Week 2 HW: DNA Read, Write and Edit
Part 0: Attend or watch all lecture and recitation videos.
Part 1: Benchling & In-silico Gel Art
- Make a free account at benchling.com
- Import the Lambda DNA.
- Simulate Restriction Enzyme Digestion with the following Enzymes:
- EcoRI
- HindIII
- BamHI
- KpnI
- EcoRV
- SacI
- SalI
- Create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks.
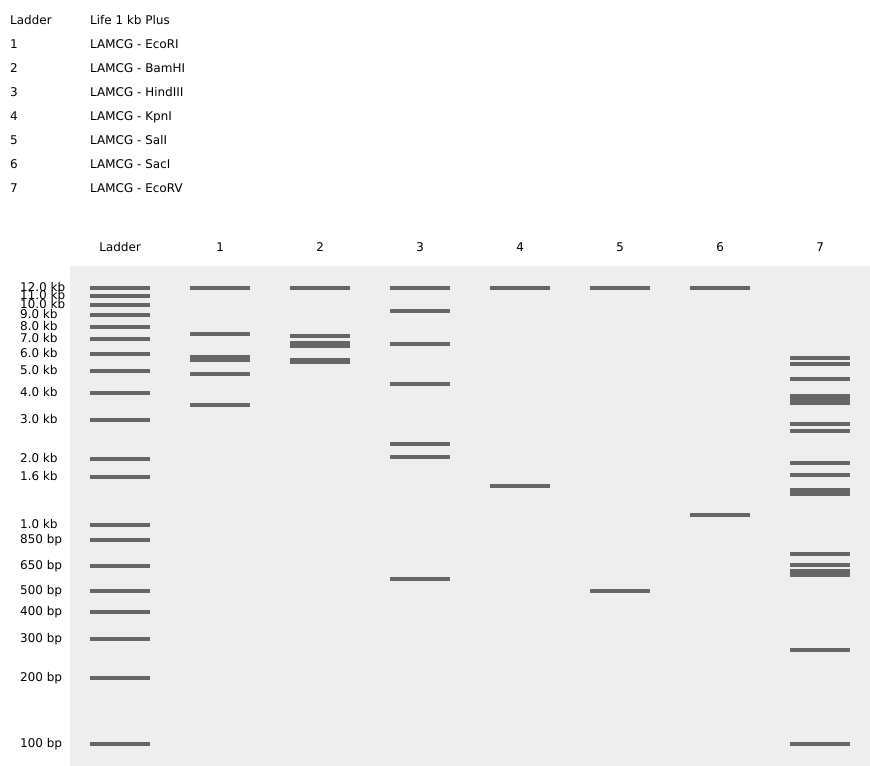




- I imagine the pattern as a hand making number one
Part 3: DNA Design Challenge
- Choose your protein.
- I chose tau protein that it’s hyperphosphorylation is involved in Alzheimer’s disease progression
- I chose UniProt to get its sequence

Reference: https://rest.uniprot.org/uniprotkb/P10636.fasta
- Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
- Tau Protein DNA Sequence:
atggcggaaccgcgccaggaatttgaagtgatggaagatcatgcgggcacctatggcctg ggcgatcgcaaagatcagggcggctataccatgcatcaggatcaggaaggcgataccgat gcgggcctgaaagaaagcccgctgcagaccccgaccgaagatggcagcgaagaaccgggc agcgaaaccagcgatgcgaaaagcaccccgaccgcggaagatgtgaccgcgccgctggtg gatgaaggcgcgccgggcaaacaggcggcggcgcagccgcataccgaaattccggaaggc accaccgcggaagaagcgggcattggcgataccccgagcctggaagatgaagcggcgggc catgtgacccaggaaccggaaagcggcaaagtggtgcaggaaggctttctgcgcgaaccg ggcccgccgggcctgagccatcagctgatgagcggcatgccgggcgcgccgctgctgccg gaaggcccgcgcgaagcgacccgccagccgagcggcaccggcccggaagataccgaaggc ggccgccatgcgccggaactgctgaaacatcagctgctgggcgatctgcatcaggaaggc ccgccgctgaaaggcgcgggcggcaaagaacgcccgggcagcaaagaagaagtggatgaa gatcgcgatgtggatgaaagcagcccgcaggatagcccgccgagcaaagcgagcccggcg caggatggccgcccgccgcagaccgcggcgcgcgaagcgaccagcattccgggctttccg gcggaaggcgcgattccgctgccggtggattttctgagcaaagtgagcaccgaaattccg gcgagcgaaccggatggcccgagcgtgggccgcgcgaaaggccaggatgcgccgctggaa tttacctttcatgtggaaattaccccgaacgtgcagaaagaacaggcgcatagcgaagaa catctgggccgcgcggcgtttccgggcgcgccgggcgaaggcccggaagcgcgcggcccg agcctgggcgaagataccaaagaagcggatctgccggaaccgagcgaaaaacagccggcg gcggcgccgcgcggcaaaccggtgagccgcgtgccgcagctgaaagcgcgcatggtgagc aaaagcaaagatggcaccggcagcgatgataaaaaagcgaaaaccagcacccgcagcagc gcgaaaaccctgaaaaaccgcccgtgcctgagcccgaaacatccgaccccgggcagcagc gatccgctgattcagccgagcagcccggcggtgtgcccggaaccgccgagcagcccgaaa tatgtgagcagcgtgaccagccgcaccggcagcagcggcgcgaaagaaatgaaactgaaa ggcgcggatggcaaaaccaaaattgcgaccccgcgcggcgcggcgccgccgggccagaaa ggccaggcgaacgcgacccgcattccggcgaaaaccccgccggcgccgaaaaccccgccg agcagcggcgaaccgccgaaaagcggcgatcgcagcggctatagcagcccgggcagcccg ggcaccccgggcagccgcagccgcaccccgagcctgccgaccccgccgacccgcgaaccg aaaaaagtggcggtggtgcgcaccccgccgaaaagcccgagcagcgcgaaaagccgcctg cagaccgcgccggtgccgatgccggatctgaaaaacgtgaaaagcaaaattggcagcacc gaaaacctgaaacatcagccgggcggcggcaaagtgcagattattaacaaaaaactggat ctgagcaacgtgcagagcaaatgcggcagcaaagataacattaaacatgtgccgggcggc ggcagcgtgcagattgtgtataaaccggtggatctgagcaaagtgaccagcaaatgcggc agcctgggcaacattcatcataaaccgggcggcggccaggtggaagtgaaaagcgaaaaa ctggattttaaagatcgcgtgcagagcaaaattggcagcctggataacattacccatgtg ccgggcggcggcaacaaaaaaattgaaacccataaactgacctttcgcgaaaacgcgaaa gcgaaaaccgatcatggcgcggaaattgtgtataaaagcccggtggtgagcggcgatacc agcccgcgccatctgagcaacgtgagcagcaccggcagcattgatatggtggatagcccg cagctggcgaccctggcggatgaagtgagcgcgagcctggcgaaacagggcctg
Reference https://www.bioinformatics.org/sms2/rev_trans.html
- Codon optimization.
- Codon optimization is conducted to increase the efficiency of expression. For example, although each amino acid has more than one codon, their efficiency varies, therefore, the optimization aims to choose the most efficient codons for increased translation efficiency and stable mRNA structure.
- I chose E.coli to optimize the codon for, because it of its fast replication and versatile applications
- Tau Protein DNA sequence with Codon-Optimization
- Tau protein optimized coding sequence
GC=59.89%, CAI=0.90
ATGGCGGAACCGCGCCAGGAGTTCGAAGTGATGGAAGATCATGCGGGCACCTATGGCCTGGGCGATCGTAAAGATCAGGGCGGCTACACGATGCATCAGGATCAGGAAGGCGATACCGATGCAGGCCTGAAAGAAAGCCCGCTGCAGACCCCGACCGAAGATGGTAGCGAAGAACCGGGCAGCGAAACCAGCGATGCGAAAAGCACCCCGACCGCCGAAGATGTTACCGCCCCTTTAGTGGATGAAGGCGCGCCGGGCAAACAGGCGGCGGCCCAGCCGCATACCGAAATTCCGGAAGGCACGACCGCGGAAGAAGCGGGCATTGGCGATACCCCGAGCCTGGAAGATGAAGCAGCGGGTCACGTGACCCAGGAACCGGAAAGCGGCAAAGTTGTGCAGGAAGGCTTTCTGCGCGAGCCGGGACCGCCCGGCCTGAGCCATCAACTGATGAGCGGCATGCCGGGTGCGCCGTTACTGCCGGAAGGCCCGCGCGAAGCCACCCGCCAGCCGAGCGGCACGGGCCCGGAAGATACCGAAGGCGGCCGTCATGCGCCGGAACTGCTGAAACATCAGCTGCTGGGCGATCTGCATCAGGAAGGCCCGCCGCTGAAAGGCGCGGGTGGCAAAGAACGTCCGGGCAGCAAAGAAGAAGTGGATGAAGATCGTGATGTGGATGAAAGCAGCCCGCAGGATAGCCCGCCGAGCAAAGCCAGCCCGGCCCAGGATGGCCGTCCGCCGCAAACCGCGGCACGTGAAGCCACCTCAATTCCGGGCTTCCCGGCGGAAGGCGCGATTCCGCTGCCGGTGGATTTCCTGAGCAAAGTGAGCACCGAAATTCCGGCGAGCGAACCGGATGGCCCGAGCGTGGGTCGCGCCAAAGGCCAGGATGCGCCGCTGGAATTCACCTTTCATGTGGAAATTACCCCGAACGTGCAGAAAGAACAGGCGCATAGCGAAGAGCATCTGGGACGCGCGGCCTTTCCGGGCGCGCCGGGTGAAGGTCCGGAAGCGCGCGGTCCGTCTCTGGGCGAAGATACGAAAGAAGCGGATCTGCCGGAACCGAGCGAAAAACAGCCGGCGGCGGCGCCGCGCGGTAAACCGGTGAGCCGCGTTCCGCAACTGAAAGCGCGCATGGTTTCGAAATCAAAAGATGGCACGGGCAGCGACGATAAAAAAGCCAAAACCAGCACCCGCAGCAGTGCCAAAACCCTGAAAAACCGCCCGTGCCTGAGCCCGAAACATCCGACGCCGGGCAGCAGCGATCCGCTGATTCAGCCGAGCTCTCCGGCGGTTTGTCCTGAACCGCCGTCAAGTCCGAAATATGTTAGCAGCGTTACCAGCCGCACCGGCTCAAGCGGCGCCAAAGAAATGAAACTGAAAGGTGCCGATGGTAAAACTAAAATTGCGACCCCGCGCGGCGCGGCCCCGCCGGGCCAGAAAGGCCAGGCGAACGCAACCCGCATTCCGGCGAAAACCCCGCCGGCGCCGAAAACCCCGCCGAGTTCAGGTGAACCGCCGAAAAGCGGCGATCGCTCAGGCTATAGTAGCCCGGGCAGCCCGGGGACCCCGGGCAGCCGTTCACGTACCCCGAGCCTGCCGACCCCGCCGACTCGTGAACCGAAAAAAGTCGCCGTGGTACGCACCCCGCCGAAAAGCCCGTCGTCGGCGAAAAGCCGCCTGCAGACCGCGCCGGTTCCGATGCCGGATCTGAAAAATGTGAAAAGCAAAATTGGCTCTACCGAAAACCTGAAACACCAGCCGGGAGGCGGCAAAGTGCAAATCATTAATAAAAAACTGGATCTGTCAAACGTGCAATCAAAATGCGGTTCGAAAGATAACATTAAACATGTTCCGGGTGGCGGCTCGGTGCAGATTGTGTATAAACCCGTGGATCTGAGCAAAGTTACCTCGAAGTGTGGATCTCTGGGCAATATCCATCATAAACCGGGCGGCGGCCAGGTTGAAGTTAAATCTGAAAAACTGGATTTTAAAGATCGCGTGCAGAGCAAAATTGGCAGCCTGGATAATATCACCCATGTGCCGGGCGGCGGCAACAAAAAAATTGAAACCCATAAACTGACCTTTCGCGAAAATGCCAAAGCGAAAACCGATCACGGTGCGGAAATTGTTTATAAAAGCCCGGTTGTTAGCGGTGATACGAGCCCGCGTCATCTGTCGAACGTTAGCTCAACCGGTAGCATTGATATGGTGGATAGCCCGCAACTGGCGACGCTGGCCGATGAAGTGTCGGCGTCCCTGGCGAAACAGGGTCTG
Reference: https://en.vectorbuilder.com/tool/codon-optimization/6781bd12-7b93-4071-a97a-e6f9b8f17287.html
Part 4: Prepare a Twist DNA Synthesis Order
- Create a Twist account and a Benchling account [DONE]
- Build Your DNA Insert Sequence
Sharing link in Benchling: https://benchling.com/s/seq-LWrWyNWwQivMCCt0o4Ra?m=slm-qbMaWSkeMRm5NZ7JzEJw
- SBOL Image

after uploading the sequence in twist, I optimized it more in twist
sharing link after exporting to benchling https://benchling.com/s/seq-wNOy6J9WVuWzZ1i4URkY?m=slm-Gj8tRUDvFaVLFxjZZbbn
image of the constructed plasmid
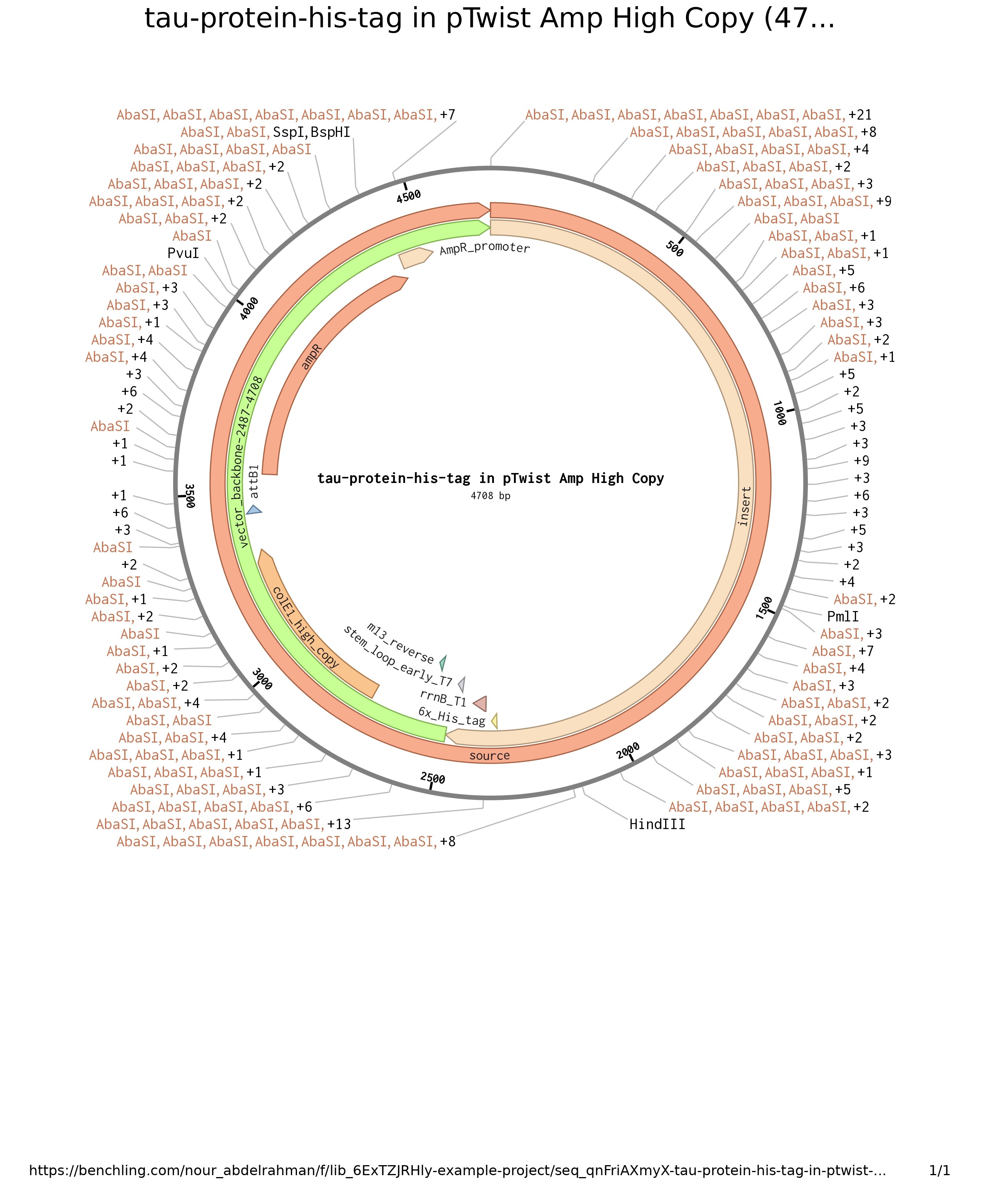

Part 5: DNA Read/Write/Edit
DNA Read
- What DNA would you want to sequence (e.g., read) and why?
- The gene I’m interested in is the APP (Amyloid Beta Precursor Protein) Gene, this gene is involved in Alzheimer disease. I chose this gene because I’m interested in using synthetic biology to understand neurodegenerative disorders, especially Alzheimer’s disease.
- In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
- I chose PacBio sequencing technology, it is a third-generation sequencing technology, that have the ability to produce long and higly accurate DNA reads. It is based on single molecule real-time (SMRT) sequencing principle.
- for preparing the input, the DNA is prepared into the SMRTbell library by ligating hairpin adapters to double-stranded DNA on both ends, forming a circular template. Primers and polymerases are added to this library, which is loaded onto the sequencing instrument that contains the SMRT Cell and ZMWs. A single template DNA is immobilized in each ZMW. As the polymerase adds fluorescently labeled nucleotides into the growing DNA strand, light is emitted. This light emission is measured in real time and these signals are converted into nucleotide sequences.
- the first step is library costruction which involves several steps to prepare DNA for sequencing:
- DNA is cleaved into fragments of the desired size and it undergoes end repair.
- Then, adaptors with hairpin structures are ligated to both ends of the DNA fragments which creates single-stranded circular structures called SMRTbell templates.
- Finally, the templates are purified and loaded onto the PacBio sequencing instrument.
- the output is fluorescent signals that are translated into base sequences then alignment and assembly are conducted
DNA Write
- What DNA would you want to synthesize (e.g., write) and why?
- I would synthesize genetic circuit that sense the presence of high amount of hyperphosphorylation in the brain for example.
- What technology or technologies would you use to perform this DNA synthesis and why?
- I would choose oxford nanopore for synthesizing the genetic circuit.
DNA Edit
- What DNA would you want to edit and why?
- I would want to edit a gene that have a disease-causing mutation.
- What technology or technologies would you use to perform these DNA edits and why?
- I would use CRISPR-Cas to edit the gene, the main steps involves designing gRNA that will guide the Cas9 to cut the specific site