Week 3 HW: Lab Automation
Opentrons Artwork
My artwork is here: https://rcdonovan.com/?id=vmns94wqt45wpqc


I used Ronan’s tool to make this. I uploaded an image of tomatoes but it didn’t render well, so I modified it significantly by hand with the editor.
Then, I attended the Saturday session on Zoom with Ronan, Michelle, and Ice at Ginkgo Bioworks. Here’s the end result:


Post-Lab Questions
- Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
I read Assembly of small silica nanoparticles using lipid-tethered DNA ‘bonds’. This paper used a novel assembly using DNA by by embedding silica nanoparticles in a lipid bilayer, embedding the cholesterol end of a DNA-cholesterol molecule within the bilayer, then assembling the nanoparticles with complementary sticky end “bridge” DNA. This is best explained with the image from the paper below:
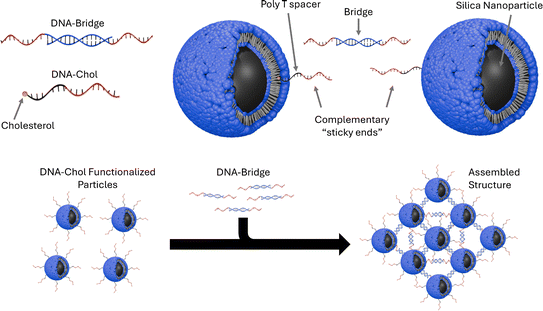

They used Opentrons to rapidly iterate on and evalute different concentrations of DNA-Chol, NaCL, and bridge DNA in the assembly mixture. These were then screened with SAXS to determine the structural qualities of each sample.
- Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more. You may reference this week’s recitation slide deck for lab automation details.
One project idea I have is genetically modifying trees for use in urban areas. I’d like to use automation tools to permute different genetic combinations. I envision custom modules that could germinate and monitor an array of seeds for different qualities.
For example, I create a module with grow lights and watering capabilities that can care for the different seed variants and cameras to compare growth rates.