Week 6 HW: Genetic Circuits Design part I
My Homework
DNA Assembly
Answer these questions about the protocol in this week’s lab:
- What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
Phusion High-Fidelity PCR master mix contains Phusion DNA polymerase, deoxynucleotides and HF reaction buffer with MgCl2. This master mix is used for long or difficult PCR amplifications and applications where high sequence fidelity is critical, such as cloning, mutagenesis, or amplicon sequencing.
2. What are some factors that determine primer annealing temperature during PCR?
It is determined by temperature of melting of the primers which depends mainly on the GC content, length of the primer and sequence composition. Salt concentrations and secondary structures could affect primer annealing temperature as well.
3. There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
When using PCR, primers must be designed to amplify the desired fragment (either the insert or the vector backbone). These primers can also include additional sequences such as overlaps for Gibson Assembly or restriction sites for downstream cloning. A high-fidelity DNA polymerase (e.g., Phusion) is typically used to minimize errors during amplification. PCR is highly flexible because it allows you to amplify any sequence regardless of the presence of restriction sites.
In contrast, restriction enzyme digestion relies on the presence of specific recognition sites in the DNA. The vector and insert must contain compatible restriction sites, and importantly, these sites should not be present within the fragment of interest. If unwanted restriction sites are present, the sequence may need to be modified (e.g., by site-directed mutagenesis or codon optimization). This makes restriction-based cloning more constrained but often more straightforward when suitable sites are available.
After generating linear fragments, the downstream workflow differs slightly between the two methods. For PCR products, it is usually necessary to purify the amplicon and often treat it with enzymes such as DpnI (to remove template DNA) or perform gel extraction to ensure specificity. For restriction digestion, fragments are typically purified after digestion, and if using traditional cloning, may require dephosphorylation of the vector to prevent self-ligation.
The assembly step also differs. PCR products are commonly used in seamless cloning methods such as Gibson Assembly, where overlapping regions allow fragments to anneal and be enzymatically joined. In contrast, restriction-digested fragments are usually ligated using DNA ligase, which joins compatible sticky or blunt ends.
In terms of when to use each method, PCR is preferable when flexibility is needed, such as when introducing mutations, adding overlaps, or working with sequences lacking suitable restriction sites. Restriction digestion is often preferred for routine cloning when appropriate sites are already available, as it can be more straightforward and reliable.
Regarding cost and time, both methods can be comparable depending on reagents, although restriction enzymes are often slightly cheaper. PCR may take longer due to amplification cycles, especially for large fragments, but it provides significantly more versatility.
4. How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
After PCR, digestion and purification, DNA fragments should be analyzed by agarose gel electrophoresis to confirm the correct size of each fragment. DNA concentration can be measured (e.g., by Nanodrop or Qubit) to ensure proper stoichiometry. It is recommended sequencing the fragment before Gibson assembly to ensure that sequence is free of errors.
5. How does the plasmid DNA enter the E. coli cells during transformation?
It is not necessary to purify Gibson assembly reactions because chemical transformation was performed. Chemical competent cells contain salts such as CaCl2 and MgCl2. Positive ions neutralizes negative charges of DNA and phospholipids in the cell membrane allowing to create a channel during heat shock. Heat-shock creates temporary pores allowing plasmid DNA enter the cells during transformation.
6. Describe another assembly method in detail (such as Golden Gate Assembly)
1. Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
2. Model this assembly method with Benchling or Asimov Kernel!
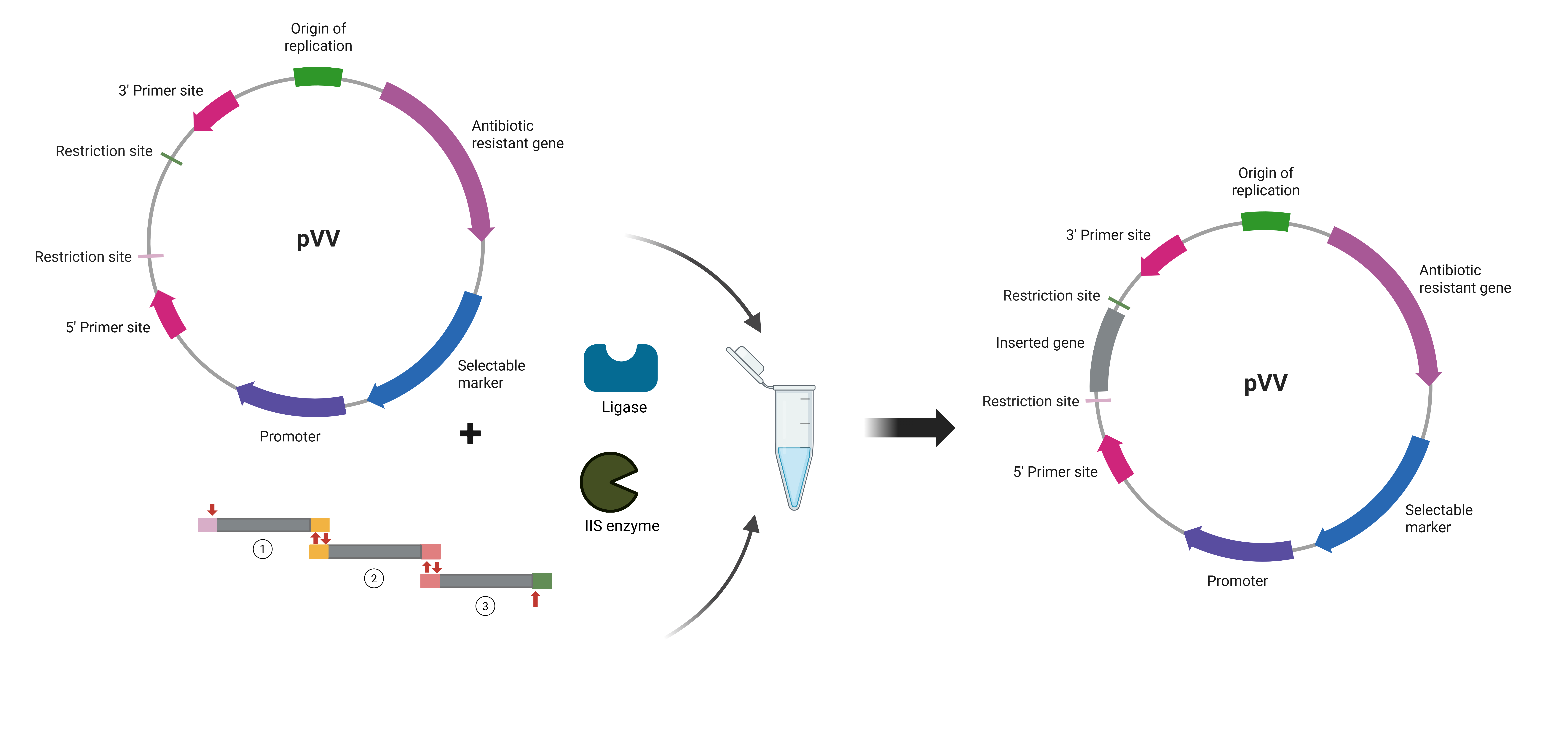
Golden Gate Assembly is a cloning method that uses Type IIS restriction enzymes, which cut DNA outside of their recognition sites. This allows the creation of custom overhangs that can be designed to guide the ordered assembly of multiple DNA fragments in a single reaction. During the process, DNA fragments and a vector are mixed with a Type IIS enzyme and DNA ligase in a one-pot reaction that cycles between digestion and ligation steps. Because the recognition sites are removed during assembly, the final construct is no longer susceptible to digestion, increasing the efficiency of correct assembly. This method enables seamless cloning without leaving extra sequences (scarless assembly) when designed appropriately. Golden Gate is particularly useful for assembling multiple fragments in a defined order, making it ideal for modular cloning and synthetic biology applications. Overall, it is a fast, efficient, and highly scalable method for constructing complex DNA assemblies.

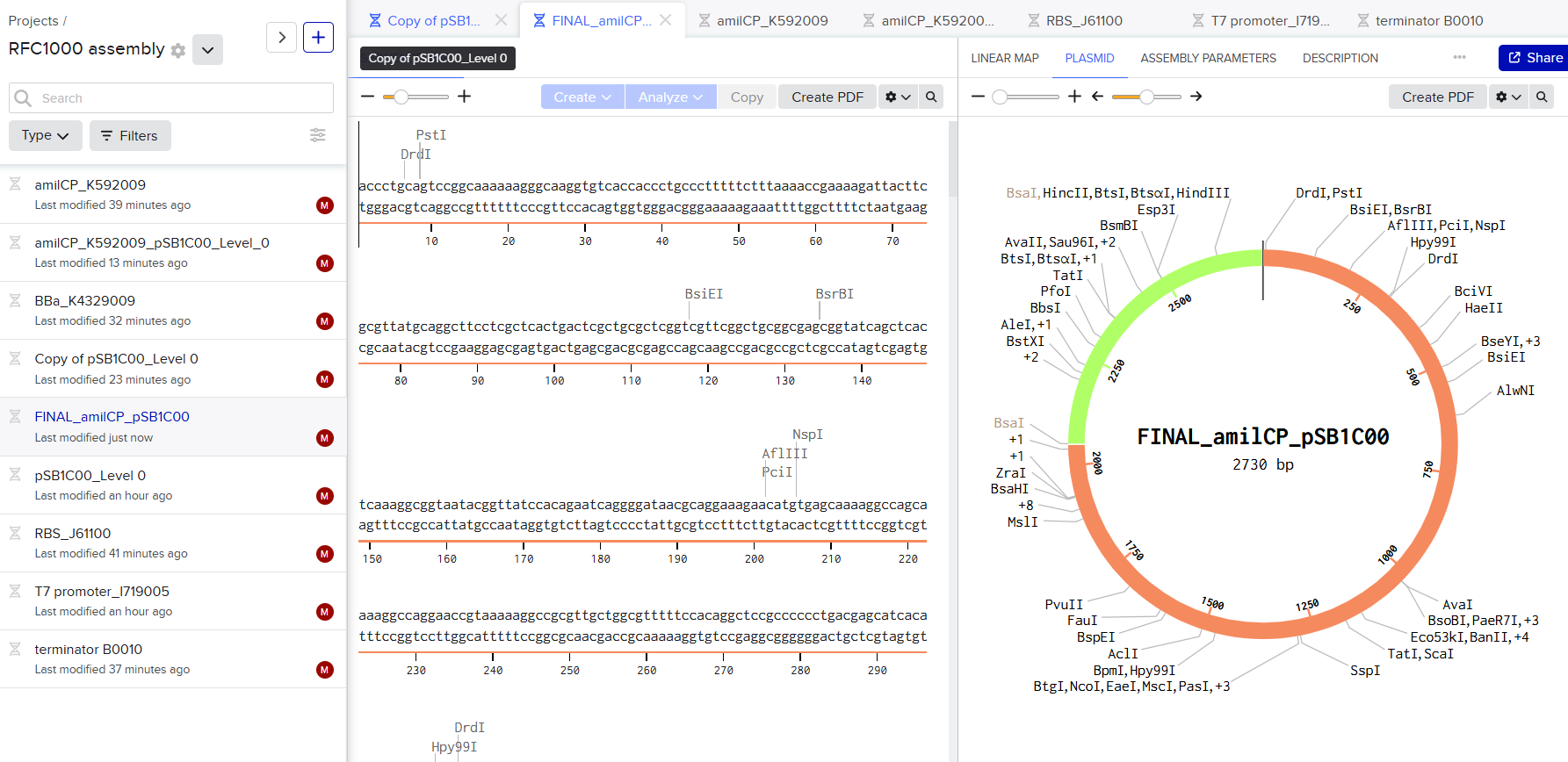
I used the iGEM RFC1000 Type IIS assembly method, which relies on restriction enzymes such as SapI and BsaI to enable hierarchical DNA assembly. In my design, I focused on creating a Level 0 construct by cloning the CDS (amilCP) into the universal acceptor vector pSB1C00 using SapI. The CDS fragment was designed with SapI recognition sites and appropriate overhangs so that, after digestion, it could be ligated into the vector while removing the RFP cassette. Importantly, the inserted CDS is flanked by BsaI recognition sites and standardized fusion sites, which prepares it for downstream assembly. These fusion sites will generate specific overhangs during BsaI digestion, allowing correct and directional assembly into a Level 1 transcriptional unit.