Week 3 HW: Lab Automation


Python Script for Opentrons Artwork
This has been the most interesting and somewhat challenging assignment so far. I chose make an artistic design based on the adrinkra symbols. The adinkra symbols are a set of visual symbols from Ghana, created by the Akan people to represent philosophical concepts, historical events, and social proverbs.
I picked the Nsaa symbol, which is a type of woven cloth renowned for its quality. It is the symbol of excellence, genuineness, and authenticity.


I used the GUI at opentrons-art.rcdonovan.com.to generate an artistic design for the Nsaa symbol.
https://opentrons-art.rcdonovan.com/?id=i96nm69hbxsi9ck
I then used the design coordinates from the Opentrons Automation Art Interface to write the code in Google Colab.
Writing the code to ensure that the colors were dispensed at the correct coordinates and there was no cross-contamination of pipette tips was very tricky. I wrote my first attempt and used the Gemini 2.5 flash in Google Colab to optimize and debug errors in the code. When writing the code, I noticed that the color wells available were red, green, and orange, so I used green and red for the design.
link to Google Colab:
Visualization of the code
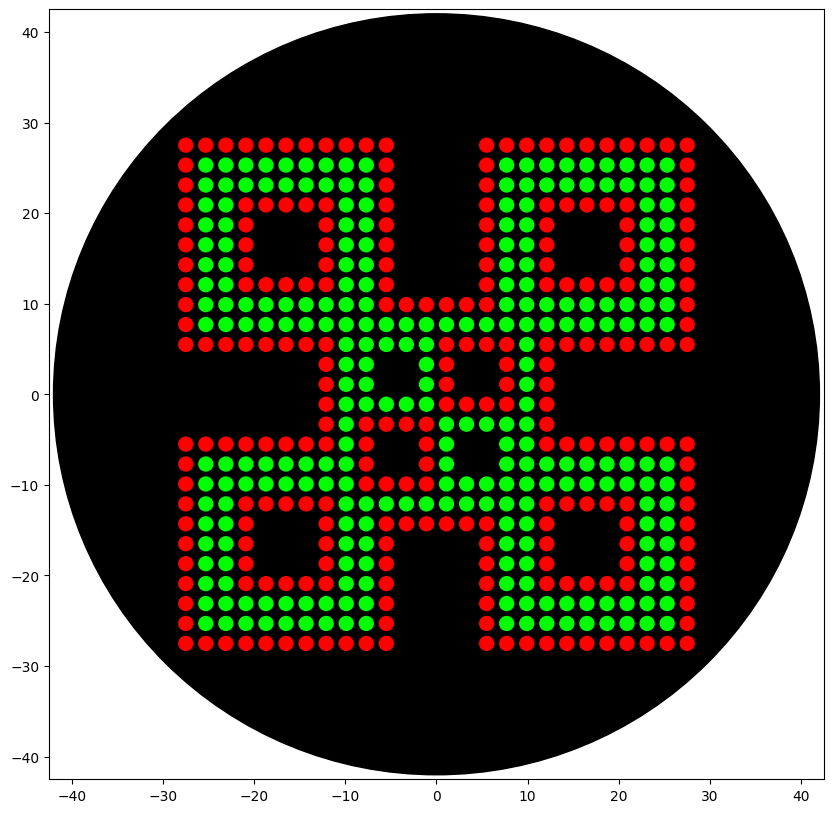

This table is the documentation of how I used Google Genimi 2.5 flash in Google Colab to help debug my code.
| Error | Prompt | Fix |
|---|---|---|
| General execution error | Please explain this error: “Sorry, I ran into an error, could you try again?” | The issue was caused by an indentation error in the for loops iterating over the ‘Green’ and ‘Red’ coordinates. The indentation was corrected and metadata fields were updated with placeholder values. |
| AttributeError: ‘Location’ object has no attribute ‘moves’ | Please explain this error | The Location object does not have a moves method. The correct Opentrons API method is move. Replacing moves with move resolved the issue. |
| Pipette dispense error (no liquid) | Please explain this error | The pipette attempted to dispense without aspirating first. The fix was to add aspiration steps before dispensing both ‘Green’ and ‘Red’ solutions. |
| Tip not dropped error | The robot is reporting that the tip was not dropped | The protocol likely stopped earlier due to incorrect aspiration logic. The aspiration volumes were revised to explicitly match dispense volumes, allowing the protocol to complete and drop the tip properly. |
| KeyError: Labware well names not found | Please explain this error | The labware 'opentrons_96_aluminumblock_generic_pcr_strip_200ul' does not use well names like ‘A1’, ‘B1’, or ‘C1’. It was replaced with 'corning_96_wellplate_360ul_flat', which supports standard 96-well naming. |
| Cross-contamination error | Please explain this error | The same pipette tip was used for both green and red solutions. The fix was to add a drop_tip() step after finishing with green and pick up a new tip before handling red. |
| Visualization color missing | Coordinate (9.9, -16.5) for green doesn’t have a color showing in the visualization. What is the fix? | There was a typo in the Green coordinate list. (9.9, 16.5) was listed instead of (9.9, -16.5). Correcting the coordinate fixed the visualization. |
I also created other designs using the GUI :
https://opentrons-art.rcdonovan.com/?id=20719mxv010c1y8
https://opentrons-art.rcdonovan.com/?id=k2p8012s471hljv
Post-Lab Questions
Question 1: Revolutionizing sample preparation: a novel autonomous microfluidic platform for serial dilution
The paper I chose is titled “Revolutionizing sample preparation: a novel autonomous microfluidic platform for serial dilution” by Dries Vloemans et al. The paper presented a novel, standalone, and fully automated microfluidic platform for the stepwise preparation of serial dilutions without the need for any active elements.
Dilution is a standard fluid operation that is widely employed in the sample preparation of many biochemical assays. It serves multiple essential functions, such as sample mixing with certain reagents at specific dilution ratios, reducing sample matrix effects, and bringing target analytes within the linear assay detection range, among many others.
Traditionally, dilution relies either on manual pipetting, which is labor-intensive and prone to human error, or automated laboratory liquid handling systems, which are bulky, expensive, and unsuitable for point-of-care use. The goal of the authors was to develop a passive, self-contained microfluidic platform that could execute serial dilution in a controlled, programmable, and reproducible manner.
The key findings of the paper include demonstrating that the proposed automated microfluidic platform can perform precise and reproducible serial dilutions without pumps or active control systems. The hydrophobic burst valves reliably metered out defined liquid volumes, enabling accurate dilution ratios such as 2X, 5X, and 10X. It also showed that effective mixing could be achieved through the incorporation of sequential expansion chambers. Which were geometrically optimized to promote passive mixing as fluids pass through them, eliminating the need for mechanical agitation. Additionally, it demonstrated the platform’s compatibility with relevant biological fluids like blood and the integration of a capillary-driven SIMPLE pumping mechanism to allow the device to operate in a fully self-powered manner and complete dilution sequences within short time frames after user activation.
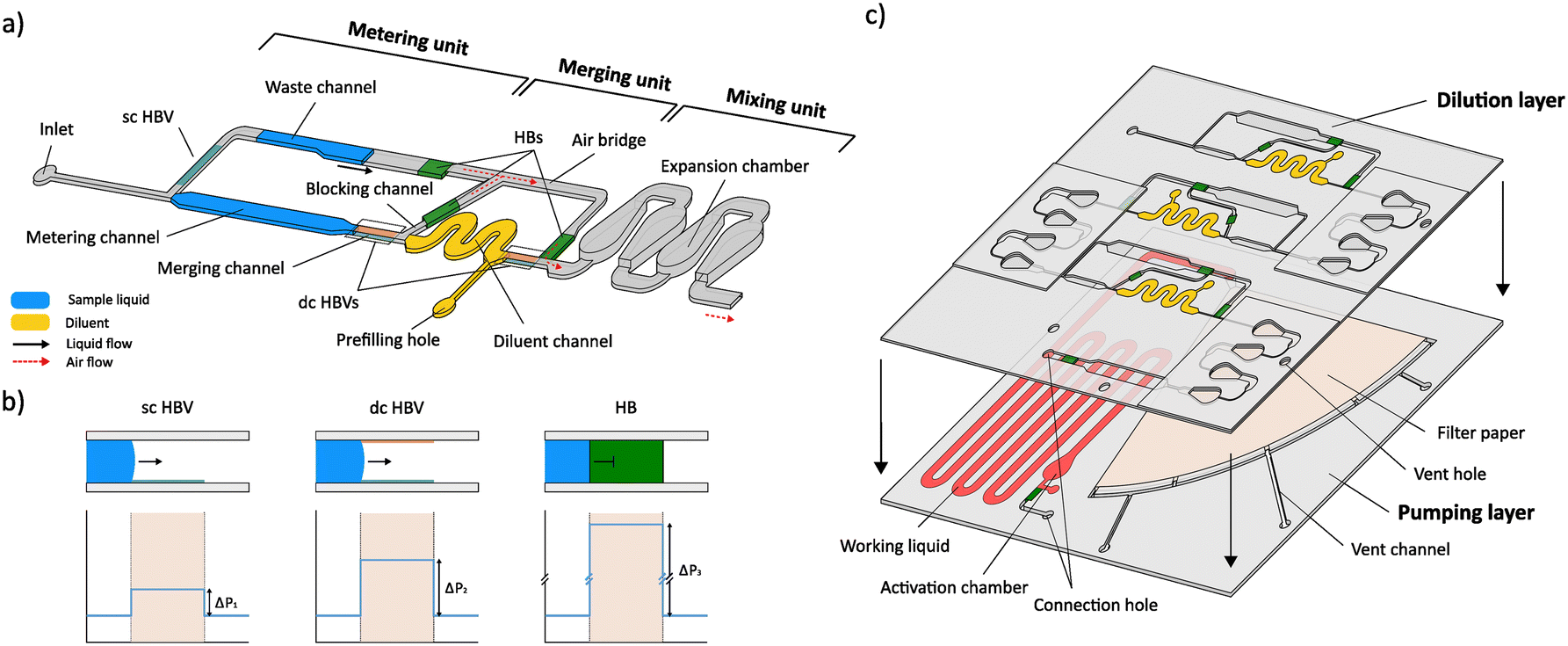

Fig. 1 a) Conceptual design of the dilution module illustrating the 3 microfluidic units that are used for plug metering, merging and mixing, and the positions of the different valving elements (single-coated (sc) and double-coated (dc) HBVs, and hydrophobic barrier (HB)). b) Configuration and working principle of the different valves with their respective theoretical burst pressure profiles. The sc HBV contains a hydrophobic coating at the bottom channel wall, while the dc HBV is treated hydrophobically at both the top and bottom walls, resulting in varying burst pressures. The HB comprises a hydrophobic-treated filter paper, which allows air passage but forms a physical barrier for the liquid, hence, inducing a very high burst pressure. c) Conceptual exploded view of the integrated microfluidic device for autonomous multistep serial dilution, illustrating the top ‘dilution’ and bottom ‘pumping’ layer. The top dilution layer comprises 3 serially coupled dilution modules (5× DF), connected with a connection hole to the bottom pumping layer, holding the prefilled working liquid and wedge-shaped filter paper (Whatman grade 598) of the SIMPLE pump unit.
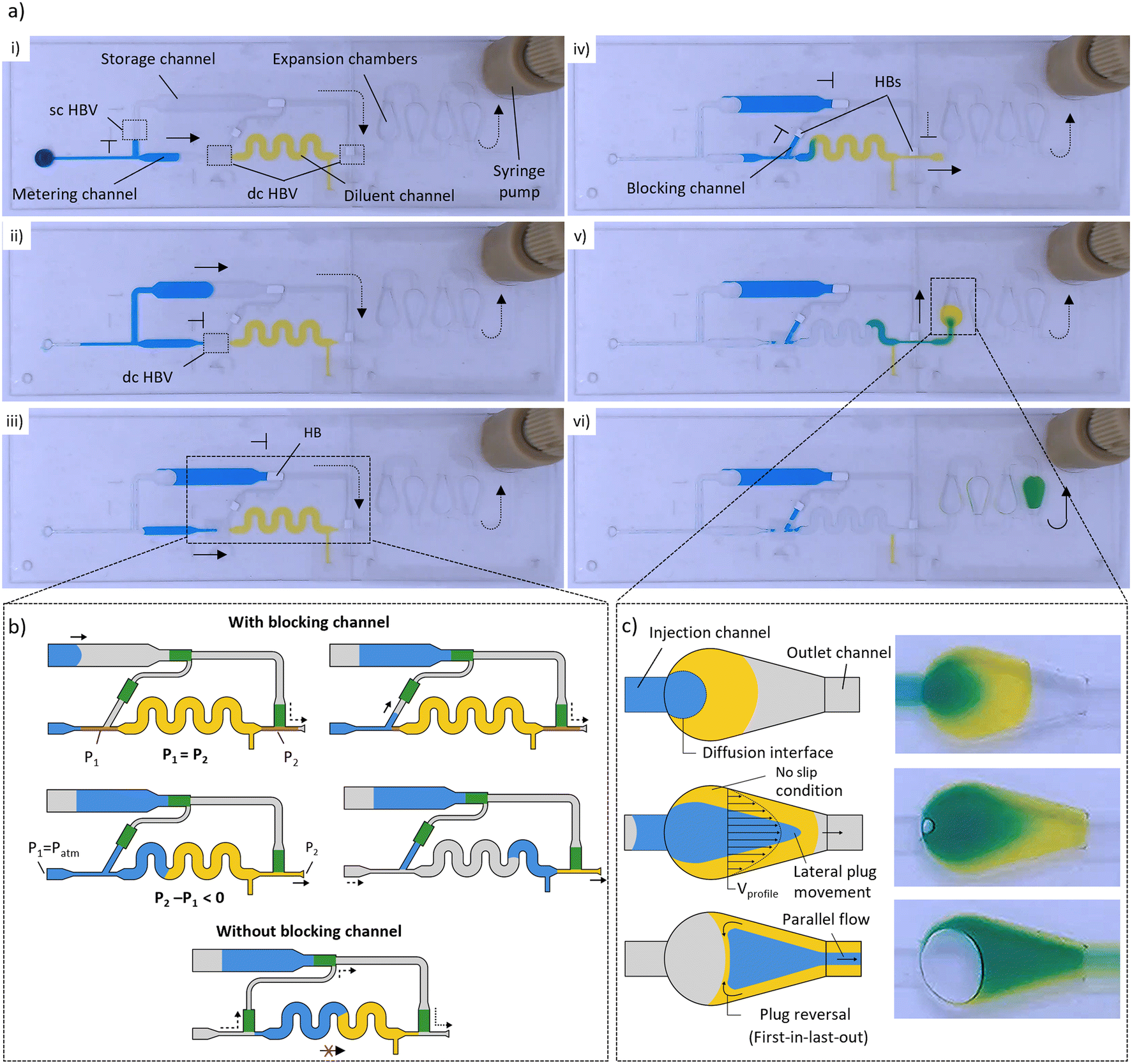

Fig. 2 a) Snapshots of the different liquid manipulations within a dilution module (DF = 5×) illustrating the working principle. (i and ii) The coordinated burst action of HBVs with different burst strengths is used to first isolate a precisely metered sample liquid (2 μL, blue), after which the excess is removed to the storage channel. (iii and iv) The metered sample liquid is next merged with a prefilled diluent (8 μL, yellow), after which (v and vi) the combined plug is sent through a sequence of expansion chambers in which it is mixed into a homogeneous solution. b) Detailed schematics of plug merging, and working principle of the microfluidic air bridge (top). Illustration of failed downstream plug manipulation when no blocking channel is used due to air intake via the microfluidic air bridge (bottom). c) Close-up of the expansion chambers, illustrating the three ongoing principles that are used within the mixing process: increase of diffusion interface, parabolic flow profile, and lateral plug distribution. Dashed and full arrows indicate air and liquid flow, respectively.
Question 2: What I intend to do with automation tools for my final project
Using my first idea, which involves developing a biosensor kit for the detection of illegal mining pollutants. The automation tools used would be a combination of Python-based liquid handling, 3D-printed assay holders, and could-based design tool like Google Nebula.
Here is a rough idea of the automation tools I might end up using:
- Using Opentrons OT-2 to dispense growth media and mix microbial cultures with chemical regents.
- Using PLateLoc to seal the plate.
- Using XPeel to remove the seal after incubation.
- Measuring fluorescence and color intensity using PHERAstar plate reader.
- Using Ginkgo Nebula to design synthetic genetic circuits for microbial biosensors and simulate sensor response behaviors.
Final Project Ideas
I have submitted my final project ideas in the slide deck that was provided for committed listeners
Reference
https://opentrons-art.rcdonovan.com/?id=i96nm69hbxsi9ck
Vloemans, D., Pieters, A., Dal Dosso, F. and Lammertyn, J., 2024. Revolutionizing sample preparation: a novel autonomous microfluidic platform for serial dilution. Lab on a Chip, 24(10), pp.2791-2801.