Week 2 HW: DNA R/W/E
My Homework
DNA ;)


HW1 — Part 0: Basics of Gel Electrophoresis
Done (new account in Benchling: @Nicorriza)
Then imported Escherichia phage Lambda complete genome
GenBank: J02459.1 — from NCBI page to Benchling
HW2 — Part 1: Benchling & In-silico Gel Art
Everything below is completed:
Created a free account at benchling.com
Imported the Lambda DNA
Simulated restriction enzyme digestion with the following enzymes:
EcoRI — GAATTC — G|AATTC
HindIII — AAGCTT — A|AGCTT
BamHI — GGATCC — G|GATCC
KpnI — GGTACC — GGTAC|C
EcoRV — GATATC — GAT|ATC (blunt)
SacI — GAGCTC — GAGCT|C
SalI — GTCGAC — G|TCGAC
Virtual digest result
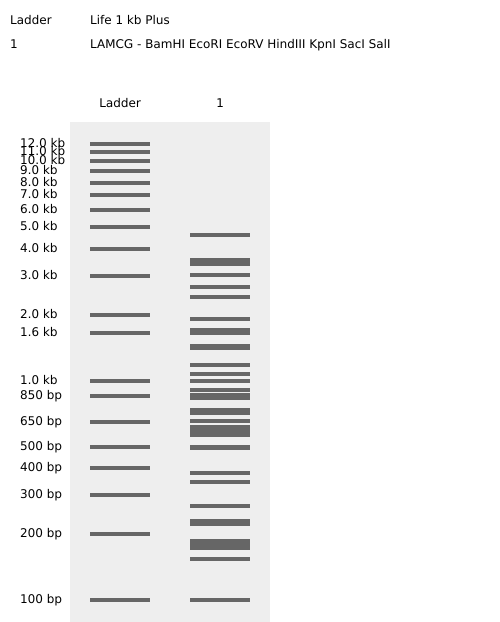

Continuing with the work:
(Create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks.
You might find Ronan’s website a helpful tool for quickly iterating on designs.)
I created a pareidolic face in an agarose gel pattern using the following restriction enzyme digests:
- Ladder — Life 1 kb Plus
- LAMCG — KpnI, PvuII
- LAMCG — PvuII
- LAMCG — PvuII, SacI
- LAMCG — PvuII
- LAMCG — PvuII, SacI
Resulting in:
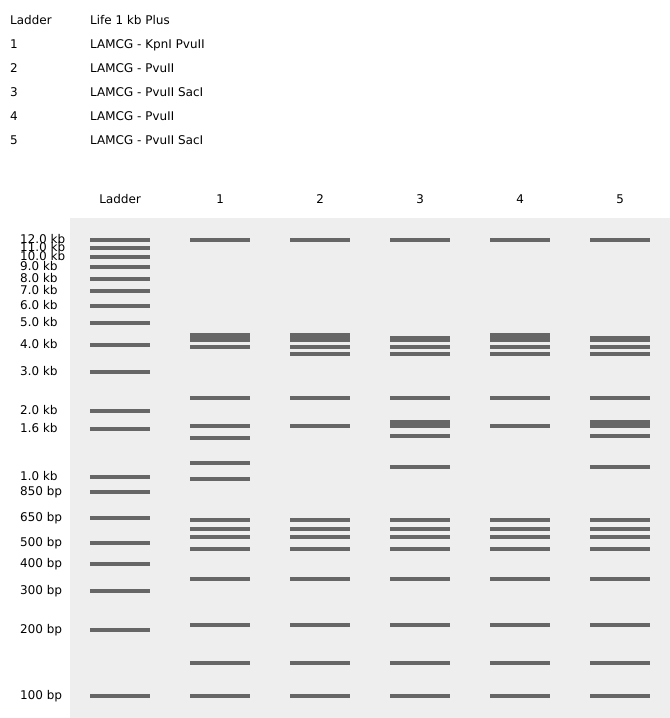


The face appears in this region of the agarose gel between 2.0 kb and 1.0 kb.


Part 3: DNA Design Challenge
3.1. Choose Your Protein
Prompt
In recitation we discussed that you should choose a protein that seems interesting for your task. Which protein did you choose and why? Using one of the tools discussed in recitation (NCBI, UniProt, Google), obtain the sequence of the protein you chose.
Answer
During the course we were introduced to FPbase, a repository of fluorescent proteins that I found very interesting. I decided to explore proteins with blue fluorescence because that color is uncommon in nature and seemed intriguing for the project. The protein I selected was HcRed1-Blue for that reason.
Below is the amino-acid sequence I obtained for the selected protein:
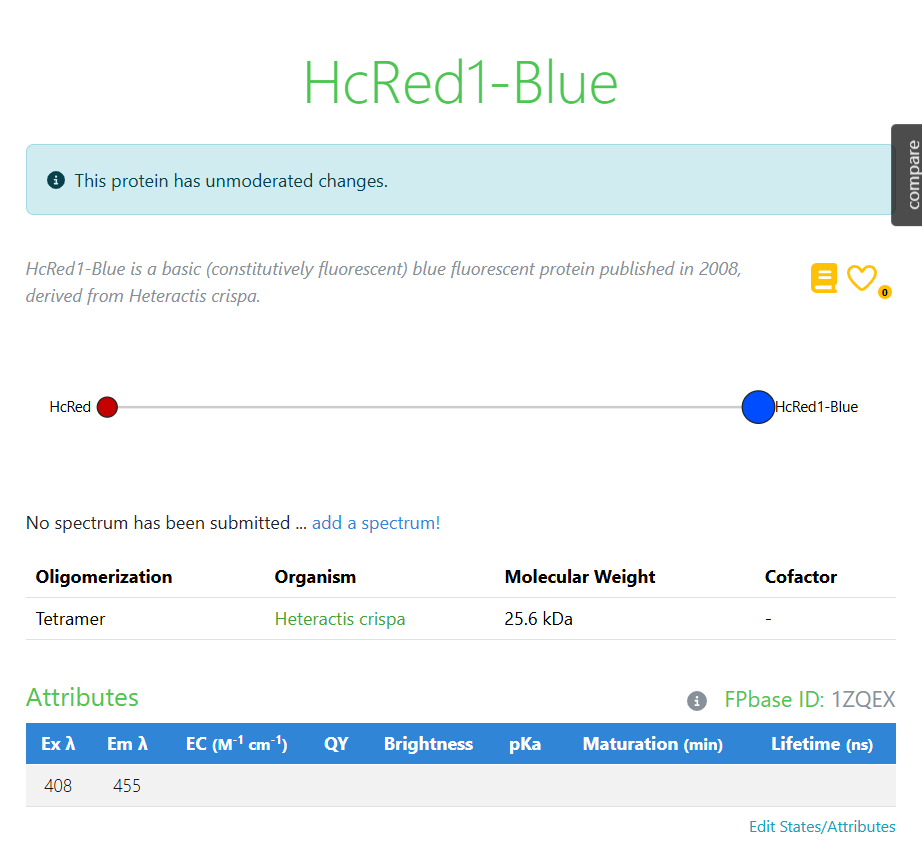
3.2. Reverse Translation: Protein → DNA
Prompt
Using tools (NCBI or online reverse-translation tools), determine the nucleotide sequence that corresponds to your chosen protein sequence.
Tool Used
I used the Reverse Translate tool at Bioinformatics.org with the default codon usage table (based on E. coli).
Result (Reverse Translate — 684 bases)
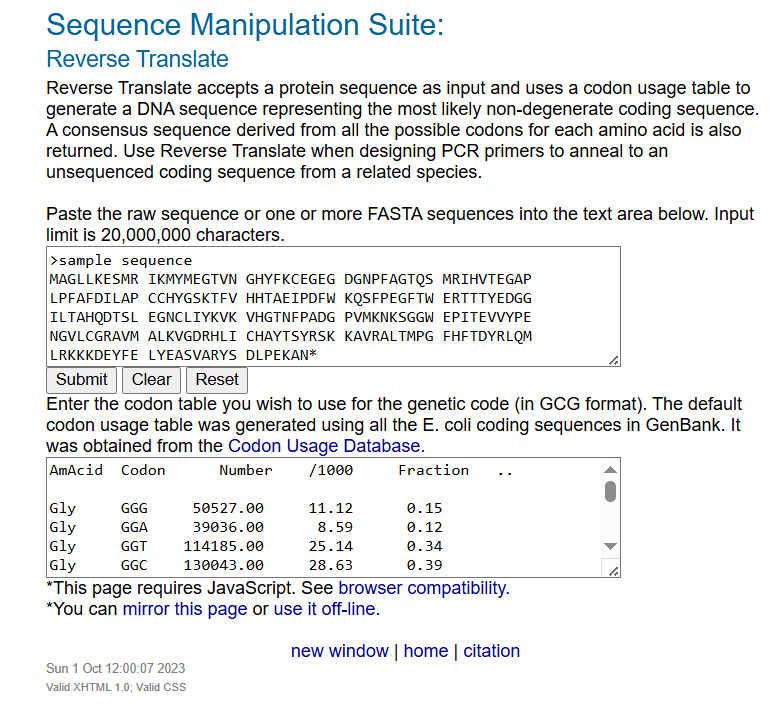
Results:
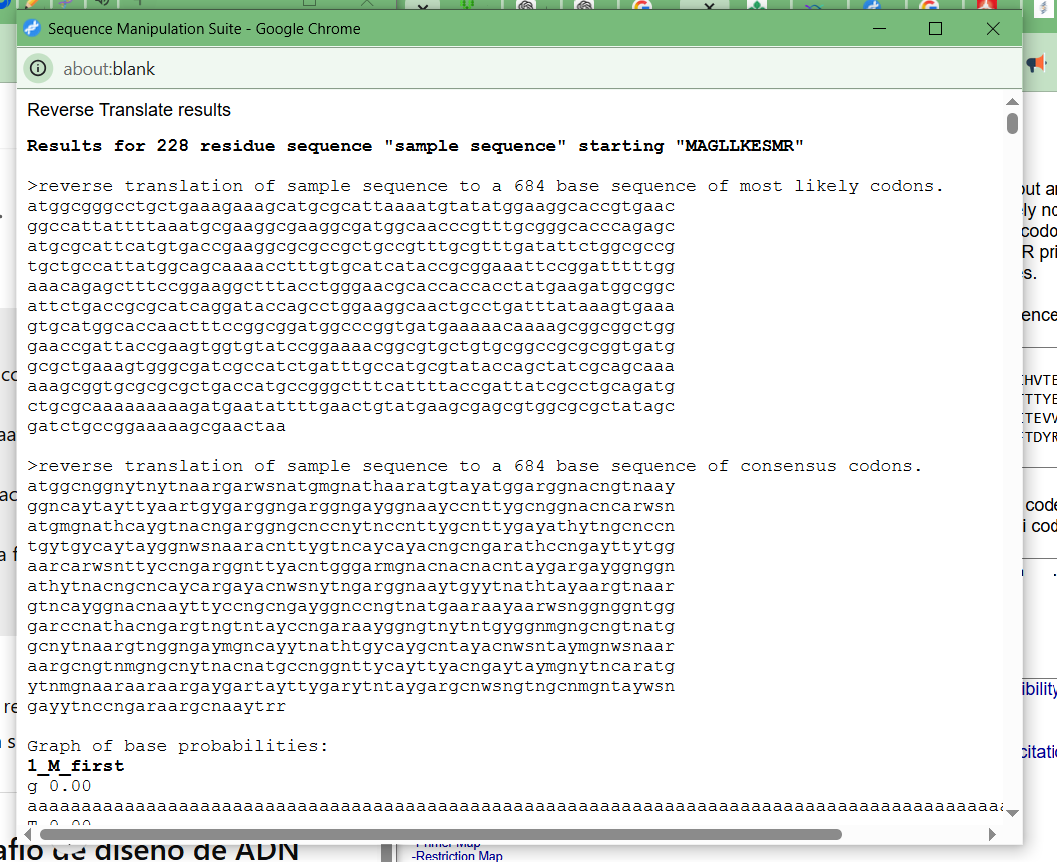
Note: This sequence corresponds to the most likely codons according to the default table (E. coli).
3.3. Codon Optimization
Prompt
Explain why you need to optimize codon usage. For which organism did you choose to optimize the sequence and why?
Answer
Why Optimize Codons?
Although the genetic code is universal, codon usage frequencies differ between organisms. Codon optimization improves translation efficiency and protein expression in the chosen host. It also helps control:
- Codon frequency (adaptation to the host)
- GC content (affects stability and transcription)
- Undesired mRNA secondary structures
- Internal restriction sites
- Repetitive sequences that complicate DNA synthesis
Organism Chosen for Optimization
For this exercise I used the default orientation toward E. coli (a practical choice for bacterial expression). Tools such as IDT provide organism-specific codon optimization options (e.g., E. coli, S. cerevisiae, Drosophila melanogaster, Mus musculus).
Example Optimized Sequence (Formatted in Codons)
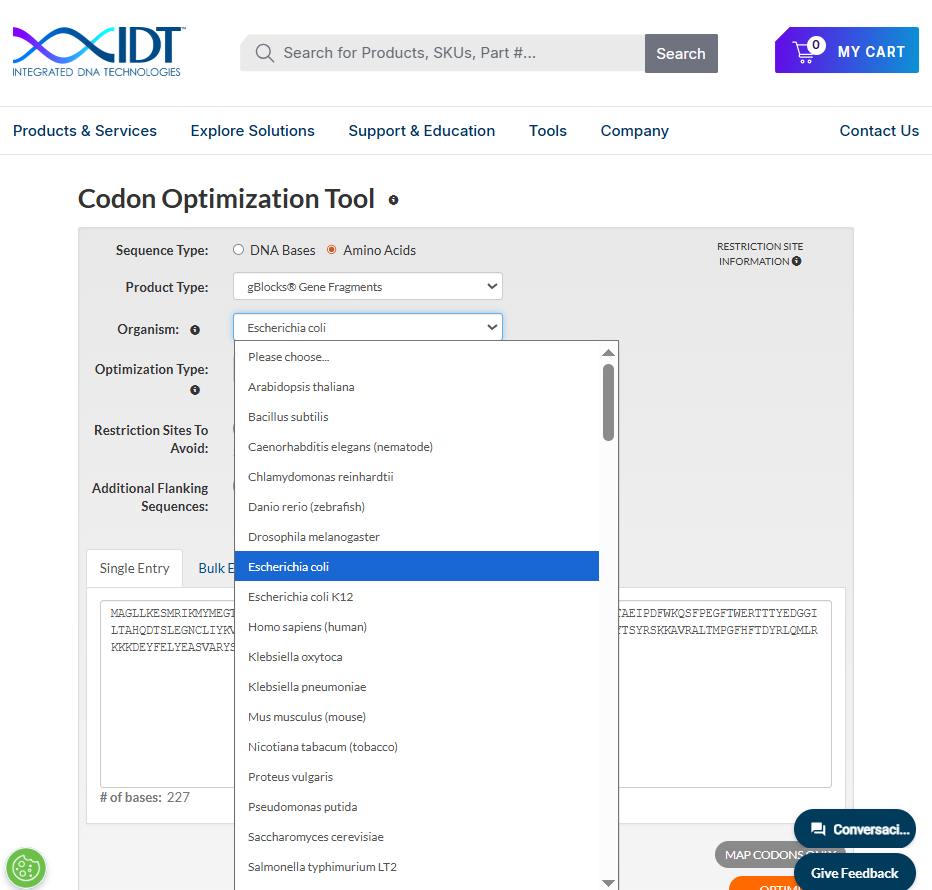
Avoiding Specific Restriction Sites (IDT Optimization Settings)
Using the IDT Codon Optimization tool, it is also possible to exclude specific restriction enzyme recognition sites from the DNA sequence. This is particularly useful when designing constructs for cloning, since it prevents unwanted internal cutting within the gene of interest.
For this design, the following restriction sites were explicitly avoided:
- EcoRI
- BamHI
- SacII
- SalI
- HindIII
- EcoRV
By removing these internal restriction sites during optimization, the sequence becomes compatible with vectors that use these enzymes in their multiple cloning site (MCS), facilitating downstream cloning and assembly steps.
Sequence Notes
- Codon optimized for E. coli
- GC content balanced : GC% = 47.09%
- Restriction sites screened
- Ready for plasmid cloning
Detected Restriction Sites (Examples to Avoid or Consider)
- BglII — AGATCT
- BspEI — TCCGGA
- DraI — TTTAAA
- MlyI — GAGTC
- PmlI — CACGTG
- PstI — CTGCAG
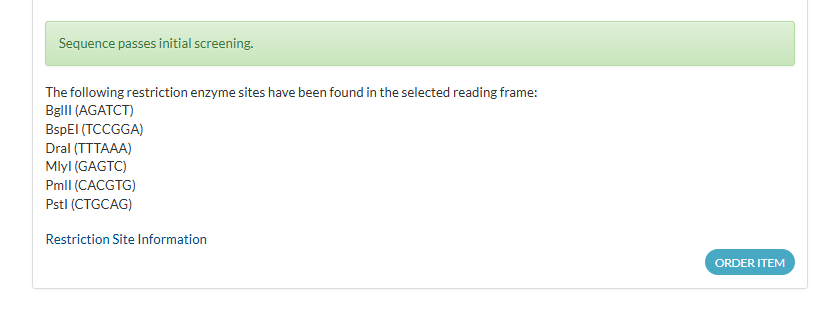
3.4. You Have a Sequence — Now What?
Prompt
Which technologies could be used to produce this protein from your DNA? Describe how the DNA sequence can be transcribed and translated into your protein (cell-dependent and/or cell-free methods).
Answer
Cell-Dependent Methods
Expression in Bacteria (e.g., E. coli)
- Clone the optimized sequence into an expression plasmid (strong promoter, RBS, terminator).
- Transform the plasmid into competent E. coli.
- Induce expression (e.g., IPTG in lac-based systems).
- The cell transcribes DNA → mRNA and translates mRNA → protein.
- After expression, the protein can be purified (e.g., chromatography).
Other possible systems include yeast (S. cerevisiae) or mammalian cells if post-translational modifications are required.