Week 2 HW — DNA Read, Write & Edit
Week 2 — DNA Read, Write & Edit
This week focused on the fundamental processes of reading, writing, and editing DNA.
Using Benchling, restriction digest simulations, and sequence design tools, I explored how DNA can be interpreted not only as biological information but also as a visual and computational medium.
Part 1 — Benchling & In-silico Gel Art
This exercise explores how restriction enzyme digestion can be simulated digitally and how gel electrophoresis patterns emerge from these cuts.
I began by creating a Benchling account and importing the Lambda DNA genome, which is commonly used as a reference molecule in molecular biology experiments. Once imported, Benchling allowed me to visualize the full DNA sequence and explore the locations where restriction enzymes cut the DNA.
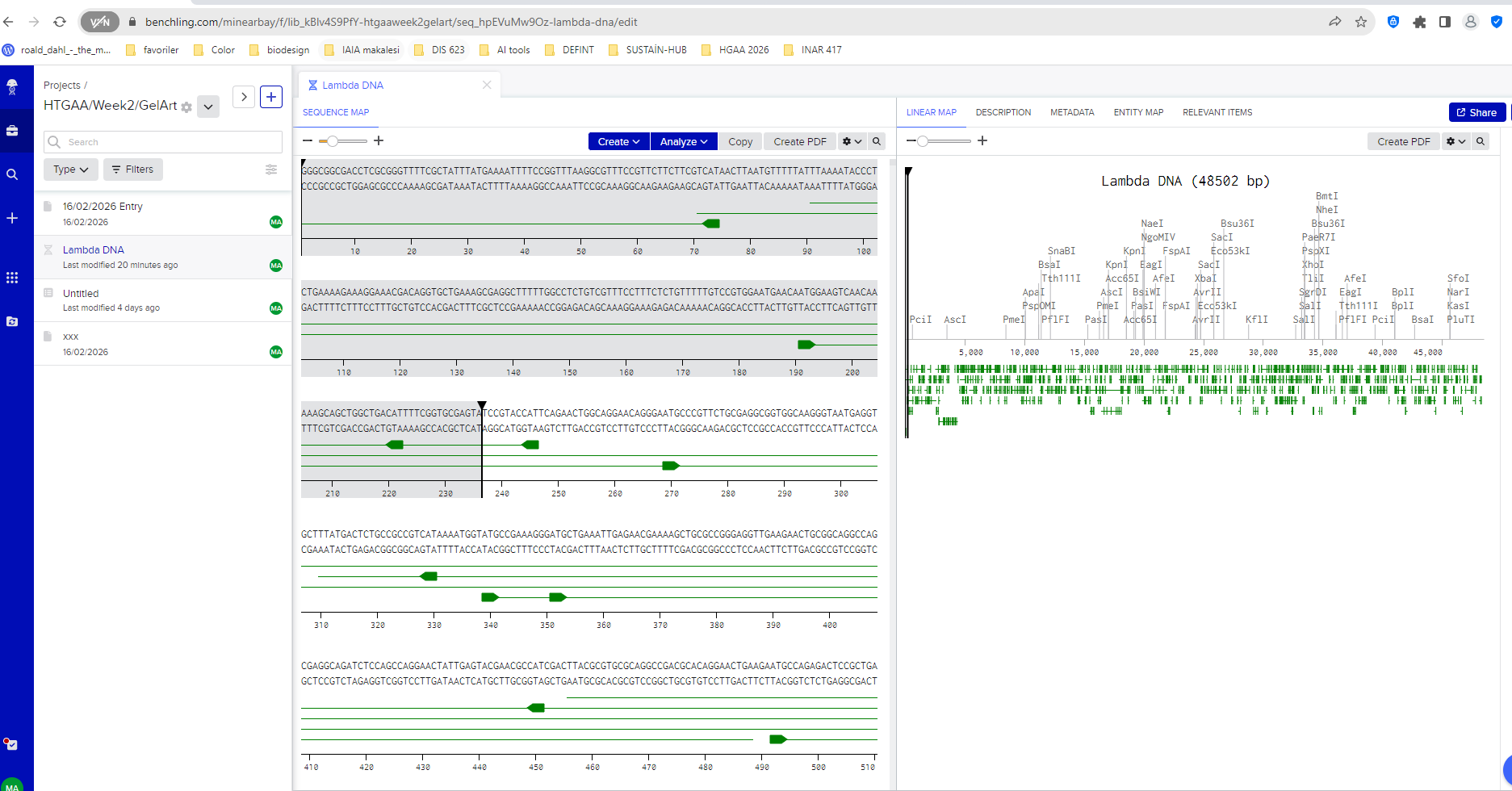
Lambda DNA sequence imported into Benchling.
After importing the sequence, I opened the restriction digest tool. The interface lists many restriction enzymes, each recognizing a specific DNA sequence and cutting the DNA at that location.
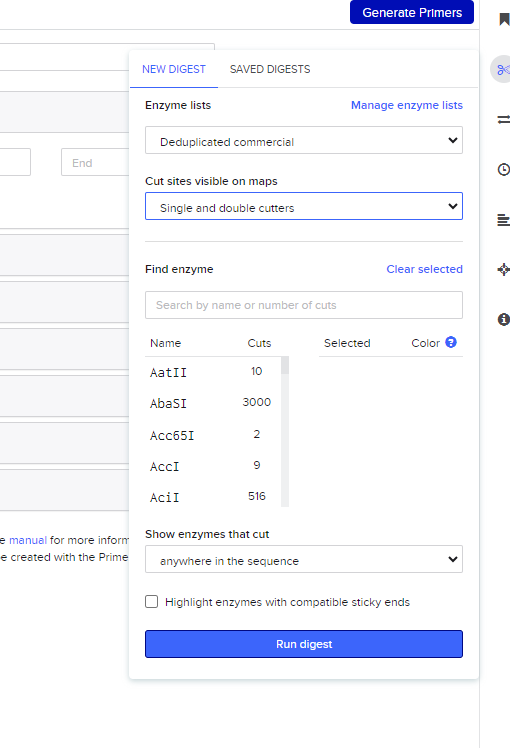
Restriction enzyme selection interface in Benchling.
For this assignment I simulated restriction digests using enzymes discussed in the lab protocol:
EcoRI
HindIII
BamHI
KpnI
EcoRV
SacI
SalI
Each enzyme cuts the DNA at specific recognition sites, generating fragments of different sizes.
Benchling then generates a virtual gel electrophoresis result, where each band corresponds to a DNA fragment produced by the digestion. Larger fragments remain closer to the top of the gel, while smaller fragments travel further down.

EcoRI restriction digest simulation.
I then experimented with multiple enzyme combinations to observe how different fragment distributions create more complex gel patterns.

Multi-enzyme restriction digest simulation.
Comparing different enzymes also revealed how each enzyme produces a unique pattern of DNA fragments.
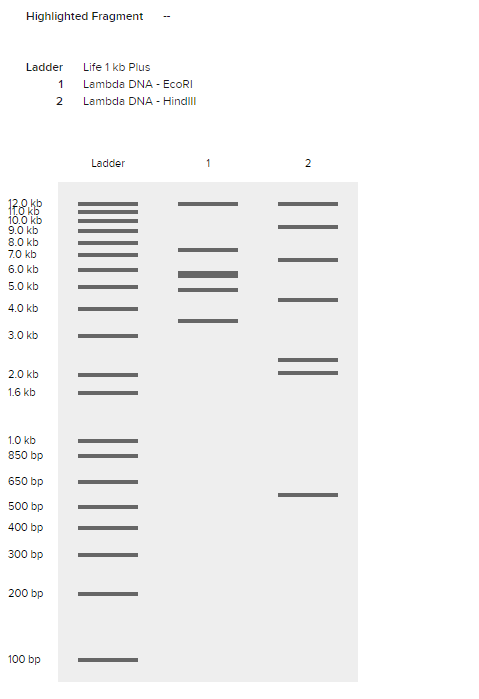
Comparison of EcoRI and HindIII digest patterns.
Inspired by Paul Vanouse’s Latent Figure Protocol, I began to interpret these gel patterns not only as analytical outputs but also as visual structures. Each gel lane behaves like a vertical column in a visual composition, where fragment size determines band position.
To experiment with pattern design more quickly, I also used Ronan Donovan’s Gel Art interface, which allows rapid testing of enzyme combinations and visualization of gel patterns.
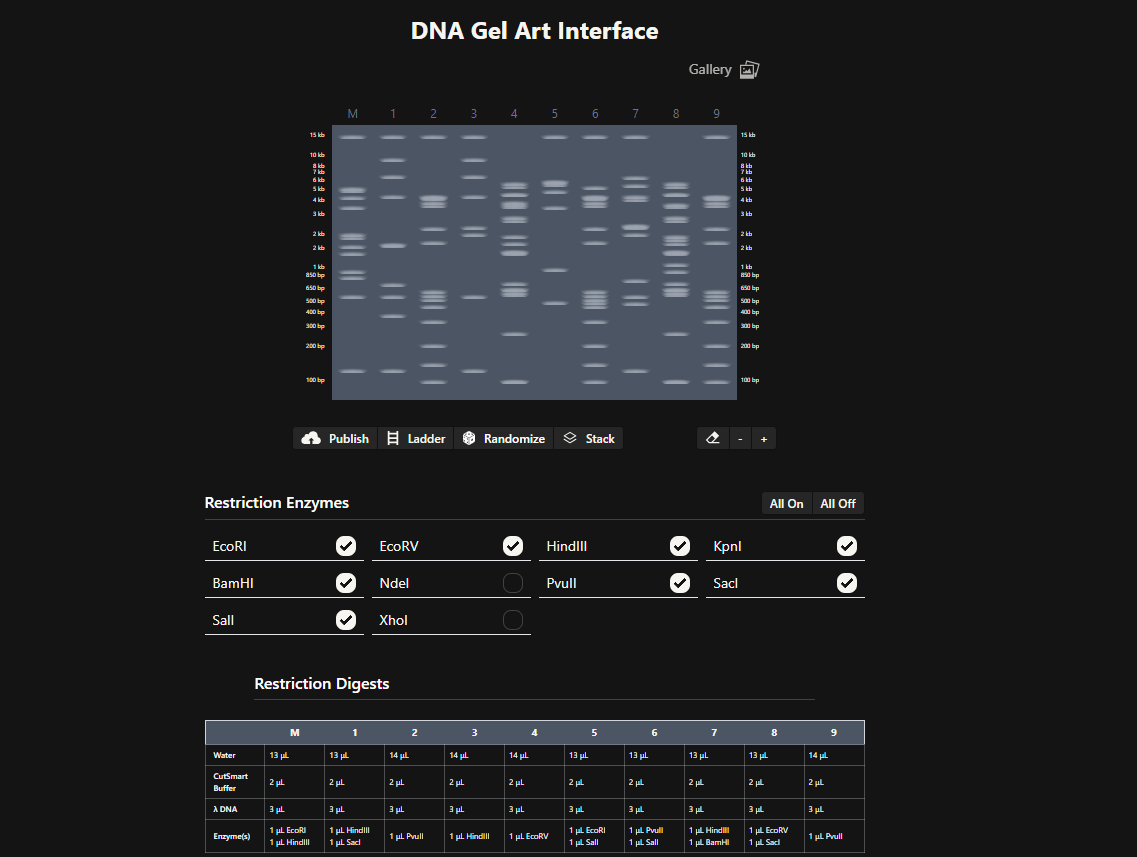
Ronan Donovan's gel-art interface used for rapid pattern exploration.
Through this process I began to see gel electrophoresis not only as a scientific analysis method, but also as a generative visual system, where biological information becomes spatial patterns.
Part 3 — DNA Design Challenge
3.1 Chosen Protein
For this assignment, I chose VioA, the first enzyme in the violacein biosynthesis pathway.
I selected this protein because it connects directly to my broader research interest in producing biological color from waste-based substrates, especially coffee waste. Violacein is a naturally occurring purple pigment, and VioA is important because it catalyzes the first step of the pathway by oxidizing L-tryptophan, initiating the sequence of reactions that leads to pigment production.
This made VioA especially relevant for me: rather than choosing a generic reporter protein, I wanted to work with a protein that is conceptually close to my own research on sustainable pigment systems and microbial color production.
For the sequence source, I used UniProt, which provides a reviewed entry for VioA from Chromobacterium violaceum.
Protein source:
UniProtKB: Q9S3V1
Entry name: VIOA_CHRVO
Protein name: Flavin-dependent L-tryptophan oxidase VioA
Protein sequence (UniProt format):
3.2 Reverse Translation
To understand how proteins are encoded in DNA, I performed a reverse translation of the VioA protein sequence.
According to the Central Dogma of Molecular Biology, DNA is transcribed into RNA and then translated into protein. Reverse translation applies this process in the opposite direction by estimating the nucleotide sequence that could produce a given protein.
Because multiple codons can encode the same amino acid, reverse translation does not produce a single unique DNA sequence but rather a plausible coding sequence.
To perform this step, I used an online reverse translation tool:
https://www.bioinformatics.org/sms2/rev_trans.html
Using the VioA amino acid sequence, I generated a nucleotide sequence that could encode the protein.
Example reverse-translated DNA sequence: ATGAAGCACAGCAGCGACATCTGCATCGTCGGCGCCATCTCCGGCCTCACCTGCGCCTCGCATCTGCTCGACTCCCCGGCGTGCCGCGGCGGCTCTCTCCCGCATCTTCGACATGCAGCAGGAGGCGGCGGGGCGCATCCGCTCCAAGATGCTGGACGGCAAAGCCAGCATCGAGCTCGGCGCCGGCCGCTACAGCCCGCAGCTGCACCCGCACTTCCAGAGCGCCATGCAGCACTACAGCCAGAAGAGCGAGGTGTACCCCTTCACCCAGCTGAAGTTCAAGAGCCACGTGCAGCAGAAGCTGAAGCGCGCGATGAACGAGCTGAGCCCGCGCCTGAAGGAGCACGGCAAGGAGAGCTTCCTGCAGTTCGTGTCCCGCTACCAGGGCCACGACTCCGCCGTGGGCATGATCCGCTCCATGGGCTACGACGCCCTGTTCCTGCCCGACATCTCCGCCGAGATGGCGTACGACATCGTCGGCAAGCACCCCGAGATCCAGAGCGTGACCGACAACGACGCCAACCAGTGGTTCGCCGAGACCGGCTTCGCCGGGCTCATCCAGGGGATCAAGGCCAAGGTGAAGGCCGCCGGCGCCCGCTTCAGCCTGGGCTACCGCCTGCTGTCCGTGCGCACCGACGGCGACGGCTACCTGCTGCAGCTGGCCGGCGACGACGGCTGGAAGCTGGAGCACCGCACCCGCCACCTGATCCTGGCCATCCCCCCGAGCGCCATGGCCGGGCTGAACGTGGACTTCCCCGAGGCCTGGTCCGGCGCGCGCTACGGCAGCCTGCCGCTGTTCAGCGGCTTCCTGACCTACGGCGAGCCGTGGTGGCTGGACTACAAGCTGGACGACCAAGTGCTCATCGTGGACAACCCCCTGCGCAAGATCTACTTCAAGGGCGACAAGTACCTGTTCCTGTACACCGACAGCGAGATGGCCAATTACCGCTGGGGCTGCGTGGCCGAGGGCGAGGACGGCTACCTGGAGCAGATCCGCACCCACCTGGCGAGCGCCCTGGGCATCGTCCGCGAGCGCATCCCGCAGCCCCTGGCGCACGTGCACAAGTACTGGGCGCACGGCGTGGAGTTCGCCGACTCCGACATCGACCACCCGTCGGCGCTGTCCCACCGCGACAGCGGCATCATCGCCTGCAGCGACGCCTACACCGAGCACTGCGGCTGGATGGAGGGCGGCCTGCTGAGCGCGCGCGAGGCCAGCCGCCTGCTGCTGCAGCGCATCGCCGCC
3.3 Codon Optimization
Once the nucleotide sequence corresponding to the VioA protein was generated through reverse translation, the next step was codon optimization.
Although different organisms can produce the same protein, they often prefer different codons for the same amino acid. This phenomenon is known as codon bias. If a DNA sequence contains codons that are rarely used by the host organism, the translation machinery may work less efficiently, resulting in lower protein expression.
To improve protein production, the DNA sequence can be codon optimized for the organism in which the protein will be expressed.
For this project, I optimized the sequence for Escherichia coli (E. coli), since it is one of the most widely used host organisms for recombinant protein production in molecular biology and synthetic biology experiments.
I used the Twist Bioscience Codon Optimization Tool to generate an optimized sequence while avoiding problematic restriction sites.
Tool used:
https://www.twistbioscience.com
The codon-optimized sequence maintains the same amino acid sequence of the VioA protein but replaces certain codons with those more frequently used in E. coli. This increases the likelihood that the protein can be efficiently produced in a bacterial expression system.
Example codon-optimized DNA sequence: ATGAAGCACAGCAGCGACATCTGCATCGTCGGCGCCATCTCCGGCCTCACCTGCGCCTCGCATCTGCTCGACTCCCCGGCGTGCCGCGGCGGCTCTCTCCCGCATCTTCGACATGCAGCAGGAGGCGGCGGGGCGCATCCGCTCCAAGATGCTGGACGGCAAAGCCAGCATCGAGCTCGGCGCCGGCCGCTACAGCCCGCAGCTGCACCCGCACTTCCAGAGCGCCATGCAGCACTACAGCCAGAAGAGCGAGGTGTACCCCTTCACCCAGCTGAAGTTCAAGAGCCACGTGCAGCAGAAGCTGAAGCGCGCGATGAACGAGCTGAGCCCGCGCCTGAAGGAGCACGGCAAGGAGAGCTTCCTGCAGTTCGTGTCCCGCTACCAGGGCCACGACTCCGCCGTGGGCATGATCCGCTCCATGGGCTACGACGCCCTGTTCCTGCCCGACATCTCCGCCGAGATGGCGTACGACATCGTCGGCAAGCACCCCGAGATCCAGAGCGTGACCGACAACGACGCCAACCAGTGGTTCGCCGAGACCGGCTTCGCCGGGCTCATCCAGGGGATCAAGGCCAAGGTGAAGGCCGCCGGCGCCCGCTTCAGCCTGGGCTACCGCCTGCTGTCCGTGCGCACCGACGGCGACGGCTACCTGCTGCAGCTGGCCGGCGACGACGGCTGGAAGCTGGAGCACCGCACCCGCCACCTGATCCTGGCCATCCCCCCGAGCGCCATGGCCGGGCTGAACGTGGACTTCCCCGAGGCCTGGTCCGGCGCGCGCTACGGCAGCCTGCCGCTGTTCAGCGGCTTCCTGACCTACGGCGAGCCGTGGTGGCTGGACTACAAGCTGGACGACCAAGTGCTCATCGTGGACAACCCCCTGCGCAAGATCTACTTCAAGGGCGACAAGTACCTGTTCCTGTACACCGACAGCGAGATGGCCAATTACCGCTGGGGCTGCGTGGCCGAGGGCGAGGACGGCTACCTGGAGCAGATCCGCACCCACCTGGCGAGCGCCCTGGGCATCGTCCGCGAGCGCATCCCGCAGCCCCTGGCGCACGTGCACAAGTACTGGGCGCACGGCGTGGAGTTCGCCGACTCCGACATCGACCACCCGTCGGCGCTGTCCCACCGCGACAGCGGCATCATCGCCTGCAGCGACGCCTACACCGAGCACTGCGGCTGGATGGAGGGCGGCCTGCTGAGCGCGCGCGAGGCCAGCCGCCTGCTGCTGCAGCGCATCGCCGCC
3.4 Expression Explanation
After obtaining a codon-optimized DNA sequence, the next step is to understand how this sequence can actually produce the VioA protein inside a biological system.
In molecular biology this process follows the Central Dogma, which describes the flow of genetic information from DNA → RNA → Protein.
First, the DNA sequence is transcribed into messenger RNA (mRNA) by the host organism’s transcription machinery. This RNA molecule carries the genetic instructions encoded in the DNA sequence.
Next, the mRNA is translated by ribosomes. During translation, the ribosome reads the RNA sequence in groups of three nucleotides called codons. Each codon corresponds to a specific amino acid. These amino acids are then assembled sequentially to form the final protein structure.
In the context of this project, the codon-optimized VioA DNA sequence could be inserted into an expression vector and introduced into a host organism such as E. coli. Once inside the cell, the bacterial transcription and translation machinery would read the inserted DNA sequence and produce the VioA enzyme.
From my perspective as a designer working with biomaterials and color systems, this process can also be understood as a form of biological material programming. Instead of shaping materials directly, we design the genetic instructions that guide a living system to produce molecules with specific properties.
In the case of the violacein pathway, the VioA enzyme initiates the biochemical transformation of L-tryptophan, which ultimately leads to the production of the purple pigment violacein. If microorganisms capable of expressing this pathway grow on substrates such as coffee waste, the metabolic process could potentially convert organic waste into biologically produced color.
This perspective connects molecular biology with design research: DNA sequences function as programmable instructions that enable living systems to generate pigments and materials through metabolic processes.
Part 4 — Twist DNA Synthesis Practice
4.1 Creating Accounts
For this exercise I created accounts on both Benchling and Twist Bioscience, which are commonly used tools for designing and synthesizing DNA sequences.
Benchling was used to design and annotate the DNA sequence, while Twist provides services for synthesizing custom DNA constructs.
4.2 Building the DNA Insert Sequence
To express the VioA protein in a bacterial system, I designed a simple expression cassette. This cassette contains the genetic elements necessary for transcription and translation inside a host organism such as E. coli.
The structure of the insert sequence follows a common synthetic biology design:
Each part of the sequence has a specific function:
Promoter
Initiates transcription and determines how strongly the gene is expressed.
Ribosome Binding Site (RBS)
Allows ribosomes to recognize where translation should begin.
Start Codon (ATG)
Signals the start of protein translation.
Coding Sequence
The codon-optimized DNA sequence encoding the VioA protein.
His Tag
A short peptide tag that allows the produced protein to be purified using affinity chromatography.
Stop Codon
Signals the end of translation.
Terminator
Stops transcription and stabilizes the mRNA transcript.
From a design perspective, this expression cassette can be understood as a programmable biological instruction set. Instead of directly shaping a material, we design the genetic instructions that guide a living system to produce a molecule—in this case, the VioA enzyme involved in pigment biosynthesis.
In principle, if such a construct were introduced into a bacterial host capable of metabolizing substrates from coffee waste, the expressed enzyme could contribute to a metabolic pathway producing colored biomolecules such as violacein.
4.3 Selecting the “Genes” Option in Twist
After preparing the insert sequence in Benchling, I navigated to the Genes section in the Twist Bioscience platform. This interface allows users to submit DNA sequences for synthesis.
Since the goal of this exercise is to simulate ordering a gene construct, the Clonal Genes option was selected. Clonal genes are delivered in a circular plasmid vector, which allows the DNA to be directly transformed into a host organism such as E. coli.
4.4 Clonal Gene Selection
Clonal genes are commonly used in synthetic biology experiments because they simplify the experimental workflow. Once the plasmid is received, it can be directly introduced into bacterial cells for protein expression.
This makes clonal genes particularly useful when testing metabolic pathways or producing proteins such as enzymes involved in pigment biosynthesis.
4.5 Importing the DNA Sequence
Next, the designed DNA insert sequence was uploaded to the Twist interface using the Upload Sequence File option.
The uploaded sequence contains the complete expression cassette:
This sequence encodes the VioA enzyme, which catalyzes the first step in the violacein pigment biosynthesis pathway.
From a design perspective, this step represents translating a conceptual biological function into a programmable genetic instruction that can be synthesized and tested experimentally.
4.6 Choosing a Vector
After uploading the sequence, a cloning vector was selected from the Twist vector catalog.
For this exercise, a standard bacterial expression vector such as pTwist Amp High Copy can be used. This vector provides:
- an origin of replication allowing the plasmid to replicate inside E. coli
- an antibiotic resistance marker (ampicillin) for selection
- a stable backbone for the inserted gene construct
Once the construct is generated, the complete plasmid sequence can be downloaded as a GenBank file and re-imported into Benchling.
This plasmid represents the final synthetic construct containing the VioA gene expression system, which in principle could be used to express the enzyme in bacteria capable of metabolizing substrates such as coffee waste for pigment-related metabolic pathways.
Part 5 — DNA Read / Write / Edit
5.1 DNA Read
If I were to sequence DNA for my research, I would focus on microbial communities growing on organic waste substrates such as coffee waste.
Coffee waste contains a rich mixture of organic compounds that can support microbial growth. Sequencing the DNA of microorganisms that naturally colonize these environments could reveal bacteria capable of producing pigments or other useful biomolecules.
Understanding which microbial species are present and which metabolic pathways they contain would help identify organisms capable of transforming waste materials into valuable biological products such as color pigments.
To perform this sequencing, I would likely use Next Generation Sequencing (NGS) technologies such as Illumina sequencing.
Technology type:
Second-generation sequencing
Input preparation steps
- DNA extraction from microbial samples
- DNA fragmentation
- Adapter ligation
- PCR amplification
- Sequencing library preparation
How sequencing works
Illumina sequencing uses a method called sequencing by synthesis. DNA fragments are attached to a flow cell and amplified. Fluorescently labeled nucleotides are then incorporated during DNA synthesis, and the emitted fluorescence is detected to identify each base.
Output
The output is a large dataset of short DNA reads that can be assembled and analyzed to identify microbial species and genes involved in pigment biosynthesis.
5.2 DNA Write
If I were to synthesize DNA, I would design genetic constructs encoding pigment-producing metabolic pathways.
In particular, I am interested in pathways such as violacein biosynthesis, which converts the amino acid tryptophan into a purple pigment.
Designing such sequences could allow microorganisms to produce color directly from biological metabolism. This approach could potentially enable sustainable color production systems, where organic waste substrates serve as feedstock for microbial pigment synthesis.
Example design goal:
The DNA constructs for these enzymes could be synthesized using commercial DNA synthesis platforms such as Twist Bioscience.
Technology used
Modern DNA synthesis typically uses phosphoramidite chemical synthesis, where nucleotides are added sequentially to build a DNA strand.
Limitations
DNA synthesis technologies face challenges such as:
- sequence length limitations
- synthesis errors
- high cost for very large constructs
However, these technologies continue to improve and enable increasingly complex genetic designs.
5.3 DNA Edit
DNA editing technologies allow scientists to modify genes inside living organisms.
For research related to biological pigments and biomaterials, DNA editing could be used to modify microorganisms so that they produce new colors, materials, or metabolic products.
One widely used technology is CRISPR-Cas9.
CRISPR works by using a guide RNA (gRNA) to direct the Cas9 enzyme to a specific location in the genome. The Cas9 enzyme then creates a cut in the DNA, allowing scientists to insert, delete, or modify genetic sequences.
Editing workflow
- Design guide RNA targeting the gene of interest
- Introduce Cas9 and gRNA into the cell
- Cas9 cuts the DNA at the target location
- DNA repair mechanisms insert or modify the sequence
Applications
DNA editing could be used to:
- enhance pigment production
- introduce new biosynthetic pathways
- engineer microorganisms capable of converting waste into useful biomaterials
From a design perspective, these technologies transform DNA into a programmable material system. Instead of shaping materials directly, we design genetic instructions that allow living systems to generate color, structure, and material properties through biological processes.
Reflection
This week shifted my perspective on DNA. Before this exercise, I mostly understood DNA as a biological concept discussed in textbooks or laboratories. Through the assignments in this week, I started to see DNA as something closer to a design medium.
Coming from an interior architecture and design background, I am used to thinking about materials in terms of structure, transformation, and visual outcomes. Working with restriction digests, genetic sequences, and expression systems revealed that DNA can also function as a kind of instruction layer for material processes.
Instead of directly shaping matter, we design the genetic information that guides living systems to produce molecules, pigments, or structures.
For my own research, which explores biological color production from organic waste such as coffee grounds, this idea is particularly meaningful. The possibility that microorganisms could transform waste substrates into pigments suggests a different way of thinking about color production — one that is metabolic rather than industrial.
This week helped me see molecular biology not only as a scientific field, but also as a design space, where biological systems can be programmed to generate new materials, colors, and spatial possibilities.