Week 2 HW: DNA - READ, WRITE & EDIT

Homework
Part 0: Basics of Gel Electrophoresis
I have watched all the lecture slides and reciatation videos.
Part 1: Benchling & In-silico Gel Art
I created a benchling account and imported the Lambda DNA
Restriction Enzyme Digest Simulations
EcoRI Restriction Enzyme Digest Simulation.
HindIII Restriction Enzyme Digest Simlution.
BamHI Restriction Enzyme Digest Simulation.
KpnI Restriction Enzyme Digest Simulation.
EcoRV Restriction Enzyme Digest Simulation.
SacI Restriction Enzyme Digest Simulation.
SalI Restriction Enzyme Digest Simulation.
EcoRI, HindIII, BamHI, KpnI, EcoRV, SacI, SalI Simultanious Restriction Enzyme Digest.
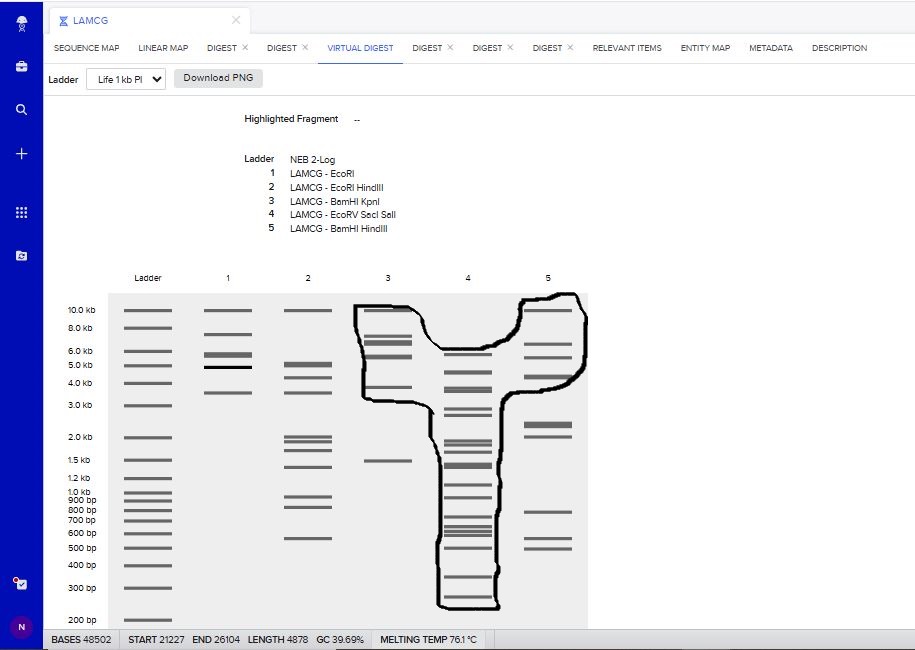
I tried to create a design in Benchling, after many trials and errors, I managed to make a pattern by using double and triple digests of restriction enzymes.
I think it looks like a Y.

Part 3: DNA Design Challenge
3.1. Choose your protein.
In recitation, we discussed that you will pick a protein for your homework that you find interesting. Which protein have you chosen and why? Using one of the tools described in the recitation (NCBI, UniProt, Google), obtain the protein sequence for the protein you chose.
Answer
I picked the BMP1 protein (Bone morphogenetic protein 1). It is a secreted metalloprotease encoded by the BMP1 gene in humans. It belongs to the astacin M12A family of proteases and plays a central role in extracellular matrix assembly by cleaving precursor proteins into the mature functional forms. Growing up, I never had a bone fracture or dislocation, but my brother had a fracture in his left hand, which made me curious about the proteins and genes that drive bone formation. (https://www.uniprot.org/uniprotkb/P13497/entry)
The protein sequence for BMP1 protein on Uniprot is an isoform that has been chosen as the canonical sequence. The sequence is as follows: MPGVARLPLLLGLLLLPRPGRPLDLADYTYDLAEEDDSEPLNYKDPCKAAAFLGDIALDEEDLRAFQVQQAVDLRRHTARKSSIKAAVPGNTSTPSCQSTNGQPQRGACGRWRGRSRSRRAATSRPERVWPDGVIPFVIGGNFTGSQRAVFRQAMRHWEKHTCVTFLERTDEDSYIVFTYRPCGCCSYVGRRGGGPQAISIGKNCDKFGIVVHELGHVVGFWHEHTRPDRDRHVSIVRENIQPGQEYNFLKMEPQEVESLGETYDFDSIMHYARNTFSRGIFLDTIVPKYEVNGVKPPIGQRTRLSKGDIAQARKLYKCPACGETLQDSTGNFSSPEYPNGYSAHMHCVWRISVTPGEKIILNFTSLDLYRSRLCWYDYVEVRDGFWRKAPLRGRFCGSKLPEPIVSTDSRLWVEFRSSSNWVGKGFFAVYEAICGGDVKKDYGHIQSPNYPDDYRPSKVCIWRIQVSEGFHVGLTFQSFEIERHDSCAYDYLEVRDGHSESSTLIGRYCGYEKPDDIKSTSSRLWLKFVSDGSINKAGFAVNFFKEVDECSRPNRGGCEQRCLNTLGSYKCSCDPGYELAPDKRRCEAACGGFLTKLNGSITSPGWPKEYPPNKNCIWQLVAPTQYRISLQFDFFETEGNDVCKYDFVEVRSGLTADSKLHGKFCGSEKPEVITSQYNNMRVEFKSDNTVSKKGFKAHFFSDKDECSKDNGGCQQDCVNTFGSYECQCRSGFVLHDNKHDCKEAGCDHKVTSTSGTITSPNWPDKYPSKKECTWAISSTPGHRVKLTFMEMDIESQPECAYDHLEVFDGRDAKAPVLGRFCGSKKPEPVLATGSRMFLRFYSDNSVQRKGFQASHATECGGQVRADVKTKDLYSHAQFGDNNYPGGVDCEWVIVAEEGYGVELVFQTFEVEEETDCGYDYMELFDGYDSTAPRLGRYCGSGPPEEVYSAGDSVLVKFHSDDTITKKGFHLRYTSTKFQDTLHSRK
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
The Central Dogma discussed in class and recitation describes the process in which DNA sequence becomes transcribed and translated into protein. The Central Dogma gives us the framework to work backwards from a given protein sequence and infer the DNA sequence that the protein is derived from. Using one of the tools discussed in class, NCBI or online tools (google “reverse translation tools”), determine the nucleotide sequence that corresponds to the protein sequence you chose above.
Answer
I used https://www.bioinformatics.org/sms2/rev_trans.html to reverse transcribe the protein to its nucleotide sequence.
The nucleotide sequence for the Bone morphogenetic protein 1 is as follows:
atgccgggcgtggcgcgcctgccgctgctgctgggcctgctgctgctgccgcgcccgggc cgcccgctggatctggcggattatacctatgatctggcggaagaagatgatagcgaaccg ctgaactataaagatccgtgcaaagcggcggcgtttctgggcgatattgcgctggatgaa gaagatctgcgcgcgtttcaggtgcagcaggcggtggatctgcgccgccataccgcgcgc aaaagcagcattaaagcggcggtgccgggcaacaccagcaccccgagctgccagagcacc aacggccagccgcagcgcggcgcgtgcggccgctggcgcggccgcagccgcagccgccgc gcggcgaccagccgcccggaacgcgtgtggccggatggcgtgattccgtttgtgattggc ggcaactttaccggcagccagcgcgcggtgtttcgccaggcgatgcgccattgggaaaaa catacctgcgtgacctttctggaacgcaccgatgaagatagctatattgtgtttacctat cgcccgtgcggctgctgcagctatgtgggccgccgcggcggcggcccgcaggcgattagc attggcaaaaactgcgataaatttggcattgtggtgcatgaactgggccatgtggtgggc ttttggcatgaacatacccgcccggatcgcgatcgccatgtgagcattgtgcgcgaaaac attcagccgggccaggaatataactttctgaaaatggaaccgcaggaagtggaaagcctg ggcgaaacctatgattttgatagcattatgcattatgcgcgcaacacctttagccgcggc atttttctggataccattgtgccgaaatatgaagtgaacggcgtgaaaccgccgattggc cagcgcacccgcctgagcaaaggcgatattgcgcaggcgcgcaaactgtataaatgcccg gcgtgcggcgaaaccctgcaggatagcaccggcaactttagcagcccggaatatccgaac ggctatagcgcgcatatgcattgcgtgtggcgcattagcgtgaccccgggcgaaaaaatt attctgaactttaccagcctggatctgtatcgcagccgcctgtgctggtatgattatgtg gaagtgcgcgatggcttttggcgcaaagcgccgctgcgcggccgcttttgcggcagcaaa ctgccggaaccgattgtgagcaccgatagccgcctgtgggtggaatttcgcagcagcagc aactgggtgggcaaaggcttttttgcggtgtatgaagcgatttgcggcggcgatgtgaaa aaagattatggccatattcagagcccgaactatccggatgattatcgcccgagcaaagtg tgcatttggcgcattcaggtgagcgaaggctttcatgtgggcctgacctttcagagcttt gaaattgaacgccatgatagctgcgcgtatgattatctggaagtgcgcgatggccatagc gaaagcagcaccctgattggccgctattgcggctatgaaaaaccggatgatattaaaagc accagcagccgcctgtggctgaaatttgtgagcgatggcagcattaacaaagcgggcttt gcggtgaacttttttaaagaagtggatgaatgcagccgcccgaaccgcggcggctgcgaa cagcgctgcctgaacaccctgggcagctataaatgcagctgcgatccgggctatgaactg gcgccggataaacgccgctgcgaagcggcgtgcggcggctttctgaccaaactgaacggc agcattaccagcccgggctggccgaaagaatatccgccgaacaaaaactgcatttggcag ctggtggcgccgacccagtatcgcattagcctgcagtttgatttttttgaaaccgaaggc aacgatgtgtgcaaatatgattttgtggaagtgcgcagcggcctgaccgcggatagcaaa ctgcatggcaaattttgcggcagcgaaaaaccggaagtgattaccagccagtataacaac atgcgcgtggaatttaaaagcgataacaccgtgagcaaaaaaggctttaaagcgcatttt tttagcgataaagatgaatgcagcaaagataacggcggctgccagcaggattgcgtgaac acctttggcagctatgaatgccagtgccgcagcggctttgtgctgcatgataacaaacat gattgcaaagaagcgggctgcgatcataaagtgaccagcaccagcggcaccattaccagc ccgaactggccggataaatatccgagcaaaaaagaatgcacctgggcgattagcagcacc ccgggccatcgcgtgaaactgacctttatggaaatggatattgaaagccagccggaatgc gcgtatgatcatctggaagtgtttgatggccgcgatgcgaaagcgccggtgctgggccgc ttttgcggcagcaaaaaaccggaaccggtgctggcgaccggcagccgcatgtttctgcgc ttttatagcgataacagcgtgcagcgcaaaggctttcaggcgagccatgcgaccgaatgc ggcggccaggtgcgcgcggatgtgaaaaccaaagatctgtatagccatgcgcagtttggc gataacaactatccgggcggcgtggattgcgaatgggtgattgtggcggaagaaggctat ggcgtggaactggtgtttcagacctttgaagtggaagaagaaaccgattgcggctatgat tatatggaactgtttgatggctatgatagcaccgcgccgcgcctgggccgctattgcggc agcggcccgccggaagaagtgtatagcgcgggcgatagcgtgctggtgaaatttcatagc gatgataccattaccaaaaaaggctttcatctgcgctataccagcaccaaatttcaggat accctgcatagccgcaaa
3.3. Codon optimization.
Once a nucleotide sequence of your protein is determined, you need to codon optimize your sequence. You may, once again, utilize Google for a “codon optimization tool”. In your own words, describe why you need to optimize codon usage. Which organism have you chosen to optimize the codon sequence for and why?
[Example from Codon Optimization Tool | Twist Bioscience while avoiding Type IIs enzyme recognition sites BsaI, BsmBI, and BbsI]
Answer
Codons need to be optimized for use due to the codon usage bias in the heterologous host organisms. The codon usage bias is due to variations in tRNA abundance in different organisms, which directly impacts translation speed and accuracy. When a gene from one organism is expressed in another, such as a human gene in bacteria, the mismatch in codon preference can cause ribosomes to stall at rare codons, leading to reduced protein yield, truncated proteins, or misfolding. Thus, codons are optimized to ensure the efficient expression of proteins in heterologous host organisms.
I optimized the Bone morphogenetic protein 1 for insertion into an E.coli plasmid using the VectorBuilder DNA Optimization tool (https://en.vectorbuilder.com/tool/codon-optimization.html).
The optimized Bone morphogenetic protein 1 nucleotide sequence for insertion into an Escherichia coli str. K-12 substr. MG1655 is as follows:
ATGCCGGGTGTTGCGCGCCTGCCGCTGCTGCTGGGCCTGCTGCTGCTGCCGCGTCCGGGCCGCCCGCTGGATCTGGCGGACTATACCTATGATCTGGCCGAAGAAGATGATAGCGAACCGCTGAACTATAAAGATCCGTGCAAAGCCGCGGCGTTTCTGGGCGATATTGCGCTGGATGAAGAAGATCTGCGCGCGTTCCAGGTGCAGCAGGCCGTGGATCTGCGCCGCCATACCGCGCGTAAAAGCAGCATTAAAGCGGCGGTCCCGGGCAACACCTCGACCCCGAGCTGCCAGAGCACCAATGGCCAGCCGCAGCGCGGTGCCTGCGGCCGCTGGCGCGGCCGCTCACGTAGCCGTCGTGCGGCCACCAGCCGCCCGGAACGTGTGTGGCCGGATGGCGTCATCCCGTTCGTGATTGGCGGCAATTTCACCGGCAGCCAGCGTGCCGTATTTCGCCAGGCGATGCGCCATTGGGAAAAACATACATGCGTGACCTTCCTGGAACGCACCGATGAAGATAGCTACATTGTGTTTACCTATCGCCCGTGCGGCTGCTGCAGCTATGTGGGCCGCCGTGGCGGCGGCCCGCAGGCGATTAGCATTGGCAAAAATTGCGACAAATTTGGTATTGTGGTGCATGAACTGGGCCATGTGGTGGGCTTTTGGCATGAACATACCCGCCCGGATCGTGATCGCCATGTTAGCATTGTGCGCGAAAACATTCAGCCGGGCCAGGAATATAATTTTCTGAAAATGGAGCCGCAGGAAGTGGAAAGCCTGGGCGAAACCTATGATTTCGATAGCATTATGCACTATGCGCGTAACACCTTCAGCCGCGGCATTTTCCTGGATACCATTGTACCGAAATACGAAGTCAATGGCGTAAAACCGCCGATTGGCCAGCGCACCCGCCTGAGCAAAGGAGATATTGCGCAGGCGCGTAAACTGTATAAATGCCCGGCGTGCGGCGAAACCCTGCAAGATAGCACCGGTAACTTCAGCAGCCCGGAATATCCGAATGGATATAGCGCCCATATGCACTGCGTGTGGCGCATTTCAGTTACCCCGGGCGAAAAAATTATTCTGAACTTTACCTCGCTGGATCTGTATCGCAGCCGCCTGTGCTGGTACGATTACGTTGAAGTGCGTGATGGCTTTTGGCGCAAAGCGCCGCTGCGCGGCCGCTTCTGTGGCAGCAAACTGCCGGAACCGATTGTCAGCACAGATAGCCGTCTGTGGGTGGAATTTCGCAGCTCAAGCAATTGGGTGGGCAAAGGCTTTTTCGCGGTATATGAAGCCATTTGCGGCGGTGATGTGAAAAAAGATTACGGCCACATTCAGAGCCCGAACTATCCGGATGATTACCGCCCGAGCAAAGTATGCATCTGGCGCATTCAGGTGAGCGAAGGTTTTCACGTGGGCCTGACCTTTCAGTCATTTGAAATTGAGCGCCATGATAGCTGTGCGTACGATTATCTGGAAGTGCGTGATGGTCATAGCGAAAGCTCAACCCTGATTGGCCGCTACTGCGGCTACGAAAAACCGGATGATATTAAAAGCACCAGCAGCCGTCTGTGGCTGAAATTTGTGAGCGATGGCAGCATTAACAAAGCGGGCTTCGCGGTTAATTTCTTCAAAGAAGTGGATGAATGCTCGCGCCCGAATCGCGGCGGCTGCGAACAGCGCTGTCTGAATACCCTGGGCAGCTATAAATGCAGCTGCGATCCGGGTTATGAACTGGCGCCGGATAAACGTCGCTGTGAAGCGGCGTGCGGCGGTTTTCTGACCAAACTGAATGGTAGCATTACGAGCCCGGGTTGGCCGAAAGAATATCCGCCGAACAAAAATTGCATCTGGCAGCTGGTGGCGCCGACCCAGTACCGCATTAGCCTGCAGTTTGATTTCTTTGAAACCGAAGGCAATGACGTCTGTAAATATGACTTCGTGGAAGTCCGCAGCGGCCTGACCGCGGATAGTAAACTGCACGGCAAATTCTGCGGCAGCGAAAAACCGGAAGTGATCACCAGCCAGTACAATAACATGCGCGTGGAATTCAAAAGCGACAACACCGTGAGCAAAAAAGGCTTTAAAGCACATTTTTTTAGCGATAAAGATGAATGTAGTAAAGATAACGGTGGCTGTCAGCAGGATTGCGTTAACACCTTTGGCAGCTACGAATGCCAGTGCCGCAGCGGTTTTGTGCTGCACGATAACAAACATGATTGTAAAGAAGCGGGTTGCGATCATAAAGTGACCAGCACCTCAGGCACCATTACCAGCCCGAACTGGCCGGATAAATATCCGAGCAAAAAAGAATGCACCTGGGCGATTAGCAGCACCCCGGGCCATCGTGTGAAACTGACCTTTATGGAAATGGATATTGAAAGCCAGCCGGAATGTGCGTACGACCATCTGGAAGTGTTTGATGGTCGCGATGCCAAAGCGCCGGTTCTGGGCCGTTTTTGCGGCAGCAAAAAACCGGAACCGGTCCTGGCGACGGGCAGCCGCATGTTCCTGCGGTTCTACAGCGATAACAGCGTGCAGCGTAAAGGTTTTCAGGCCAGCCATGCGACCGAATGCGGTGGCCAGGTGCGTGCCGATGTGAAAACCAAAGATCTGTACAGCCATGCGCAGTTTGGCGATAACAACTATCCGGGCGGCGTGGATTGCGAATGGGTGATCGTGGCGGAAGAAGGCTATGGCGTGGAACTTGTGTTTCAGACCTTTGAAGTGGAAGAAGAAACCGATTGTGGTTACGACTACATGGAACTGTTCGATGGCTATGACAGCACCGCCCCGCGCCTGGGCCGTTATTGCGGCAGCGGCCCGCCGGAAGAAGTGTACTCCGCCGGCGATAGCGTTCTGGTGAAGTTTCATAGCGATGATACCATTACCAAGAAAGGCTTTCACCTGCGCTATACCAGCACCAAATTTCAGGATACCCTGCACAGCCGCAAA
3.4. You have a sequence! Now what?
What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both.
Answer
In a cell-dependent system, the Bone morphogenetic protein 1 can be produced using recombinant plasmid cloning technology. This would work by inserting the DNA sequence coding for the BMP1 protein into a plasmid such as E.coli. The DNA sequence should be optimised for the chosen plasmid. The plasmid should have a promoter, start and stop codons, regulator sequences, and a terminator. The plasmid is then introduced into bacteria via transformation. Inside the bacteria, RNA polymerase will bind to the promoter and transcribe the DNA coding region in RNA. Which then binds to the ribosome and tRNA reads the codons and assembles amino acids and peptide chains fold into the BMP1 protein.
Part 4: Prepare a Twist DNA Synthesis Order
4.1 Create a twist account I created a twist account
I chose to build an insert sequence for the luciferase. The luciferase gene encodes an enzyme that catalyzes a bioluminescent reaction, producing light in the presence of its substrate (luciferin), ATP, and oxygen. It is widely used as a reporter gene to study gene expression.
I first started the build by using the T7 promoter and added the Shine-Dalgarno sequence as the ribosome binding site.
I added a start codon
I imported the luciferase gene from NCBI and tried to copy out the coding sequence for luciferase. I used Benchling to optimize the coding sequence for insertion into E.coli plasmid.
I then inserted the optimized luciferase coding sequence into the build, added a 6x his tag,a stop codon, and a T7 terminator
The structure of the insert was as follows:
- T7 Promoter
- RBS
- Start Codon
- Luciferase Coding sequence
- 6x His Tag
- Stop Codon
- T7 Terminator
Benchling link: https://benchling.com/s/seq-udkiPsJ6nw8LkBmPBwlf?m=slm-X0Qj5SONejIma2zXxUQ8
I uploaded the sequence into Twist and chose the pTwist Amp High Copy vector. I then downloaded the construct as GenBack and imported it into Benchling.
Benchling link: https://benchling.com/s/seq-kstnI8Jy48TISQPIJUoh?m=slm-d5lB8uOEDoqk2WyY47fS
Part 5: DNA Read/Write/Edit
(i) What DNA would you want to sequence (e.g., read) and why?
Answer
I would like to sequence the whole genome and transcriptome of rice varieties grown in Ghana. I would focus on genes involved in nitrogen use efficiency, drought tolerance, and yield stability. Coming from an agricultural biotechnology background, sequencing rice DNA for nitrogen use efficiency will improve food security by reducing fertilizer dependency while maintaining yield levels. Additionally, nitrogen metabolism is tightly linked to drought stress and carbon metabolism, thus sequencing can reveal alleles that enhance resilience under variable rainfall.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
Answer
I would use a hybrid sequencing strategy combining third-generation PacBio HiFi sequencing (developed by Pacific Biosciences) and second-generation Illumina sequencing (developed by Illumina) because each technology addresses different challenges of plant genomics. The genome of Oryza sativa contains many repetitive elements and duplicated gene families, which are difficult to assemble accurately using short reads alone. PacBio HiFi produces long, highly accurate reads that can span repetitive regions and resolve structural variants such as insertions, deletions, and gene duplications. This is especially important when studying nitrogen use efficiency genes, which may exist in multiple similar copies or be influenced by regulatory structural variation. Long-read sequencing therefore enables the creation of a high-quality de novo assembly of locally adapted rice varieties, ensuring that important region-specific alleles are not missed.
However, PacBio alone is not sufficient for large-scale comparative or expression studies. Illumina sequencing provides extremely high depth at a lower cost per base, making it ideal for population-level SNP discovery, genome polishing, and RNA sequencing. Since nitrogen use efficiency is strongly influenced by gene regulation, Illumina RNA-seq would allow precise quantification of gene expression under different nitrogen treatments. Combining long-read structural resolution with high-depth short-read accuracy ensures reliable variant detection, strong transcriptomic analysis, and cost efficiency. Together, this hybrid approach provides the comprehensive genomic insight needed to improve nitrogen use efficiency, enhance sustainable fertilizer management, and support precision breeding strategies in rice.
For PacBio HiFi sequencing, the input would be high-molecular-weight genomic DNA (15-25 kb fragments).
Library Preparation:
- Extract intact genomic DNA.
- Size selection.
- Ligate hairpin adapters to create circular SMRTbell templates.
- Polymerase binding (no amplification required).
Sequencing & Base Calling:
- DNA polymerase is immobilized in Zero-Mode Waveguides (ZMWs).
- Fluorescently labeled nucleotides are incorporated.
- Each base emits a distinct fluorescence signal.
- Circular consensus sequencing (multiple passes) improves accuracy to >99.9% (Wenger et al., 2019).
Output:
- Long high-fidelity reads (10–25 kb)
- FASTQ files with quality scores
- Ideal for resolving repetitive plant genome regions
- Long-read sequencing is especially important because plant genomes are repeat-rich and structurally complex (Michael & VanBuren, 2020).
For Illumina Sequencing, the input would be fragmented DNA of 300-500 bp or cDNA for RNA sequencing.
Library Preparation:
- DNA fragmentation
- End repair and adapter ligation
- PCR amplification
- Cluster generation via bridge amplification
Sequencing & Base Calling: -Sequencing-by-synthesis
- Reversible terminator nucleotides incorporated one at a time
- Fluorescent imaging determines base identity
- High per-base accuracy (>99%)
Output: Millions to billions of short reads (100–150 bp), which are ideal for gene expression quantification and polishing assemblies
5.2 DNA Write
I would design and synthesize a synthetic nitrogen-sensing genetic circuit that could be introduced into rice to improve nitrogen uptake and fertilizer responsiveness.
Instead of simply overexpressing a transporter gene (which can cause metabolic imbalance), I would engineer a smart, feedback-controlled genetic circuit that activates nitrogen uptake genes only under low-nitrogen conditions. The genetic circuit would consist of:
- A low-nitrogen inducible promoter.
- A synthetic transcriptional activator module.
- A nitrogen transporter gene.
- A fluorescent reporter for monitoring.
- A terminator sequence.
The core gene would be NRT2.1 (high-affinity nitrate transporter, which is involved in nitrate uptake under nitrogen-limited conditions.
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
Answer
To synthesize the nitrogen-responsive genetic circuit containing the NRT2.1 module, I would use phosphoramidite-based solid-phase DNA synthesis combined with enzymatic assembly methods such as Gibson Assembly.
In this approach, short DNA oligonucleotides are chemically synthesized base-by-base through iterative cycles of deprotection, nucleotide coupling, capping, and oxidation. Because individual oligos are typically limited to ~200 bp, overlapping fragments would then be assembled into the full-length construct using enzymatic assembly in a single reaction using Gibson assembly.
This method is precise, scalable, and well-suited for modular plant genetic circuit design. However, limitations include length constraints requiring multi-fragment assembly, potential synthesis errors that accumulate with longer sequences, and challenges with high-GC or repetitive regions. Therefore, the final construct would require sequence verification to ensure accuracy before plant transformation.
5.3 DNA Edit
(i) What DNA would you want to edit and why?
Beyond plants, I would be very interested in editing the genome of Aedes aegypti, the mosquito species that transmits diseases such as dengue, Zika, and yellow fever. The goal would be to reduce the transmission of these viruses through gene drives or other targeted genome editing strategies. Specifically, I would target genes involved in fertility or pathogen susceptibility, such as those encoding reproductive proteins or viral receptor proteins in the mosquito midgut. For example, disrupting a key fertility gene could reduce mosquito population density, while modifying viral receptor genes could make mosquitoes resistant to virus infection, breaking the disease transmission cycle.
My rationale for editing Aedes aegypti is both public health and environmental impact. Vector-borne diseases affect millions worldwide, especially in tropical regions, and current control methods (insecticides, habitat elimination) are often insufficient, costly, or ecologically harmful. Gene editing offers a precise, sustainable solution that can complement traditional control strategies.
(ii) What technology or technologies would you use to perform these DNA edits and why?
For this, I would use CRISPR-Cas9-based gene drives, which allow a targeted gene to be copied preferentially to offspring, ensuring rapid spread of the desired trait through wild populations. CRISPR-Cas9, which is currently the most precise and widely used genome editing technology for insects and other organisms. CRISPR-Cas9 enables targeted modifications of DNA by creating double-strand breaks at specific genomic locations, which are then repaired by the cell’s own repair machinery, allowing for insertions, deletions, or gene replacement. This technology is ideal for engineering traits such as reduced fertility or virus resistance in mosquitoes.
However, careful containment, ecological risk assessment, and ethical considerations would be critical because of the potential for irreversible effects in wild populations.
How CRISPR-Cas9 Edits DNA
- Targeting – A single-guide RNA is designed to complement a specific DNA sequence in the mosquito genome, adjacent to a protospacer adjacent motif.
- Cutting – Cas9 endonuclease binds the sgRNA and introduces a double-strand break at the targeted site.
- Repair – The mosquito cell repairs the break via:
- Non-Homologous End Joining → introduces small insertions/deletions (indels), which can disrupt gene function.
- Homology-Directed Repair → if a DNA template is provided, precise sequence changes can be introduced, e.g., inserting a virus-resistance allele.
Preparation and Inputs
Before editing, careful design and preparation are required:
Design steps
- Identify the target genes critical for fertility or viral susceptibility.
- Design sgRNAs that minimize off-target effects using computational tools.
- Design donor DNA templates if precise sequence insertion is needed.
Inputs
- sgRNA – synthesized guide RNA targeting the mosquito gene.
- Cas9 protein or Cas9-expressing plasmid/mRNA.
- Donor DNA template.
- Embryos or cultured mosquito cells – Aedes aegypti embryos are typically microinjected with these components.
The delivery will be a microinjection of CRISPR components into fertilized mosquito eggs, which is standard for germline editing, ensuring heritable changes.
CRISPR-Cas9 has some limitations, which are editing efficiency can be low because not all injected embryos survive, and only a fraction carries the intended mutation.
The precision of the edits is also variable: when DNA breaks are repaired through non-homologous end joining, unpredictable insertions or deletions can occur, and homology-directed repair is often inefficient in embryos. Off-target effects are another concern, as the guide RNA may bind unintended genomic sites, causing unwanted mutations.
Additionally, while CRISPR-based gene drives can spread edited traits through populations, they require careful ecological risk assessment to avoid unintended consequences, and scaling up edits for population-level interventions demands extensive breeding and monitoring. Despite these challenges, CRISPR remains the most practical and validated method for achieving heritable and targeted genetic modifications in mosquitoes.
References
Michael, T.P. & VanBuren, R., 2020. Building near-complete plant genomes. Current Opinion in Plant Biology, 54, pp.26–33.
Thomsen, H.C. et al., 2014. Glutamine synthetase: role in nitrogen metabolism and crop productivity. Frontiers in Plant Science, 5, p.465. The sequence of sequencers: The history of sequencing DNA https://pmc.ncbi.nlm.nih.gov/articles/PMC4727787/
Oxford Nanopore Technologies – Official Documentation https://nanoporetech.com](https://nanoporetech.com/
Wang, W. et al., 2018. Genetic variation in ARE1 mediates grain yield by modulating nitrogen utilization in rice. Nature Communications, 9, p.735.
Wenger, A.M. et al., 2019. Accurate circular consensus long-read sequencing improves variant detection and genome assembly. Nature Biotechnology, 37, pp.1155–1162.
Xu, G. et al., 2012. Plant nitrogen assimilation and use efficiency. Annual Review of Plant Biology, 63, pp.153–182.
I also used Chat-gtp to guide me in the steps, and design procedures used in read , write and edit DNA questions.