Week 3 HW: Lab Automation
Assignment: Python Script for Opentrons Artwork
Python file to run on an Opentrons liquid handling robot
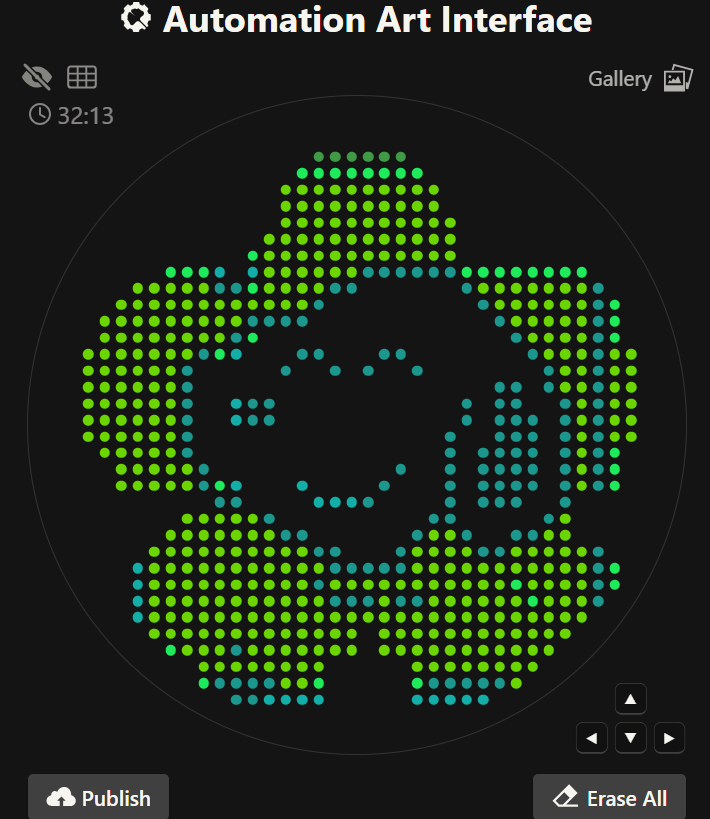

Opentrons Colab
https://colab.research.google.com/drive/1FSLFjeZfWudyaujskQb60dIuhlPAv9Nq?usp=sharing
Python Script from GUI
OTDesign_02-24-26_10-43-25.py
from opentrons import types
import string
metadata = {
'protocolName': '{YOUR NAME} - Opentrons Art - HTGAA',
'author': 'HTGAA',
'source': 'HTGAA 2026',
'apiLevel': '2.20'
}
Z_VALUE_AGAR = 2.0
POINT_SIZE = 0.75
venus_points = [(-3.3,23.1), (-1.1,23.1), (1.1,23.1), (3.3,23.1), (5.5,23.1), (7.7,23.1), (9.9,23.1), (12.1,23.1), (-3.3,20.9), (-1.1,20.9), (-7.7,18.7), (-5.5,18.7), (16.5,18.7), (-16.5,16.5), (-14.3,16.5), (-12.1,16.5), (-9.9,16.5), (-7.7,16.5), (18.7,16.5), (-16.5,14.3), (20.9,14.3), (-23.1,12.1), (-20.9,12.1), (-18.7,12.1), (23.1,12.1), (-23.1,9.9), (25.3,9.9), (-25.3,7.7), (-25.3,5.5), (27.5,5.5), (-25.3,3.3), (29.7,3.3), (-25.3,1.1), (29.7,1.1), (-25.3,-1.1), (29.7,-1.1), (-25.3,-3.3), (29.7,-3.3), (-23.1,-5.5), (29.7,-5.5), (-20.9,-7.7), (29.7,-7.7), (-18.7,-9.9), (29.7,-9.9), (-16.5,-12.1), (-14.3,-12.1), (27.5,-12.1), (-12.1,-14.3), (9.9,-14.3), (25.3,-14.3), (-9.9,-16.5), (-7.7,-16.5), (7.7,-16.5), (12.1,-16.5), (20.9,-16.5), (23.1,-16.5), (-5.5,-18.7), (14.3,-18.7), (16.5,-18.7), (18.7,-18.7), (-3.3,-20.9), (-1.1,-20.9), (1.1,-20.9), (3.3,-20.9), (5.5,-20.9), (7.7,-20.9), (-5.5,-23.1), (3.3,-23.1), (-5.5,-25.3), (-3.3,-25.3), (-1.1,-25.3), (1.1,-25.3), (3.3,-25.3), (5.5,-25.3)]
mjuniper_points = [(1.1,20.9), (3.3,20.9), (5.5,20.9), (7.7,20.9), (9.9,20.9), (12.1,20.9), (-3.3,18.7), (-1.1,18.7), (14.3,18.7), (-5.5,16.5), (16.5,16.5), (-14.3,14.3), (-12.1,14.3), (-9.9,14.3), (-7.7,14.3), (18.7,14.3), (-16.5,12.1), (20.9,12.1), (-20.9,9.9), (-7.7,9.9), (-5.5,9.9), (3.3,9.9), (5.5,9.9), (23.1,9.9), (-23.1,7.7), (-9.9,7.7), (-3.3,7.7), (1.1,7.7), (7.7,7.7), (25.3,7.7), (-23.1,5.5), (18.7,5.5), (20.9,5.5), (25.3,5.5), (-23.1,3.3), (-14.3,3.3), (-12.1,3.3), (14.3,3.3), (18.7,3.3), (20.9,3.3), (27.5,3.3), (-23.1,1.1), (-14.3,1.1), (-12.1,1.1), (14.3,1.1), (18.7,1.1), (20.9,1.1), (23.1,1.1), (27.5,1.1), (-23.1,-1.1), (12.1,-1.1), (18.7,-1.1), (20.9,-1.1), (23.1,-1.1), (27.5,-1.1), (-23.1,-3.3), (12.1,-3.3), (16.5,-3.3), (18.7,-3.3), (20.9,-3.3), (23.1,-3.3), (27.5,-3.3), (-20.9,-5.5), (5.5,-5.5), (12.1,-5.5), (16.5,-5.5), (18.7,-5.5), (20.9,-5.5), (23.1,-5.5), (27.5,-5.5), (3.3,-7.7), (12.1,-7.7), (16.5,-7.7), (18.7,-7.7), (20.9,-7.7), (23.1,-7.7), (27.5,-7.7), (-16.5,-9.9), (1.1,-9.9), (12.1,-9.9), (16.5,-9.9), (18.7,-9.9), (20.9,-9.9), (27.5,-9.9), (-12.1,-12.1), (9.9,-12.1), (12.1,-12.1), (25.3,-12.1), (-9.9,-14.3), (-7.7,-14.3), (7.7,-14.3), (12.1,-14.3), (14.3,-14.3), (20.9,-14.3), (23.1,-14.3), (-5.5,-16.5), (-3.3,-16.5), (5.5,-16.5), (14.3,-16.5), (16.5,-16.5), (18.7,-16.5), (-3.3,-18.7), (-1.1,-18.7), (1.1,-18.7), (3.3,-18.7), (5.5,-18.7), (-5.5,-20.9), (-3.3,-23.1), (-1.1,-23.1), (1.1,-23.1), (5.5,-23.1), (7.7,-23.1)]
sfgfp_points = [(14.3,20.9), (16.5,20.9), (-14.3,12.1), (-18.7,9.9), (-18.7,-7.7)]
mturquoise2_points = [(-16.5,9.9), (-16.5,3.3), (-16.5,1.1), (-7.7,-7.7), (-5.5,-9.9), (-3.3,-9.9), (-1.1,-9.9)]
point_name_pairing = [("venus", venus_points),("mjuniper", mjuniper_points),("sfgfp", sfgfp_points),("mturquoise2", mturquoise2_points)]
# Robot deck setup constants
TIP_RACK_DECK_SLOT = 9
COLORS_DECK_SLOT = 6
AGAR_DECK_SLOT = 5
PIPETTE_STARTING_TIP_WELL = 'A1'
# Place the PCR tubes in this order
well_colors = {
'A1': 'sfGFP',
'A2': 'mRFP1',
'A3': 'mKO2',
'A4': 'Venus',
'A5': 'mKate2_TF',
'A6': 'Azurite',
'A7': 'mCerulean3',
'A8': 'mClover3',
'A9': 'mJuniper',
'A10': 'mTurquoise2',
'A11': 'mBanana',
'A12': 'mPlum',
'B1': 'Electra2',
'B2': 'mWasabi',
'B3': 'mScarlet_I',
'B4': 'mPapaya',
'B5': 'eqFP578',
'B6': 'tdTomato',
'B7': 'DsRed',
'B8': 'mKate2',
'B9': 'EGFP',
'B10': 'mRuby2',
'B11': 'TagBFP',
'B12': 'mChartreuse_TF',
'C1': 'mLychee_TF',
'C2': 'mTagBFP2',
'C3': 'mEGFP',
'C4': 'mNeonGreen',
'C5': 'mAzamiGreen',
'C6': 'mWatermelon',
'C7': 'avGFP',
'C8': 'mCitrine',
'C9': 'mVenus',
'C10': 'mCherry',
'C11': 'mHoneydew',
'C12': 'TagRFP',
'D1': 'mTFP1',
'D2': 'Ultramarine',
'D3': 'ZsGreen1',
'D4': 'mMiCy',
'D5': 'mStayGold2',
'D6': 'PA_GFP'
}
volume_used = {
'venus': 0,
'mjuniper': 0,
'sfgfp': 0,
'mturquoise2': 0
}
def update_volume_remaining(current_color, quantity_to_aspirate):
rows = string.ascii_uppercase
for well, color in list(well_colors.items()):
if color == current_color:
if (volume_used[current_color] + quantity_to_aspirate) > 250:
# Move to next well horizontally by advancing row letter, keeping column number
row = well[0]
col = well[1:]
# Find next row letter
next_row = rows[rows.index(row) + 1]
next_well = f"{next_row}{col}"
del well_colors[well]
well_colors[next_well] = current_color
volume_used[current_color] = quantity_to_aspirate
else:
volume_used[current_color] += quantity_to_aspirate
break
def run(protocol):
# Load labware, modules and pipettes
protocol.home()
# Tips
tips_20ul = protocol.load_labware('opentrons_96_tiprack_20ul', TIP_RACK_DECK_SLOT, 'Opentrons 20uL Tips')
# Pipettes
pipette_20ul = protocol.load_instrument("p20_single_gen2", "right", [tips_20ul])
# PCR Plate
temperature_plate = protocol.load_labware('opentrons_96_aluminumblock_generic_pcr_strip_200ul', 6)
# Agar Plate
agar_plate = protocol.load_labware('htgaa_agar_plate', AGAR_DECK_SLOT, 'Agar Plate')
agar_plate.set_offset(x=0.00, y=0.00, z=Z_VALUE_AGAR)
# Get the top-center of the plate, make sure the plate was calibrated before running this
center_location = agar_plate['A1'].top()
pipette_20ul.starting_tip = tips_20ul.well(PIPETTE_STARTING_TIP_WELL)
# Helper function (dispensing)
def dispense_and_jog(pipette, volume, location):
assert(isinstance(volume, (int, float)))
# Go above the location
above_location = location.move(types.Point(z=location.point.z + 2))
pipette.move_to(above_location)
# Go downwards and dispense
pipette.dispense(volume, location)
# Go upwards to avoid smearing
pipette.move_to(above_location)
# Helper function (color location)
def location_of_color(color_string):
for well,color in well_colors.items():
if color.lower() == color_string.lower():
return temperature_plate[well]
raise ValueError(f"No well found with color {color_string}")
# Print pattern by iterating over lists
for i, (current_color, point_list) in enumerate(point_name_pairing):
# Skip the rest of the loop if the list is empty
if not point_list:
continue
# Get the tip for this run, set the bacteria color, and the aspirate bacteria of choice
pipette_20ul.pick_up_tip()
max_aspirate = int(18 // POINT_SIZE) * POINT_SIZE
quantity_to_aspirate = min(len(point_list)*POINT_SIZE, max_aspirate)
update_volume_remaining(current_color, quantity_to_aspirate)
pipette_20ul.aspirate(quantity_to_aspirate, location_of_color(current_color))
# Iterate over the current points list and dispense them, refilling along the way
for i in range(len(point_list)):
x, y = point_list[i]
adjusted_location = center_location.move(types.Point(x, y))
dispense_and_jog(pipette_20ul, POINT_SIZE, adjusted_location)
if pipette_20ul.current_volume == 0 and len(point_list[i+1:]) > 0:
quantity_to_aspirate = min(len(point_list[i:])*POINT_SIZE, max_aspirate)
update_volume_remaining(current_color, quantity_to_aspirate)
pipette_20ul.aspirate(quantity_to_aspirate, location_of_color(current_color))
# Drop tip between each color
pipette_20ul.drop_tip()
Post-Lab Questions
- Use of automation in final project
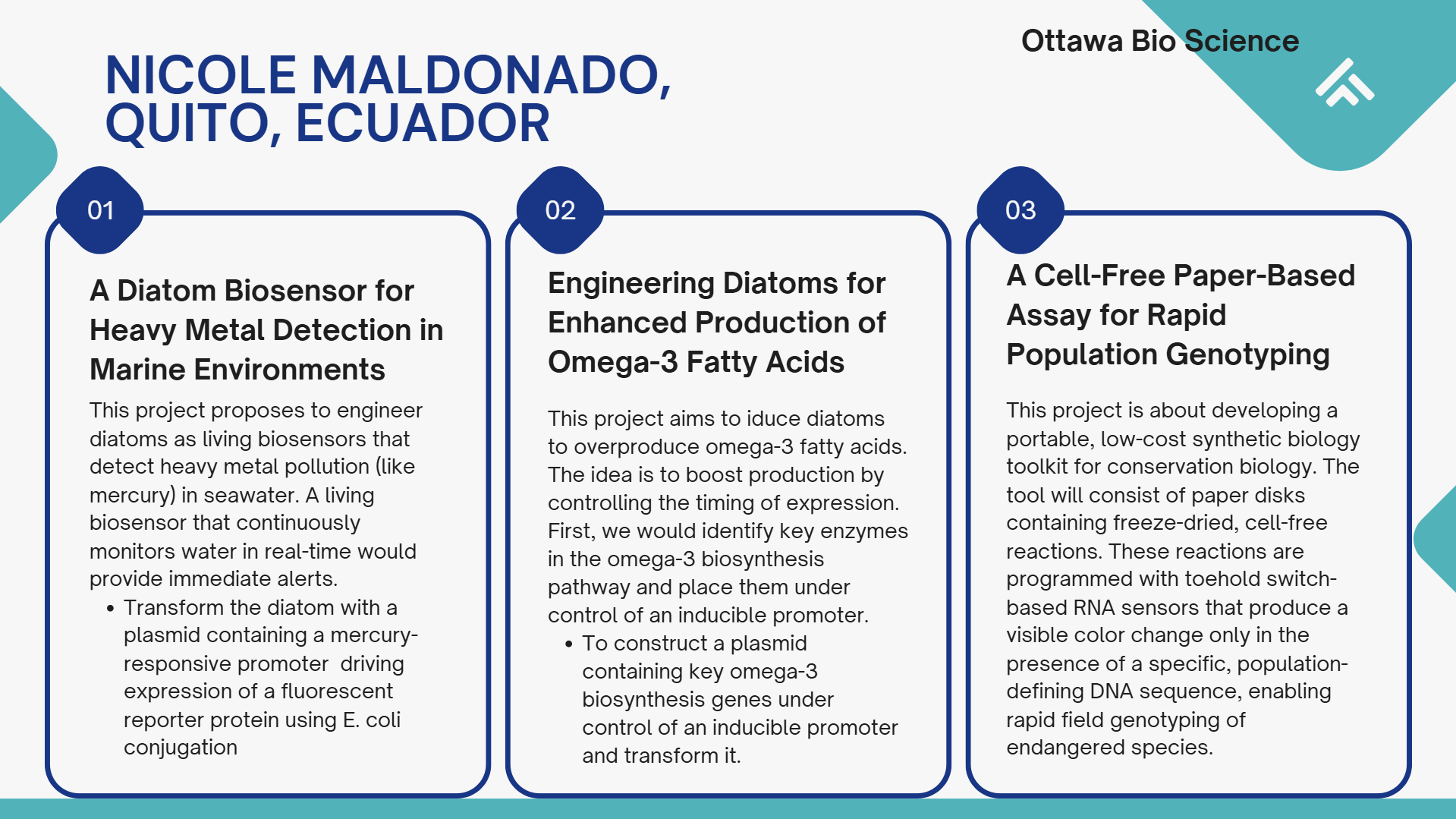
Project: A Cell-Free Paper-Based Assay for Rapid Population Genotyping
This project will utilize cloud laboratory automation to streamline the design and optimization of toehold switch RNA sensors. The core idea is to move from a manual, low-throughput testing process to an automated, high-throughput workflow that can rapidly evaluate hundreds of sensor variants against multiple target conditions. This approach will accelerate the identification of highly specific sensors that can distinguish between populations based on single nucleotide polymorphisms (SNPs).
The automated workflow will systematically test sensor variants against target and non-target sequences to identify the most specific and sensitive candidates for field deployment. By automating the iterative design-build-test cycle, we can rapidly screen hundreds of sensor designs that would be impractical to test manually.
Use Ginkgo’s automated DNA design tools to generate a library of toehold switch variants targeting a population-specific SNP in humpback whale mitochondrial DNA
Each variant will incorporate sequence variations around the SNP to optimize discrimination
Expected Outcomes:
The automated workflow will deliver a validated set of 3-5 highly specific toehold sensors ready for field deployment. Ultimately we would be making advanced genetic analysis accessible for conservation applications in remote locations.
- Summary of paper
Paper DOI: 10.1016/j.mcpro.2024.100790
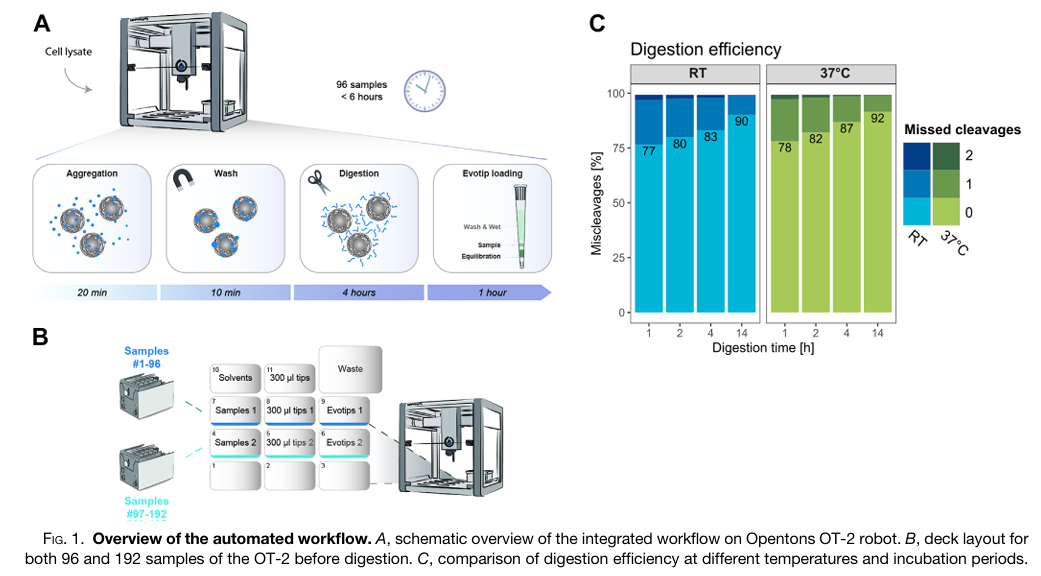
Fully Automated Workflow for Integrated Sample Digestion and Evotip Loading Enabling High-Throughput Clinical Proteomics
The study aimed to address a major bottleneck in clinical proteomics, meaning the manual, time-consuming, and often variable preparation of samples for mass spectrometry analysis. The authors sought to develop and validate a fully automated, high-throughput workflow using the affordable and flexible Opentrons OT-2 liquid handling robot. Their primary goal was to create an end-to-end process that could handle hundreds of samples simultaneously, from protein digestion to final loading onto Evotips for LC-MS/MS analysis, with minimal human intervention. They also aimed to expand this automated workflow to include phosphopeptide enrichment, enabling large-scale studies of cellular signaling.
To achieve this, the researchers designed a protocol on the OT-2 that integrates magnetic bead-based protein aggregation capture (PAC) digestion with direct loading of the resulting peptides onto Evotips. This method replaces traditional centrifugation steps with positive pressure, allowing the robot to process up to 192 samples in parallel within approximately six hours. They rigorously tested the system’s performance using HeLa cell lysates and human plasma samples, analyzing the results with high-speed Orbitrap mass spectrometers. Furthermore, they adapted the protocol to incorporate an automated step for enriching phosphorylated peptides using magnetic Ti-IMAC beads, demonstrating its utility in a time-course study of cellular response to drug treatment.
The automated workflow proved to be highly robust and reproducible, consistently identifying approximately 8,000 protein groups and 130,000 peptides from just 1 µg of HeLa protein digest across 192 samples. In a pilot clinical study with plasma samples from 48 metastatic melanoma patients, the robot-enabled preparation allowed for the quantification of over 600 protein groups, revealing potential biomarker candidates that differentiated responders from non-responders to immune therapy. The integrated phosphoproteomics workflow also successfully identified over 8,000 phosphorylation sites from as few as 40,000 cells and accurately mapped dynamic signaling changes in response to anisomycin treatment. The use of the Opentrons OT-2 robot was central to this research, dramatically increasing throughput and reproducibility while reducing reagent costs and hands-on time, thereby demonstrating its potential to unlock large-scale, clinically-relevant proteomic and phosphoproteomic studies.

Final Project ideas