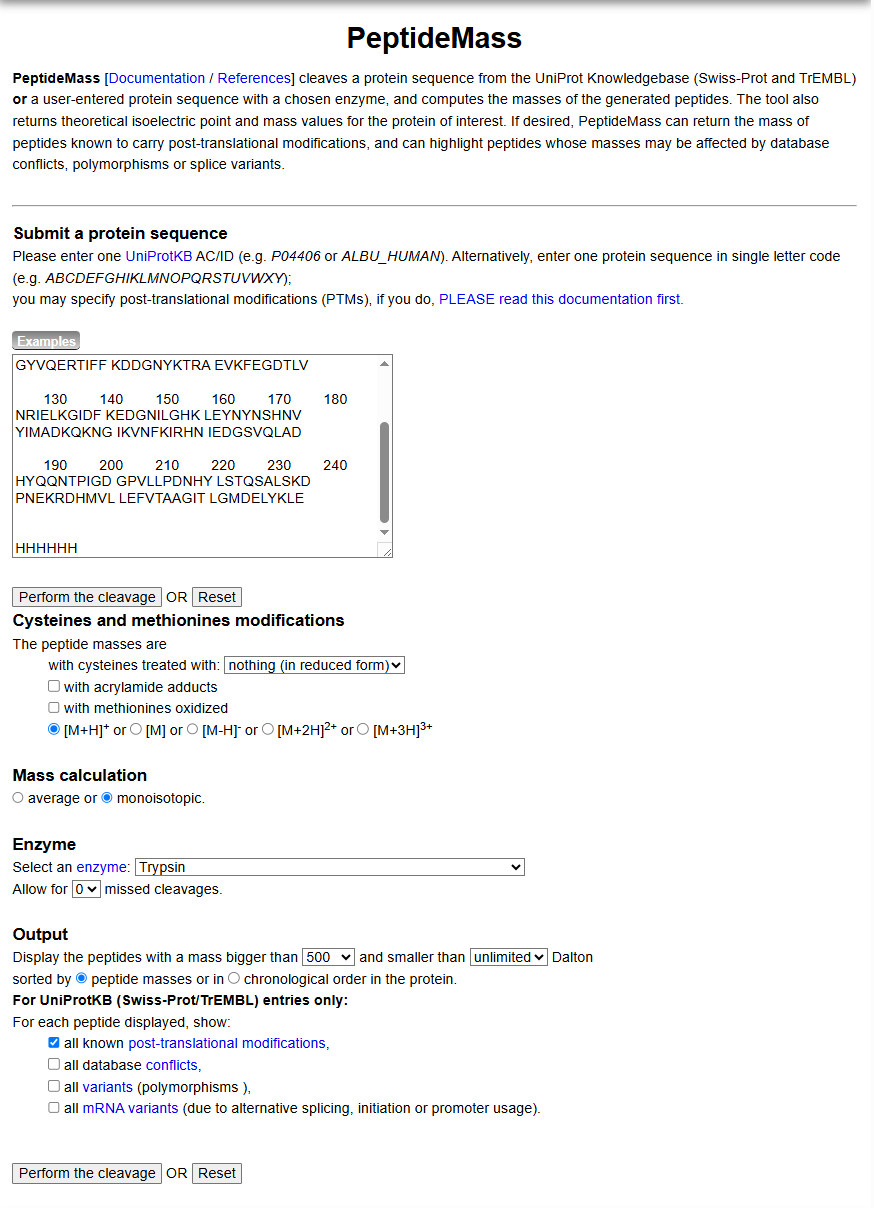

d. Click “Perform the Cleavage” button in the PeptideMass tool and report the number of peptides generated when using trypsin to perform the digest.