Week 2 HW: DNA read and Write
Part 1 Benchling & In-Silico Gel Art
I simulated restriction enzyme digestion of Escherichia phage Lambda (complete genome) using the listed enzymes. Here are the results of the virtual digestion:
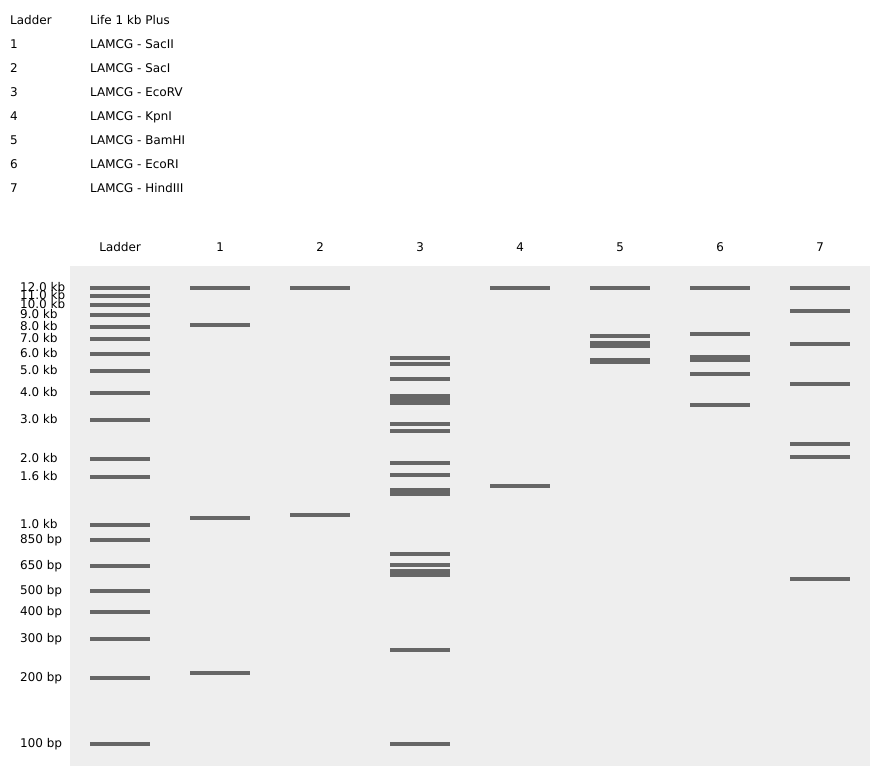

and I tried quick design using Ronan’s website:


Part 3 DNA Design Challenge
I choose Glutathione S-transferase Mu 1 (GSTM1). GSTM1 is a gene that encodes a Phase II detoxification enzyme, crucial for metabilizing xenobiotics, environmental toxins, and carcinogens by conjugating them with reduced glutathione. This gene located on chromosome 1p31 that protects cells from oxidative stress.
Reverse translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
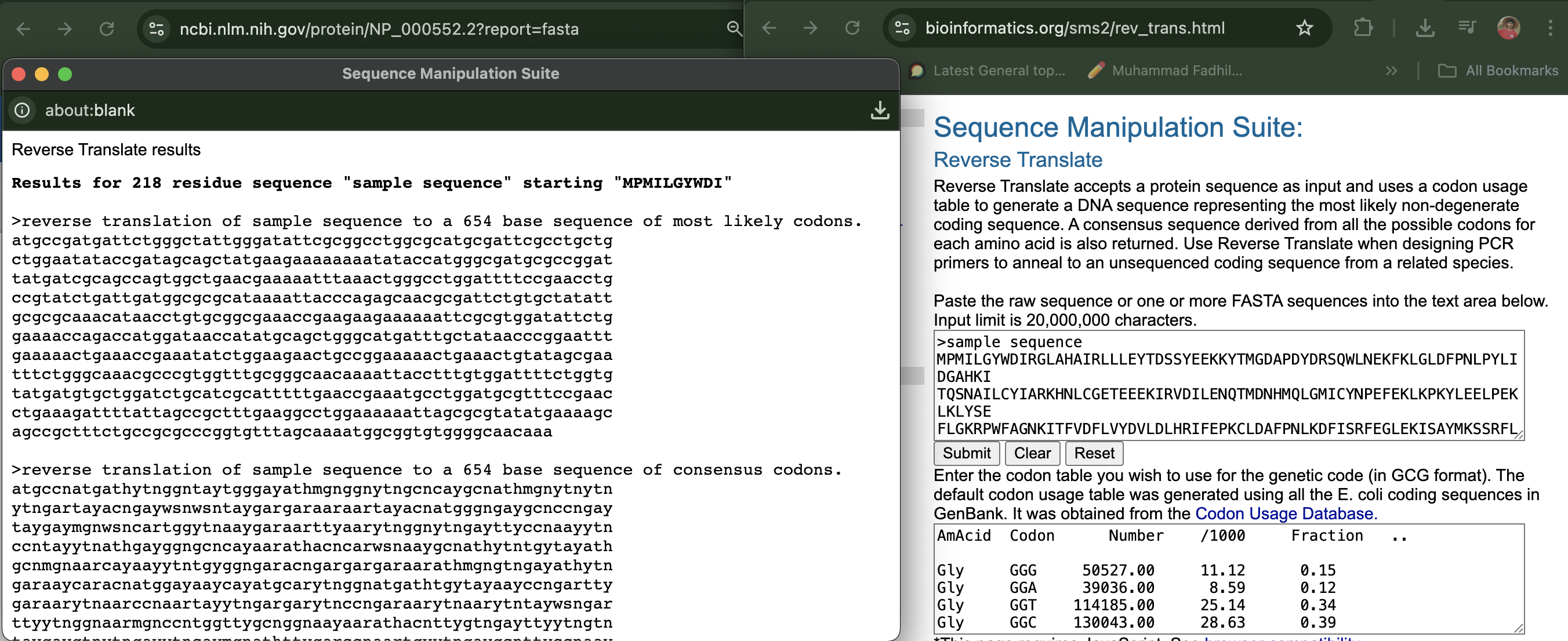

I use www.bioinformatics.org website to translate
Results for 218 residue sequence “sample sequence” starting “MPMILGYWDI”
reverse translation of sample sequence to a 654 base sequence of most likely codons. atgccgatgattctgggctattgggatattcgcggcctggcgcatgcgattcgcctgctg ctggaatataccgatagcagctatgaagaaaaaaaatataccatgggcgatgcgccggat tatgatcgcagccagtggctgaacgaaaaatttaaactgggcctggattttccgaacctg ccgtatctgattgatggcgcgcataaaattacccagagcaacgcgattctgtgctatatt gcgcgcaaacataacctgtgcggcgaaaccgaagaagaaaaaattcgcgtggatattctg gaaaaccagaccatggataaccatatgcagctgggcatgatttgctataacccggaattt gaaaaactgaaaccgaaatatctggaagaactgccggaaaaactgaaactgtatagcgaa tttctgggcaaacgcccgtggtttgcgggcaacaaaattacctttgtggattttctggtg tatgatgtgctggatctgcatcgcatttttgaaccgaaatgcctggatgcgtttccgaac ctgaaagattttattagccgctttgaaggcctggaaaaaattagcgcgtatatgaaaagc agccgctttctgccgcgcccggtgtttagcaaaatggcggtgtggggcaacaaa
reverse translation of sample sequence to a 654 base sequence of consensus codons. atgccnatgathytnggntaytgggayathmgnggnytngcncaygcnathmgnytnytn ytngartayacngaywsnwsntaygargaraaraartayacnatgggngaygcnccngay taygaymgnwsncartggytnaaygaraarttyaarytnggnytngayttyccnaayytn ccntayytnathgayggngcncayaarathacncarwsnaaygcnathytntgytayath gcnmgnaarcayaayytntgyggngaracngargargaraarathmgngtngayathytn garaaycaracnatggayaaycayatgcarytnggnatgathtgytayaayccngartty garaarytnaarccnaartayytngargarytnccngaraarytnaarytntaywsngar ttyytnggnaarmgnccntggttygcnggnaayaarathacnttygtngayttyytngtn taygaygtnytngayytncaymgnathttygarccnaartgyytngaygcnttyccnaay ytnaargayttyathwsnmgnttygarggnytngaraarathwsngcntayatgaarwsn wsnmgnttyytnccnmgnccngtnttywsnaaratggcngtntggggnaayaar
Codon optimization
Codon optimization is a way to ensure that the translation of our sequence’s codons is compatible with the host organism. If codon usage is incompatible with the host, protein expression will be inefficient. I try to optimized the GSTM1 coding sequence for expression in Escherichia coli, since it is widely used as a host organism.
Technology to produce this protein
I used cell-dependent system to optimized GSTM1 DNA. The gene is cloned into an expression plasmid using a T7 promoter, transformed into E.coli and induce the protein expression with IPTG (Isopropyl B-D-1-thiogalactopyranoside). The Recombinant GSTM1 protein would subsequently be purified using a His-tag through affinity chromatography.
How does it work in Biological System? the GSTM1 coding DNA sequence ATG CCT ATG ATC CTG GGC is transcribed into mRNA as AUG CCU AUG AUC CUG GGC, which is translated into the amino acid sequence Met–Pro–Met–Ile–Leu–Gly.
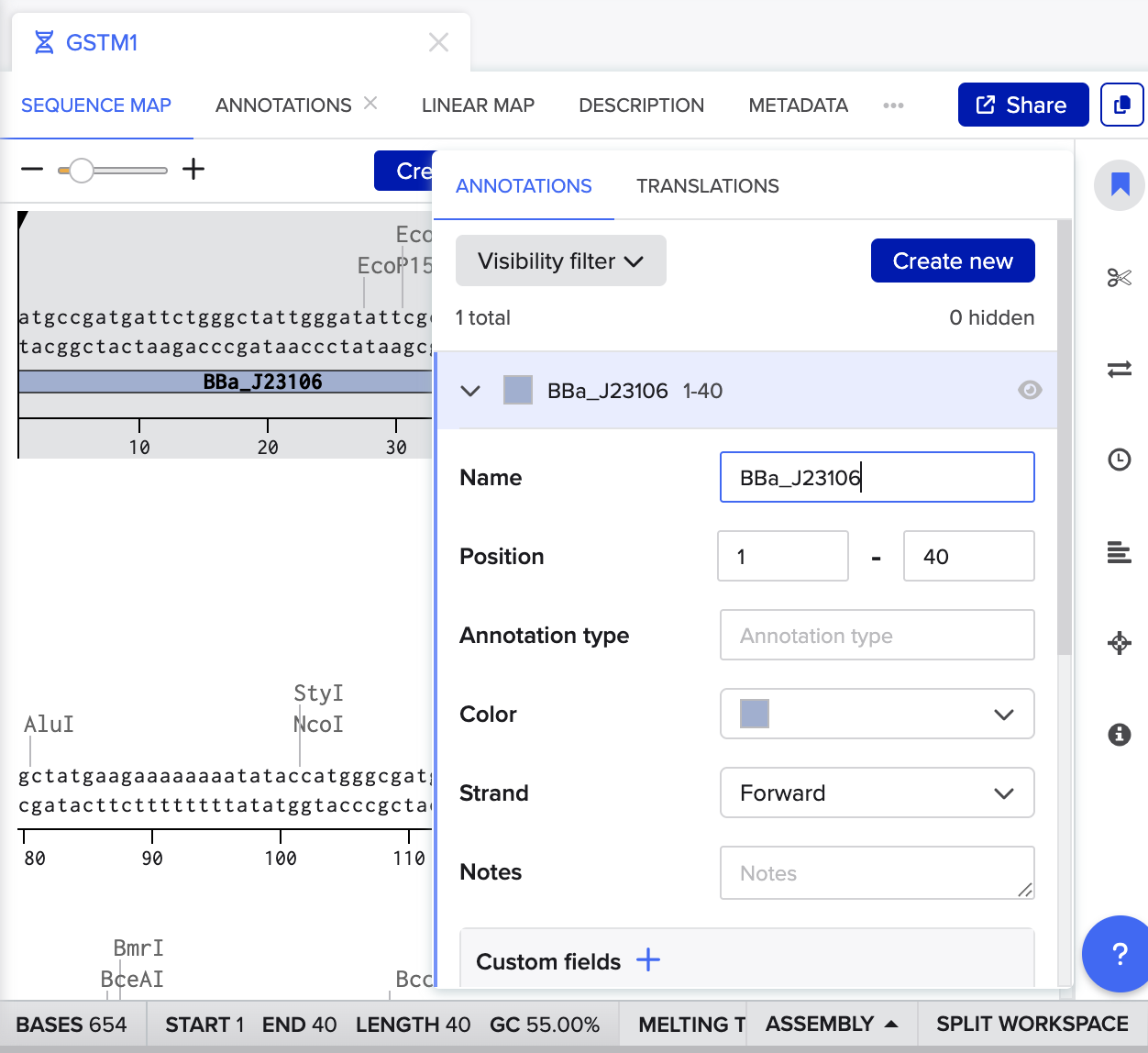
Part 4 Twist DNA Synthesis (Order?)
I have a problem following the istruction, after I put my bases and create sequence. I don’t understand how to make annotation

Part 5 DNA Read/Write/Edit
What DNA would you want to sequence (e.g., read) and why?
I would like to sequence the GSTM1 coding region and promoter region from individuals exposed to lead to assess potential mutations, deletions, or regulatory changes associated with heavy metal exposure. Because lead is known to induce oxidative stress and genomic instability, sequencing detoxification genes may reveal genetic susceptibility factors contributing to impaired xenobiotic elimination.
The method I would use is PCR amplification followed by –> Sanger sequencing to analyze GSTM1 gene (1st generation). The input would be genomic DNA extracted from blood samples of industrial workers exposed to lead. The GSTM1 gene region would be amplified using specific primers through PCR. The amplified product then undergo a sequencing reaction using fluorescently labeled ddNTPS.
During sequencing, incorporation of ddNTP terminates DNA synthesis. DNA fragments are separated by capillary electrophoresis. A laser detects fluorescent signals corresponding to each nucleotide, allowing base calling and reconstruction of the DNA sequence. The output consists of a chromatogram and a nucleotide sequence that can be compared with a reference sequence to identify mutations or variations.
- What DNA would you want to synthesize and why?
I would like to make Synthetic Detoxification Gene Construct for Heavy metal response by optimizing GSTM1 gene under a strong inducible promoter in E.coli. This construct would include a strong T7 promoter, ribosome binding site, the GSTM1 coding sequence, a 6xHis purification tag, and a transcription terminator. The objectives of synthesizing this DNA is to study detoxification mechanisms under heavy metal exposure, particularly lead-induced oxidative stress. Since GSTM1 catalyzes glutathione conjugation reactions, overexpression of this enzyme may help investigate protective mechanisms against xenobiotic toxicity. The synthesized coding sequence begins with ATG (start codon) and encodes the GSTM1 amino acid sequence, followed by a His-tag (CATCACCATCACCATCAC) for affinity purification. This construct could be used for protein expression studies or detoxification research
- What DNA would you want to edit and why?
I want to edit GSTM1 in human cells using CRISPR-Cas9 editing. The cell repairs the break either through non-homologous end joining (NHEJ) or homology-directed repair (HDR). For gene restoration, I would provide a donor DNA template to enable HDR-mediated insertion of a functional GSTM1 gene. The required inputs include guide RNA, Cas9 protein or plasmid, donor DNA template, and target cells. Although CRISPR is highly effective, limitations include off-target effects, low HDR efficiency, and delivery challenges in vivo.