Homework
Weekly homework submissions:
Week 1 HW: Principles and Practices
Project Proposal: Bio-IDE and Governance Architecture.
Week 1 HW: Principles and Practices
Project Proposal: Bio-IDE and Governance Architecture.
The Problem: Currently, designing a cloning experiment requires a fragmented, complex workflow. A researcher must search AddGene for vectors, use NEBcutter for enzyme selection, and switch to desktop software like SnapGene for visualization. This fragmentation leads to data compatibility errors and high barriers to entry for students, labbers, and hobbyists who cannot afford expensive software licenses.
The Solution: Bio-IDE is a browser-based “Co-Pilot” that unifies these tools into one intelligent workflow:
Why: By shifting from fragmented desktop tools to a unified web app, we democratize access to high-level biological design. A student in a resource-constrained lab can design industrial-grade experiments on a simple laptop.
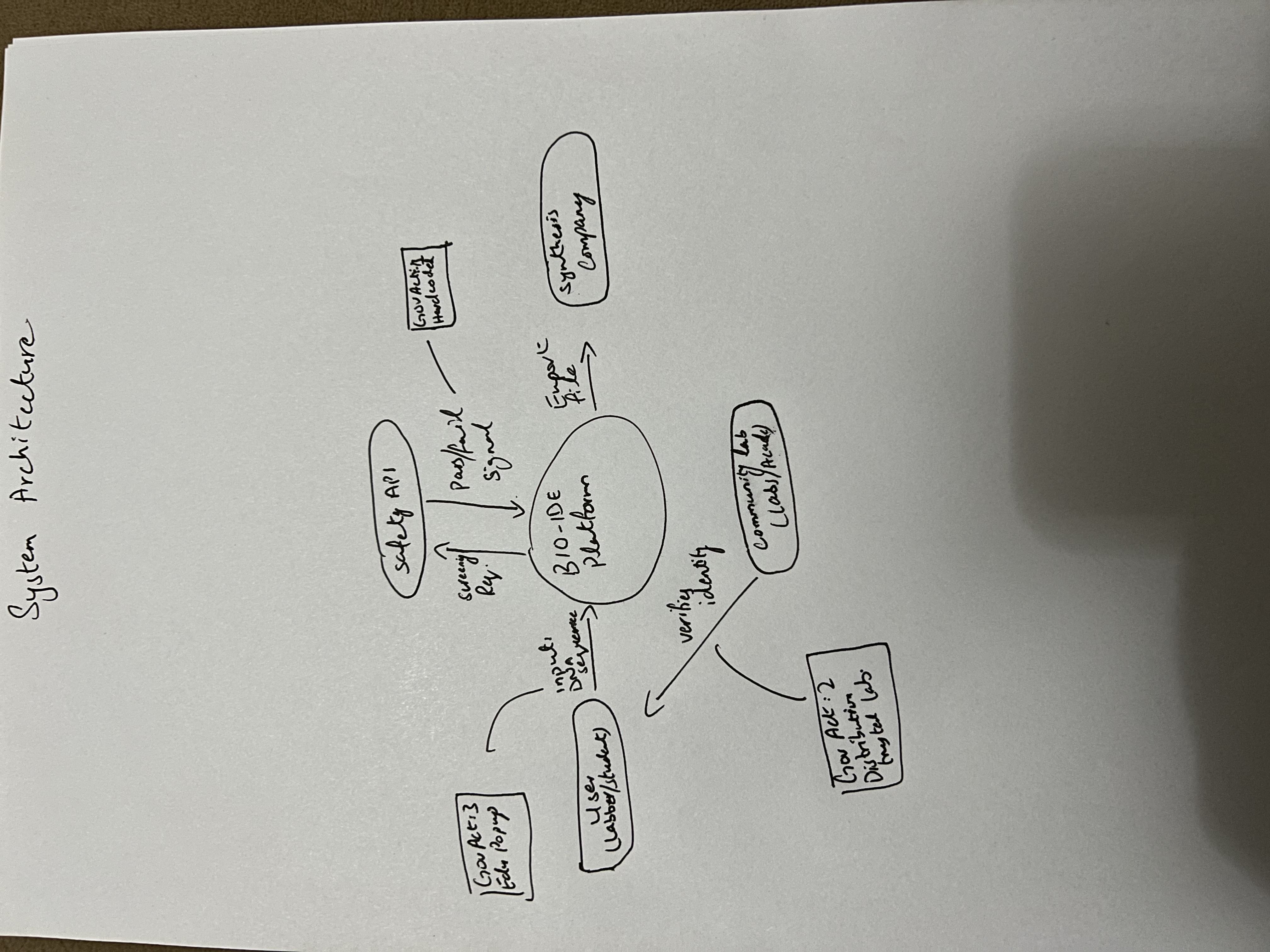
 Figure 1: Governance Ecosystem Sketch
Figure 1: Governance Ecosystem Sketch
To ensure Bio-IDE contributes to an ethical future, I propose the following goals:
Option 1: The “Embedded Safety” API (Hard Control)
Option 2: The “Community Verification” Web (Social Trust)
Option 3: The “Educational Pop-Up” (Informed Consent)
(Score Key: 1 = Best / Strongest Impact, 3 = Weakest Impact)
| Does the option: | Option 1 (Safety API) | Option 2 (Trust Web) | Option 3 (Edu-Popup) |
|---|---|---|---|
| Enhance Biosecurity | 1 | 2 | 3 |
| • By preventing incidents | 1 | 2 | 3 |
| • By helping respond | 1 | 1 | 2 |
| Foster Lab Safety | 1 | 3 | 1 |
| • By preventing incident | 1 | 3 | 1 |
| • By helping respond | n/a | 2 | 1 |
| Protect the environment | 2 | 3 | 1 |
| • By preventing incidents | 2 | 3 | 1 |
| • By helping respond | n/a | n/a | 1 |
| Other considerations | |||
| • Minimizing costs and burdens to stakeholders | 1 | 3 | 1 |
| • Feasibility? | 1 | 3 | 1 |
| • Not impede research | 2 | 1 | 2 |
| • Promote constructive applications | 1 | 1 | 1 |
I prioritize Option 1 (Embedded Safety) supported by Option 3 (Educational Quiz).
Reasoning: As a solo developer, I need scalable solutions. Option 1 provides the “Hard Wall” against bioweapons, while Option 3 provides the “Soft Skills” to ensure students handle even safe organisms correctly. Both can be implemented in code without requiring a massive human support team (unlike Option 2, which requires managing a complex human trust network).
New Concern: The “Black Box” Problem Reflecting on this week’s lectures, I worry that automating the design process might lead to “Scientific Deskilling.” If the AI chooses the plasmid and the enzymes, the student might never learn why those choices were made.
Proposed Governance: To counter this, Bio-IDE will implement an “Explainable AI” (XAI) interface. It will not just say “Use pUC19.” It will say: “I recommend pUC19 because your gene is small (500bp) and you requested high copy number for maximum yield.” This turns the tool into a mentor, not just a machine.

 Figure 2: My calculations for Polymerase error rates and Oligo synthesis yields.
Figure 2: My calculations for Polymerase error rates and Oligo synthesis yields.
How biology deals with the error discrepancy: As calculated above, 3,200 errors per division is too high. Biology solves this with a Mismatch Repair (MMR) system (MutS/MutL). This system scans the DNA after the polymerase passes, finding “bumps” (mismatches) and cutting them out. This improves accuracy by another 1000x, bringing the final error rate to ~10^-9 (less than 1 error per genome).
Why coding for proteins is difficult (Secondary Structure): Even if a DNA sequence theoretically codes for the right amino acids, it might fail in practice due to Secondary Structure. If the mRNA folds into a tight hairpin loop (as seen in NUPACK simulations), the ribosome cannot bind to the “Start” signal, and translation never happens.
Most Common Synthesis Method: The industry standard is Phosphoramidite Chemistry. It is a 4-step cycle (De-blocking –> Coupling –> Capping –> Oxidation) performed on a solid support (like a silicon chip or glass beads).
Why we can’t print 2000bp genes: As shown in my calculations (Figure 2), the yield for a 2000bp sequence drops to 0.004%. This means the test tube would contain almost entirely “trash” (truncated fragments). To build a 2000bp gene, we must synthesize short oligos (e.g., 60-100nt) and stitch them together using enzymes (Gene Assembly).
Question: What are the 10 essential amino acids, and how does this affect the “Lysine Contingency”?