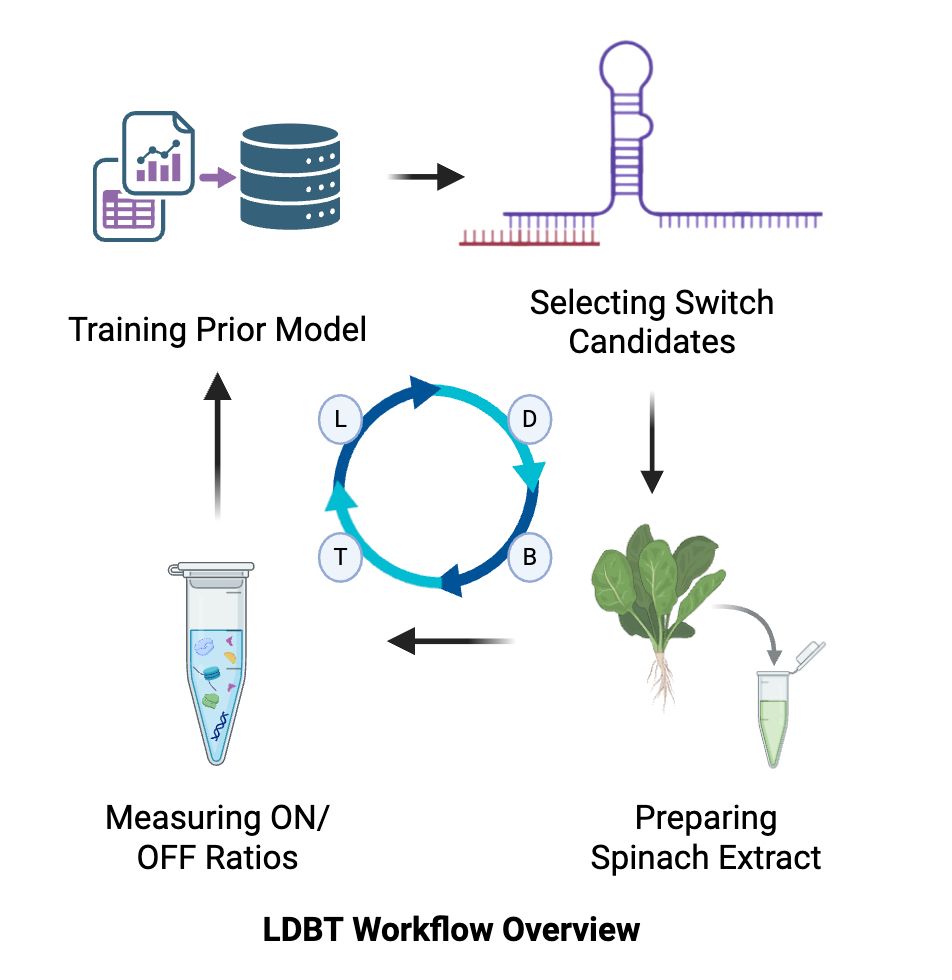
Mapping the Thermodynamic Rules of Toehold Switch Function in Spinach Chloroplast Cell-Free Expression: an LDBT Approach Abstract Chloroplast cell-free expression (CFE) systems have recently been established as powerful rapid-prototyping platforms for plastid genetic parts, yet whether these systems can support synthetic RNA logic remains entirely untested. Toehold switches — de novo-designed riboregulators that activate translation in response to specific trigger RNAs — represent the most sophisticated programmable RNA gates in synthetic biology. Machine learning models trained on E. coli CFE data have begun to extract sequence-structure features predictive of switch performance using frameworks like SANDSTORM (Riley et al., 2025), but whether those learned relationships hold in a chloroplast ribosome context is unknown. This project addresses that gap directly.