Week 1 HW: Principles and Practices
Class assignment
1. First, describe a biological engineering application or tool you want to develop and why.
I’m heavily inspired by Professor Jacobson’s call for a “bio-FPGA” tool, as well as his lecture about cellular automata. I’d like to develop a bio-FPGA that can be programmed to grow into arbitrary 2D patterns on a petri dish, using the machine learning technique mentioned in the lecture to reverse learn the cellular automata rules for growing a specific pattern. The learned CA rules can be encoded by genetically programming the bio-FPGA then using bacteria with the genes to grow an actual cell culture into the pattern, like the butterfly wing letter patterns in the lecture. If this is feasible, 3D patterns would be the next step, and one might even imagine a wild future of programmable plants that grow into the shapes of houses and furniture.
2. Next, describe one or more governance/policy goals related to ensuring that this application or tool contributes to an “ethical” future, like ensuring non-malfeasance (preventing harm). Break big goals down into two or more specific sub-goals.
My primary goal is to protect health and safety. The sub-goals are:
- Prevent the development of biological weapons.
- Prevent outbreak of harmful bacteria.
- Maximize productive use-cases.
A bio-FPGA has the possibility to be used for great benefit with many applications, but could also be abused.
3. Next, describe at least three different potential governance “actions” by considering the four aspects below (Purpose, Design, Assumptions, Risks of Failure & “Success”).
Action 1: Engineer a genetic “off switch” into the bio-FPGA to stop proliferation any time.
- Purpose: We would build a genetic off switch to immediately turn off all genes of the bio-FPGA.
- Design: This would involve researchers and industry as actors to build this prior to releasing the bio-FPGA. The government could also regulate by requiring all bio-FPGA and adjacent tools have such a fail-safe.
- Assumptions: This assumes that such a technological solution could be reliably engineered and triggered.
- Risks: The risks are that the technical fail-safe does not work, or could even cause problems if it does work because it could be abused to disable legitimate use cases.
Action 2: Regulate against use for biological warfare.
- Purpose: Although there are already regulations in place, we could craft regulation to specifically account for bio-FPGA technology.
- Design: This would involve the government to understand the technology, the dangers, and pass appropriate laws preventing malicious use of bio-FPGAs.
- Assumptions: This assumes that lawmakers would be motivated to pass regulation and that the public would be accepting of such regulation. It also assumes that lawmakers are able to craft good laws or adapt accordingly.
- Risks: The risk is that excessive regulation could stifle adoption and research for beneficial use cases. Another risk is that lawmakers don’t understand the science and pass inappropriate laws.
Action 3: Host a conference for researchers and industry to share new developments.
- Purpose: To share beneficial use cases, foster collaboration, and disseminate research learnings.
- Design: This requires coordinating and organizing the research and industry community, as well as raising funds to host a venue.
- Assumptions: I assume that researchers would be interested in attending and discussing.
- Risks: The conference could be used to develop malicious use-cases, or ethics could be overlooked in favor of scientific progress at all costs.
4. Next, score (from 1-3 with, 1 as the best, or n/a) each of your governance actions against your rubric of policy goals.
| Does the action: | Action 1 | Action 2 | Action 3 |
|---|---|---|---|
| Enhance Biosecurity | |||
| • By preventing incidents | 3 | 1 | 2 |
| • By helping respond | 1 | 3 | 3 |
| Foster Lab Safety | |||
| • By preventing incident | 3 | 1 | 2 |
| • By helping respond | 1 | 3 | 3 |
| Protect the environment | |||
| • By preventing incidents | 3 | 1 | 2 |
| • By helping respond | 1 | 3 | 3 |
| Other considerations | |||
| • Minimizing costs and burdens to stakeholders | 2 | 3 | 1 |
| • Feasibility? | 2 | 1 | 1 |
| • Not impede research | 1 | 3 | 1 |
| • Promote constructive applications | 3 | 1 | 1 |
Week 2 lecture prep
Homework Questions from Professor Jacobson
- Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy?
According to the slides, the error rate of polymerase is 1:106 (online it suggest that it can be even worse depending on the polymerase), or 1 in 1 million. The length of the human genome is 3.2 Gbp (3.2 * 109), so at that rate there would be ~3.2 * 103 (3,200) errors in the human genome per copy. That would be a lot of errors, but there are additional pathways that perform error correction, such as MutS.
- How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice what are some of the reasons that all of these different codes don’t work to code for the protein of interest?
The slides mention the average human protein is 1036 base pairs. That is 345 codons, so 345 amino acids.
Amino acids can have multiple corresponding codons. There are 20 amino acids and 4 possible nucleotides, so there are about 3 possible codons per amino acid.
That is an estimate of 3345 ways to code for a given protein, a huge number.
However, despite a synonymous codon coding for the same amino acid, the base pairs choice can affect the chemical bonds of the mRNA structure, affecting RNA cleavage rules.
Another view is that there are 4^1035 possible nucleotides, which are very unlikely to code for the specific protein even with synonymous codons due to sheer possibility space.
Homework Questions from Dr. LeProust
- What’s the most commonly used method for oligo synthesis currently?
The phosphoramidite method.
- Why is it difficult to make oligos longer than 200nt via direct synthesis?
The coupling step is not possible to have perfect efficiency. That step is repeated per cycle, and each additional base requires the cycle to repeat. This means that longer oligos become dramatically harder to make, even with extremely high efficiencies:
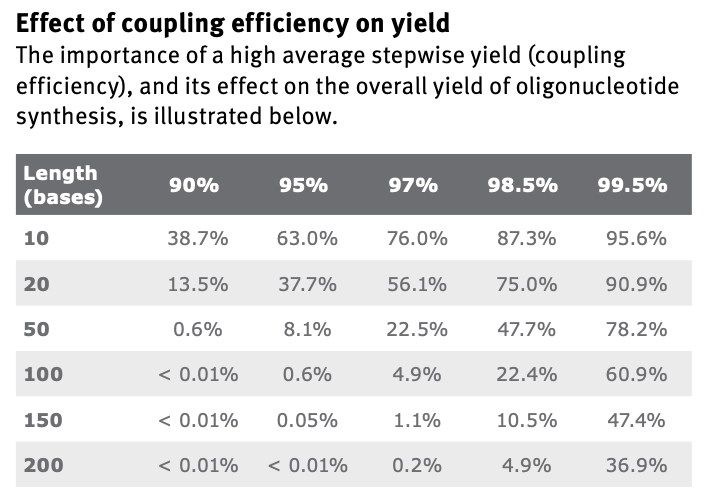

I found the table from this PDF.
- Why can’t you make a 2000bp gene via direct oligo synthesis?
The above answer explains why we can’t synthesize longer oligos. At 2000bp, the probabilities become near impossible even at the highest efficiencies.
Homework Question from George Church
Choose ONE of the following three questions to answer; and please cite AI prompts or paper citations used, if any.
I’m answering question 1.
- [Using Google & Prof. Church’s slide #4] What are the 10 essential amino acids in all animals and how does this affect your view of the “Lysine Contingency”?
The ten essential amino acids are:
- Histidine (H)
- Isoleucine (I)
- Leucine (L)
- Lysine (K)
- Methionine (M)
- Phenylalanine (F)
- Threonine (T)
- Tryptophan (W)
- Valine (V)
- Arginine (R), sometimes considered conditionally essential.
The “Lysine Contingency” is apparently from Jurassic Park, which was a plot element in the movie that was a genetic modification to make the dinosaurs not to be able to produce Lysine so they would die off without human provided Lysine supplements.
However, Lysine is one of the 10 essential amino acids so animals cannot produce it, making this is a scientifically dubious plot point (the genetic modification would have done nothing).
- [Given slides #2 & 4 (AA:NA and NA:NA codes)] What code would you suggest for AA:AA interactions?
- [(Advanced students)] Given the one paragraph abstracts for these real 2026 grant programs sketch a response to one of them or devise one of your own: