Week 3: Lab Automation & Protest BioArt
1. Artistic Concept: “Woman Life Freedom”


This project is more than a laboratory exercise; it is a piece of Protest Art. It is dedicated to the current situation in Iran, where thousands of people have been killed or continue to face violence in the streets while fighting for their fundamental freedoms.
The design features a raised fist—a universal symbol of resistance—accompanied by the slogan “Woman, Life, Freedom.” By using living, glowing bacteria to form this image, I aim to represent the resilience and the “living” spirit of the struggle for liberty in my country.
Color & Protein Mapping:
- Blue (TagBFP): Used for the main outline, hair, and text to provide a strong, deep contrast.
- Pink (mRFP1): Used for highlighting the eye and cheek details, representing vitality.
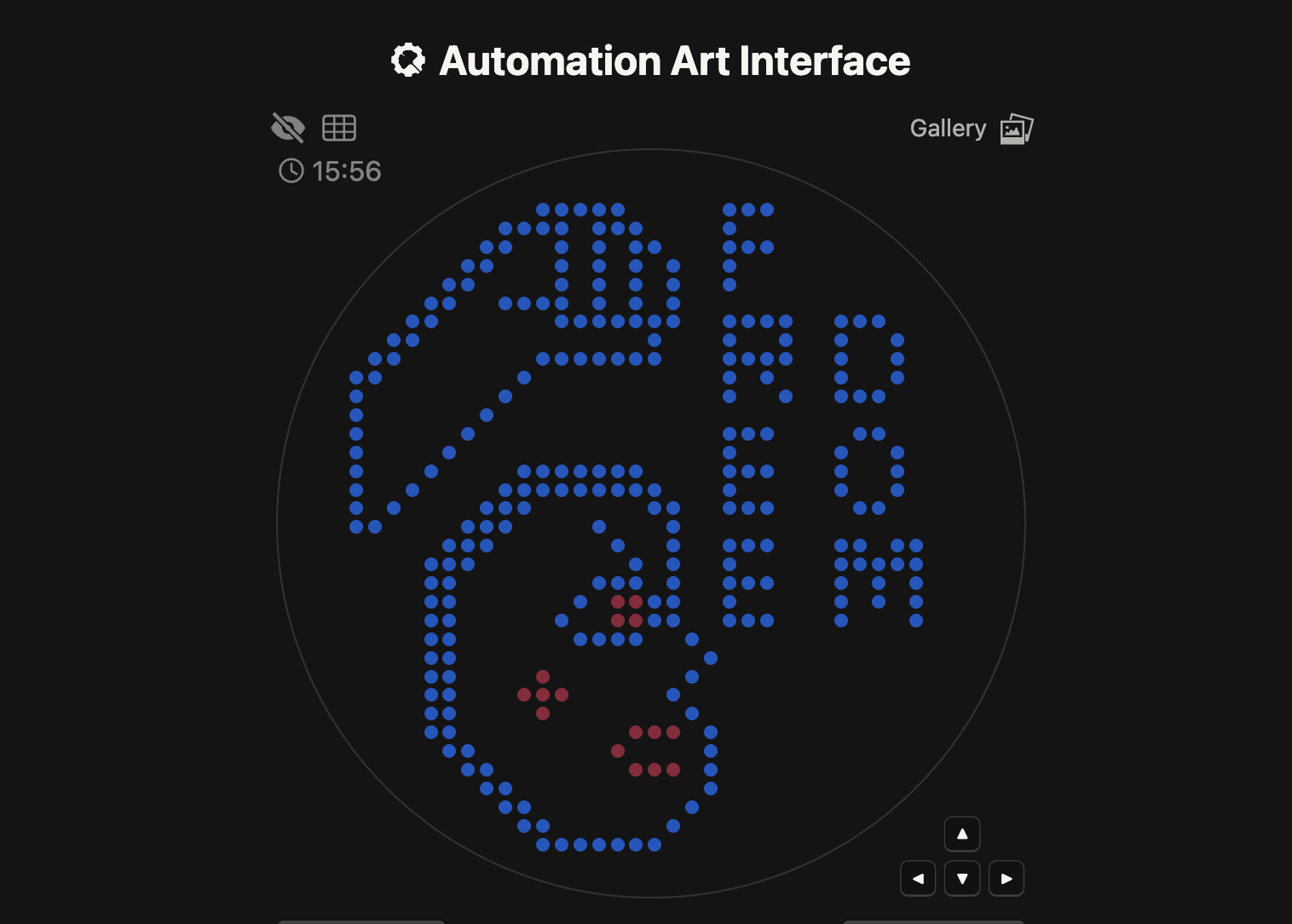

2. Technical Implementation
For this assignment, I utilized the Automation Art Interface (GUI) to map my design onto a 90mm circular Petri dish.
- Code Source: As permitted by the assignment instructions for those focusing on the art concept, I have used the Full Python Script generated by the GUI website (opentrons-art.rcdonovan.com).
- Labware Configuration: The script is configured for a 96-well PCR plate to hold the fluorescent bacterial strains.
- Robotic Protocol: The Opentrons robot uses
pick_up_tip(),aspirate(), and the specializeddispense_and_jog()function to deposit 0.75µL droplets without damaging the charcoal agar surface. - Spacing: A 2.2mm spacing was chosen for optimal resolution and clarity under UV light.
3. Design Files
- Coordinates List:
Blue Points (TagBFP) - 194 points
tagbfp_points = [(-13.2, 37.4), (-11, 37.4), (-8.8, 37.4), (-6.6, 37.4), (-4.4, 37.4), (8.8, 37.4), (11, 37.4), (13.2, 37.4), (-17.6, 35.2), (-15.4, 35.2), (-13.2, 35.2), (-11, 35.2), (-6.6, 35.2), (-4.4, 35.2), (-2.2, 35.2), (8.8, 35.2), (-19.8, 33), (-17.6, 33), (-11, 33), (-6.6, 33), (-2.2, 33), (0, 33), (8.8, 33), (11, 33), (13.2, 33), (-22, 30.8), (-19.8, 30.8), (-11, 30.8), (-6.6, 30.8), (-2.2, 30.8), (2.2, 30.8), (8.8, 30.8), (-24.2, 28.6), (-22, 28.6), (-11, 28.6), (-6.6, 28.6), (-2.2, 28.6), (2.2, 28.6), (8.8, 28.6), (-26.4, 26.4), (-24.2, 26.4), (-17.6, 26.4), (-15.4, 26.4), (-13.2, 26.4), (-11, 26.4), (-6.6, 26.4), (-2.2, 26.4), (2.2, 26.4), (-28.6, 24.2), (-26.4, 24.2), (-11, 24.2), (-8.8, 24.2), (-6.6, 24.2), (-4.4, 24.2), (-2.2, 24.2), (0, 24.2), (2.2, 24.2), (8.8, 24.2), (11, 24.2), (13.2, 24.2), (15.4, 24.2), (22, 24.2), (24.2, 24.2), (26.4, 24.2), (-30.8, 22), (-28.6, 22), (0, 22), (8.8, 22), (15.4, 22), (22, 22), (28.6, 22), (-33, 19.8), (-30.8, 19.8), (-13.2, 19.8), (-11, 19.8), (-8.8, 19.8), (-6.6, 19.8), (-4.4, 19.8), (-2.2, 19.8), (0, 19.8), (8.8, 19.8), (11, 19.8), (13.2, 19.8), (15.4, 19.8), (22, 19.8), (28.6, 19.8), (-35.2, 17.6), (-33, 17.6), (-15.4, 17.6), (8.8, 17.6), (13.2, 17.6), (22, 17.6), (28.6, 17.6), (-35.2, 15.4), (-17.6, 15.4), (8.8, 15.4), (15.4, 15.4), (22, 15.4), (24.2, 15.4), (26.4, 15.4), (-35.2, 13.2), (-19.8, 13.2), (-35.2, 11), (-22, 11), (8.8, 11), (11, 11), (13.2, 11), (24.2, 11), (26.4, 11), (-35.2, 8.8), (-24.2, 8.8), (8.8, 8.8), (22, 8.8), (28.6, 8.8), (-35.2, 6.6), (-26.4, 6.6), (-15.4, 6.6), (-13.2, 6.6), (-11, 6.6), (-8.8, 6.6), (-6.6, 6.6), (-4.4, 6.6), (-2.2, 6.6), (8.8, 6.6), (11, 6.6), (13.2, 6.6), (22, 6.6), (28.6, 6.6), (-35.2, 4.4), (-28.6, 4.4), (-17.6, 4.4), (-15.4, 4.4), (-13.2, 4.4), (-11, 4.4), (-8.8, 4.4), (-6.6, 4.4), (-4.4, 4.4), (-2.2, 4.4), (0, 4.4), (8.8, 4.4), (22, 4.4), (28.6, 4.4), (-35.2, 2.2), (-30.8, 2.2), (-19.8, 2.2), (-17.6, 2.2), (-15.4, 2.2), (0, 2.2), (2.2, 2.2), (8.8, 2.2), (11, 2.2), (13.2, 2.2), (24.2, 2.2), (26.4, 2.2), (-35.2, 0), (-33, 0), (-22, 0), (-19.8, 0), (-17.6, 0), (-6.6, 0), (2.2, 0), (-24.2, -2.2), (-22, -2.2), (-19.8, -2.2), (-4.4, -2.2), (2.2, -2.2), (8.8, -2.2), (11, -2.2), (13.2, -2.2), (22, -2.2), (24.2, -2.2), (28.6, -2.2), (30.8, -2.2), (-26.4, -4.4), (-24.2, -4.4), (-22, -4.4), (-2.2, -4.4), (2.2, -4.4), (8.8, -4.4), (22, -4.4), (24.2, -4.4), (26.4, -4.4), (28.6, -4.4), (30.8, -4.4), (-26.4, -6.6), (-24.2, -6.6), (-6.6, -6.6), (-4.4, -6.6), (-2.2, -6.6), (2.2, -6.6), (8.8, -6.6), (11, -6.6), (13.2, -6.6), (22, -6.6), (26.4, -6.6), (30.8, -6.6), (-26.4, -8.8), (-24.2, -8.8), (-8.8, -8.8), (0, -8.8), (2.2, -8.8), (8.8, -8.8), (22, -8.8), (26.4, -8.8), (30.8, -8.8), (-26.4, -11), (-24.2, -11), (-11, -11), (0, -11), (2.2, -11), (8.8, -11), (11, -11), (13.2, -11), (22, -11), (30.8, -11), (-26.4, -13.2), (-24.2, -13.2), (-8.8, -13.2), (-6.6, -13.2), (-4.4, -13.2), (-2.2, -13.2), (4.4, -13.2), (-26.4, -15.4), (-24.2, -15.4), (6.6, -15.4), (-26.4, -17.6), (-24.2, -17.6), (4.4, -17.6), (-26.4, -19.8), (-24.2, -19.8), (2.2, -19.8), (-26.4, -22), (-24.2, -22), (4.4, -22), (-26.4, -24.2), (-24.2, -24.2), (6.6, -24.2), (-24.2, -26.4), (-22, -26.4), (6.6, -26.4), (-22, -28.6), (-19.8, -28.6), (6.6, -28.6), (-19.8, -30.8), (-17.6, -30.8), (6.6, -30.8), (-17.6, -33), (-15.4, -33), (4.4, -33), (-15.4, -35.2), (-13.2, -35.2), (2.2, -35.2), (-13.2, -37.4), (-11, -37.4), (-8.8, -37.4), (-6.6, -37.4), (-4.4, -37.4), (-2.2, -37.4), (0, -37.4)]
Pink Points (mRFP1) - 16 points
mrfp1_points = [(-4.4, -8.8), (-2.2, -8.8), (-4.4, -11), (-2.2, -11), (-13.2, -17.6), (-15.4, -19.8), (-13.2, -19.8), (-11, -19.8), (-13.2, -22), (-2.2, -24.2), (0, -24.2), (2.2, -24.2), (-4.4, -26.4), (-2.2, -28.6), (0, -28.6), (2.2, -28.6)]
Python Script
Post Lab Question 1: Automation Strategy for Final Projects
1. Project Overview: Caff-Ink
For my final project, Caff-Ink, I am developing a silk-based bio-tattoo that monitors systemic caffeine levels using cell-free synthetic biology. The success of this project relies on the precise formulation of bio-inks and the high-definition printing of these inks onto thin silk membranes.
2. Intent for Automation
I intend to use the Opentrons OT-2 liquid handling robot to move beyond manual experimentation into a reproducible manufacturing process. Manual pipetting of cell-free reagents is often inconsistent and wasteful due to the high viscosity and low volumes involved.
Key Automation Goals:
- High-Throughput Ink Screening: I will automate the mixing of various ratios of silk fibroin, cell-free extracts, and caffeine-responsive DNA riboswitches to identify the most stable “Bio-ink”.
- Automated Bio-Printing: By utilizing the robot’s X-Y-Z precision, I will program the OT-2 to “draw” specific artistic patterns onto silk substrates, ensuring uniform thickness and chemical distribution.
3. Hardware and 3D Printed Components
To adapt the Opentrons deck for my specific needs, I will design and 3D print the following:
- Silk Patch Stabilizer: A custom plate that holds multiple silk membranes under tension, preventing them from shifting during the dispensing process.
- Micro-Reagent Rack: A specialized holder for 1.5ml tubes that minimizes dead volume, ensuring every microliter of expensive cell-free reagent is utilized.
4. Opentrons Python Protocol (Pseudocode)
The following script illustrates the logic for automated printing of a 10-point gradient tattoo pattern on a silk substrate:
project 2
Automation Strategy for Project: Dia-Lens
** 1. Intent for Automation:** For the Dia-Lens project, I will use the Opentrons OT-2 to automate the precise mixing of engineered S. epidermidis with the hydrogel matrix. Automation is essential to ensure a consistent microbial load across all lenses, which is critical for both lens transparency and accurate therapeutic dosing.
** 2. 3D Printed Components:**
- Ocular Mold Rack: A custom 3D-printed holder designed to secure 12 contact lens molds on the Opentrons deck, ensuring the pipette can dispense the bio-ink into the center of each mold with sub-millimeter precision.
** 3. Python Pseudocode Logic:**
project 3
Automation Strategy for Project: The Living Glow
The Living Glow focuses on a personalized probiotic system using engineered Saccharomyces cerevisiae to address specific skin conditions via the gut-skin axis. Unlike “one-size-fits-all” supplements, this project aims to create custom yeast cocktails tailored to an individual’s skin profile—such as dehydration, acne-prone, or inflammatory conditions.
1. Intent for Automation: The “Digital Pharmacist”
I will use the Opentrons OT-2 as an automated formulation station to create personalized probiotic doses. Automation is the only way to achieve the precise mixing ratios required for multi-strain therapies.
Key Automation Goals:
- Personalized Strain Blending: The robot will receive a digital “Skin Profile” and automatically mix different engineered yeast strains (e.g., Hyaluronic Acid producers for dry skin vs. Antimicrobial Peptide producers for acne) in specific ratios.
- High-Precision Encapsulation: Automated dispensing of these custom ratios into capsule shells to ensure dose consistency and prevent cross-contamination between different patient formulations.
2. Hardware and 3D Printed Components
- Dynamic Capsule Matrix: A 3D-printed rack designed to hold various capsule sizes, allowing the Opentrons to fill individual “personalized prescriptions” in a high-throughput manner.
Part 2: Summary of a Published Automation Paper
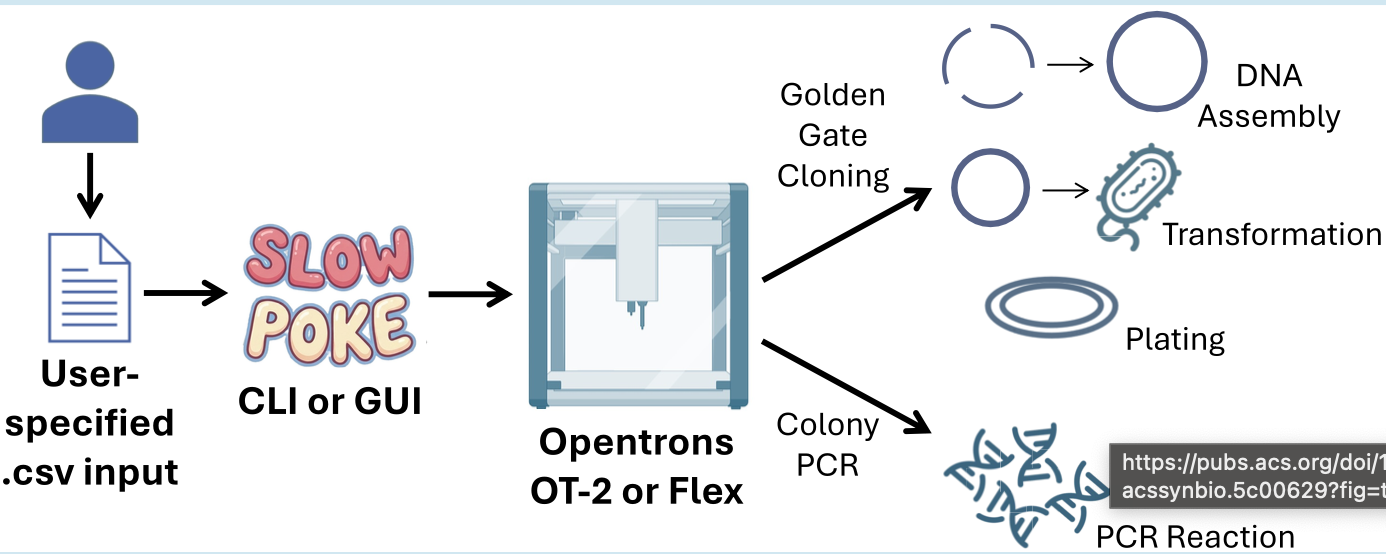

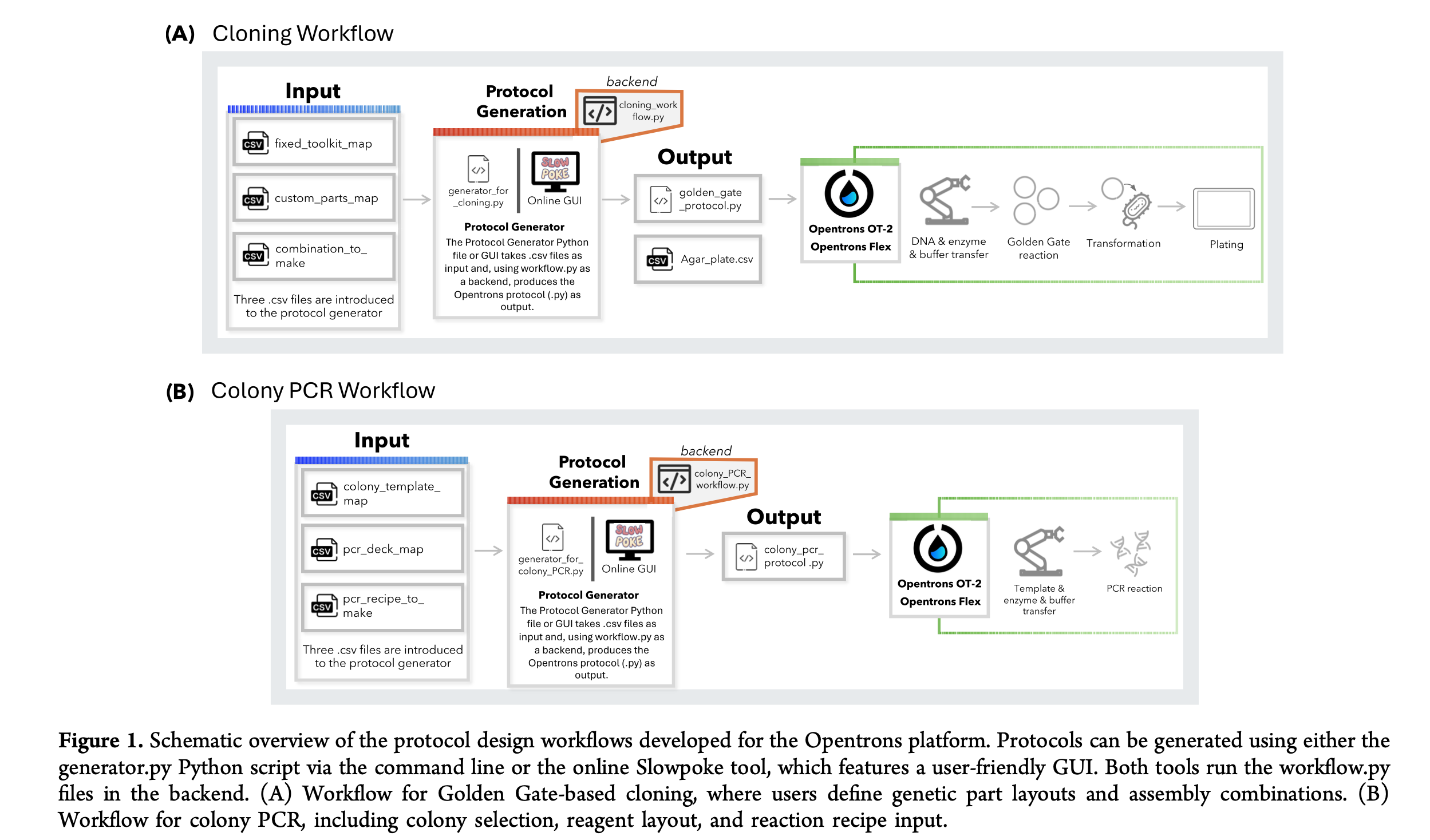
Paper Reference: Malcı, K., et al. (2026). Slowpoke: An Automated Golden Gate Cloning Workflow for Opentrons OT-2 and Flex. ACS Synthetic Biology.
General Overview
Synthetic biology relies heavily on DNA assembly, a routine but labor-intensive process that is often prone to human error when performed manually. To address these challenges, the authors developed Slowpoke, an open-source and modular automation workflow specifically designed for the accessible Opentrons OT-2 and Flex liquid-handling platforms. The system automates the most critical steps of the Golden Gate cloning cycle, including DNA assembly, Escherichia coli transformation, agar plating, and colony PCR.
A key innovation of this research is the introduction of a free Graphical User Interface (GUI) that allows researchers to generate complex Python protocols without any coding knowledge. By simply uploading a CSV file, the GUI converts biological parameters into instructions the robot can execute, requiring manual intervention only for colony picking and plate transfers. This significantly lowers the barrier to high-throughput genetic engineering for laboratories with limited programming expertise.
Findings
The validation of the Slowpoke workflow demonstrated its high reliability across different biological systems and automation hardware. Key results include:
- Broad Compatibility: The effectiveness of the system was validated using established genetic toolkits for both yeast (S. cerevisiae) and bacteria (B. subtilis).
- Assembly Precision: The Opentrons OT-2 platform achieved a 100% success rate (17/17 positive colonies) for basic transcription unit assemblies.
- High-Throughput Scalability: In a complex assembly test involving six DNA parts and 57 different combinations, 55 out of 57 resulted in the correct genetic constructs.
- System Robustness: These results confirm that Slowpoke provides a standardized and scalable solution, making it highly reliable for modern synthetic biology laboratories.
Relevant Figures