Week 4 HW: Protein Design Pt. 1

Part A: Conceptual Questions
How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons)
Why do humans eat beef but do not become a cow, eat fish but do not become fish?
Why are there only 20 natural amino acids? Ref: https://www.chemistryworld.com/features/why-are-there-20-amino-acids/3009378.article
Can you make other non-natural amino acids? Design some new amino acids.
Where did amino acids come from before enzymes that make them, and before life started?
If you make an α-helix using D-amino acids, what handedness (right or left) would you expect?
Can you discover additional helices in proteins?
Why are most molecular helices right-handed?
Why do β-sheets tend to aggregate? What is the driving force for β-sheet aggregation?
Why do many amyloid diseases form β-sheets? Can you use amyloid β-sheets as materials?
Design a β-sheet motif that forms a well-ordered structure.
Part B: Protein Analysis and Visualization
- Briefly describe the protein you selected and why you selected it.
I select tau protein. It is a microtubule-associated protein that promotes microtubule assembly and stability, and might be involved in the establishment and maintenance of neuronal polarity. In neurodegeneration, this protein becomes hyperphosphorylated, detaches from microtubules, and aggregates into toxic, insoluble neurofibrillary tangles (NFTs). Since I’m interested in using synthetic biology to understand more neurodegenerative disorders, this protein is of interest.
Another protein is Amyloid-beta precursor protein.
It may play a role in postsynaptic function. The C-terminal gamma-secretase processed fragment, ALID1, activates transcription activation through APBB1 (Fe65) binding. Couples to JIP signal transduction through C-terminal binding. May interact with cellular G-protein signaling pathways. Can regulate neurite outgrowth through binding to components of the extracellular matrix such as heparin and collagen I. The gamma-CTF peptide, C30, is a potent enhancer of neuronal apoptosis.
Identify the amino acid sequence of your protein.
Tau Protein
Sequence Length: 758 amino acids
The most common amino acid is: Proline, which appears 93 times.
Amino Acid Frequencies: P: 93 (12.27%) G: 82 (10.82%) S: 79 (10.42%) K: 64 (8.44%) A: 60 (7.92%) E: 59 (7.78%) T: 50 (6.60%) D: 43 (5.67%) L: 43 (5.67%) V: 41 (5.41%) Q: 33 (4.35%) R: 30 (3.96%) H: 20 (2.64%) I: 20 (2.64%) N: 13 (1.72%) M: 9 (1.19%) F: 9 (1.19%) Y: 6 (0.79%) C: 4 (0.53%)
there are 10 protein homologs
yes, it belongs to the microtubule-associated protein family
APP
Sequence length: 650 Amino Acids (fasta file: https://rest.uniprot.org/uniprotkb/P51693.fasta)
The most common amino acid is L appearing 70 times
Amino Acid Frequencies: L: 70 E: 67 P: 61 R: 57 G: 51 A: 51 Q: 46 S: 43 V: 36 D: 26 T: 23 H: 22 M: 16 I: 16 F: 14 K: 14 C: 12 Y: 11 N: 10 W: 4
250 homolgs (reference: https://www.uniprot.org/blast/uniprotkb/ncbiblast-R20260307-104740-0382-64643955-p1m/overview)
it belongs to the APP family.
- Identify the structure page of your protein in RCSB
Tau Protein
- the structure was solved/released in 2015-07-08
- it is a high-quality NMR structure
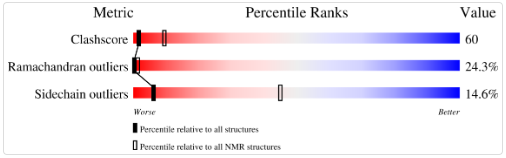

- No, it doesn’t belong to any structure classification family
APP Protein
- Good resoltuion. Resolution: 2.60 Å
References: https://www.uniprot.org/uniprotkb/P10636/entry https://www.ebi.ac.uk/pdbe/scop/search?t=txt;q=tau%20protein
- Open the structure of your protein in any 3D molecule visualization software
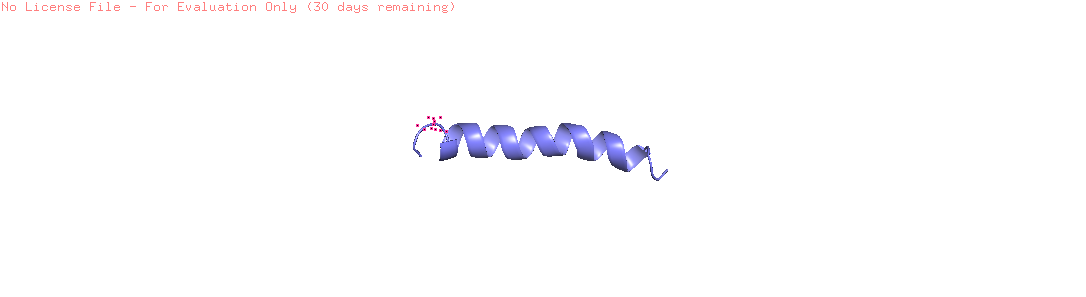
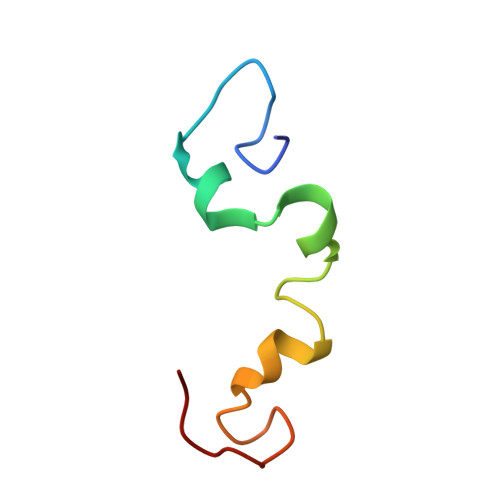
Visualize the protein as “cartoon”, “ribbon” and “ball and stick”
the protein that will be visualized is APP (Amyloid-beta precursor protein)
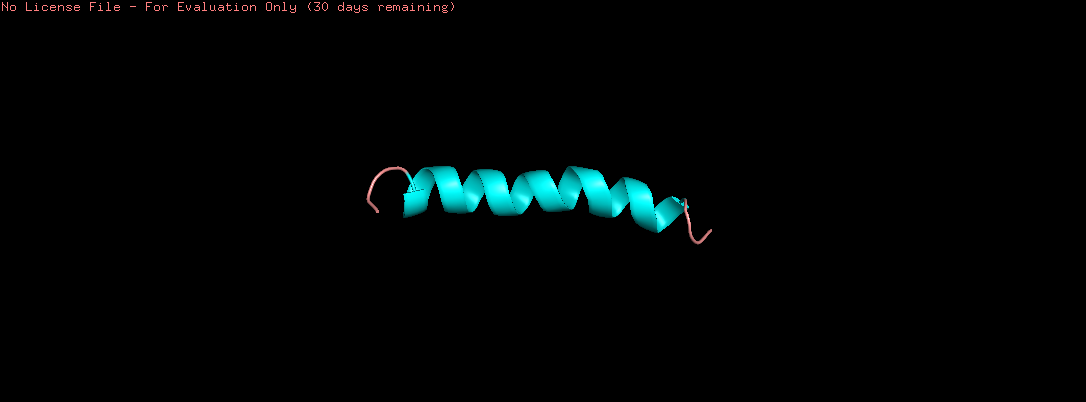
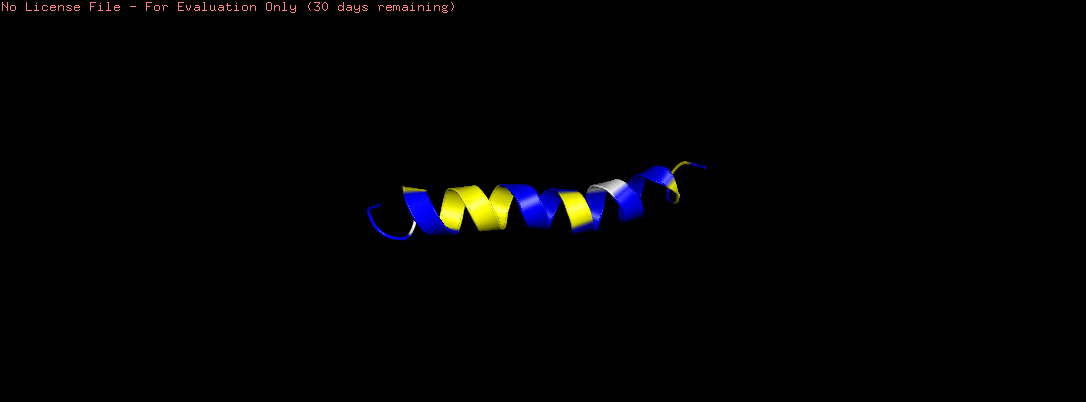
Color the protein by secondary structure. Does it have more helices or sheets?
The helix is colored in cyan and the sheet in magenta, from the image the helices are more
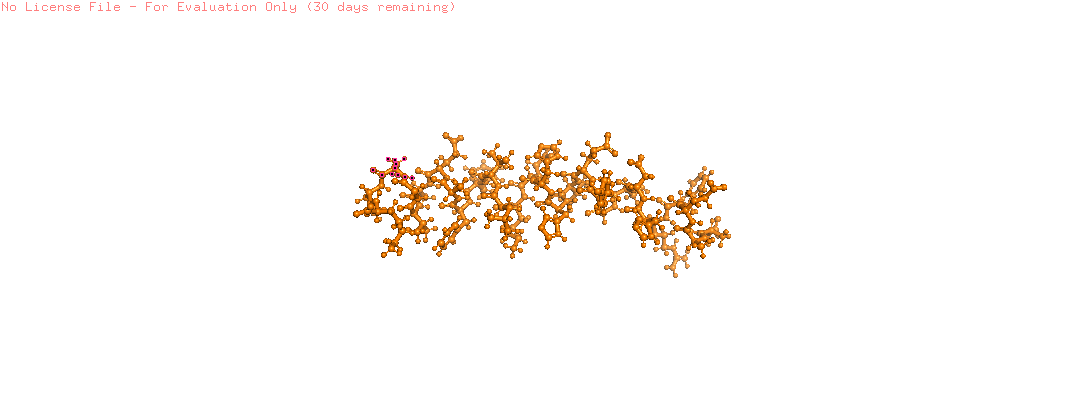
- Color the protein by residue type. What can you tell about the distribution of hydrophobic vs hydrophilic residues?
- The hydrophobic residues (ALA, VAL, LEU, ILE, MET, PHE, TRP, PRO) are colored in yellow
- The hydrophilic residues (SER, THR, ASN, GLN, TYR, CYS, LYS, ARG, HIS, ASP, GLU) are colored in blue
- GLY (neutral) is colored in white
- From the image the hydrophilic residues are grouped together and they are more then the hydrophobic ones
- Visualize the surface of the protein. Does it have any “holes” (aka binding pockets)? No, it doesn’t have any holes
Part C: Using ML-Based Protein Design Tools
I chose the Amyloid Beta-Peptide protein
C1. Protein Language Modeling
Deep Mutational Scans
- Use ESM2 to generate an unsupervised deep mutational scan of your protein based on language model likelihoods.
- Can you explain any particular pattern? (choose a residue and a mutation that stands out)
Fasta File
1AMC_1|Chain A|AMYLOID BETA-PEPTIDE|Homo sapiens (9606) old
DAEFRHDSGYEVHHQKLVFFAEDVGSNK old
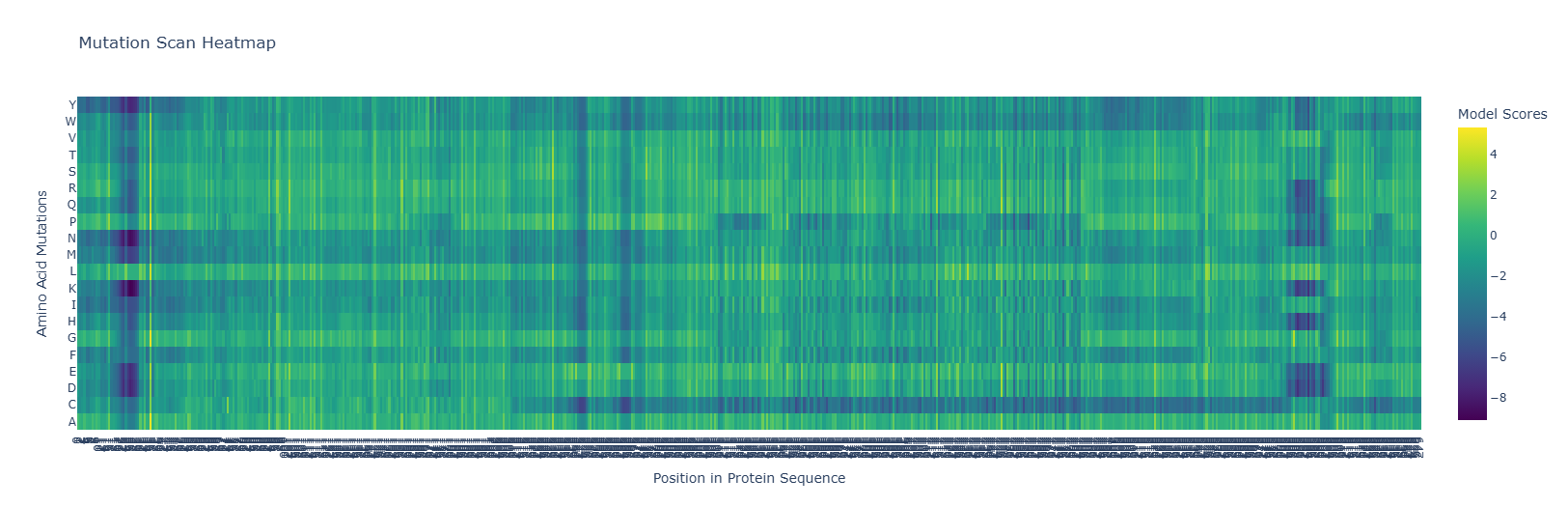

The heatmap shows hotspots for mutations that are beneficial or detrimental to the function of the protein. Notice the dark blue regions, where the LLR values are negative, indicating that mutations that are likely detrimental to function, and lighter yellow regions where the LLR values are positive, indicating mutations that are likely beneficial to the function of the protein. Also, note how there are dark bands running vertically indicating regions which are likely evolutionarily conserved, and brighter bands running vertically indicating regions of the protein which may in fact be preferable over the wild-type sequence. Note also, for some regions of the protein, there are amino acid mutations which are likely to be detrimental to functioning for entire regions of the protein, indicated by dark bands running horizontally along most of the protein. Similarly, we see brighter bands of yellow running horizontally, indicating almost any residue mutated to that amino acid would be preferential to the wild type.
for example replacing by is prefereable. mutations at as well are favourable
Latent Space Analysis
- Use the provided sequence dataset to embed proteins in reduced dimensionality.
- Analyze the different formed neighborhoods: do they approximate similar proteins?
- Place your protein in the resulting map and explain its position and similarity to its neighbors.
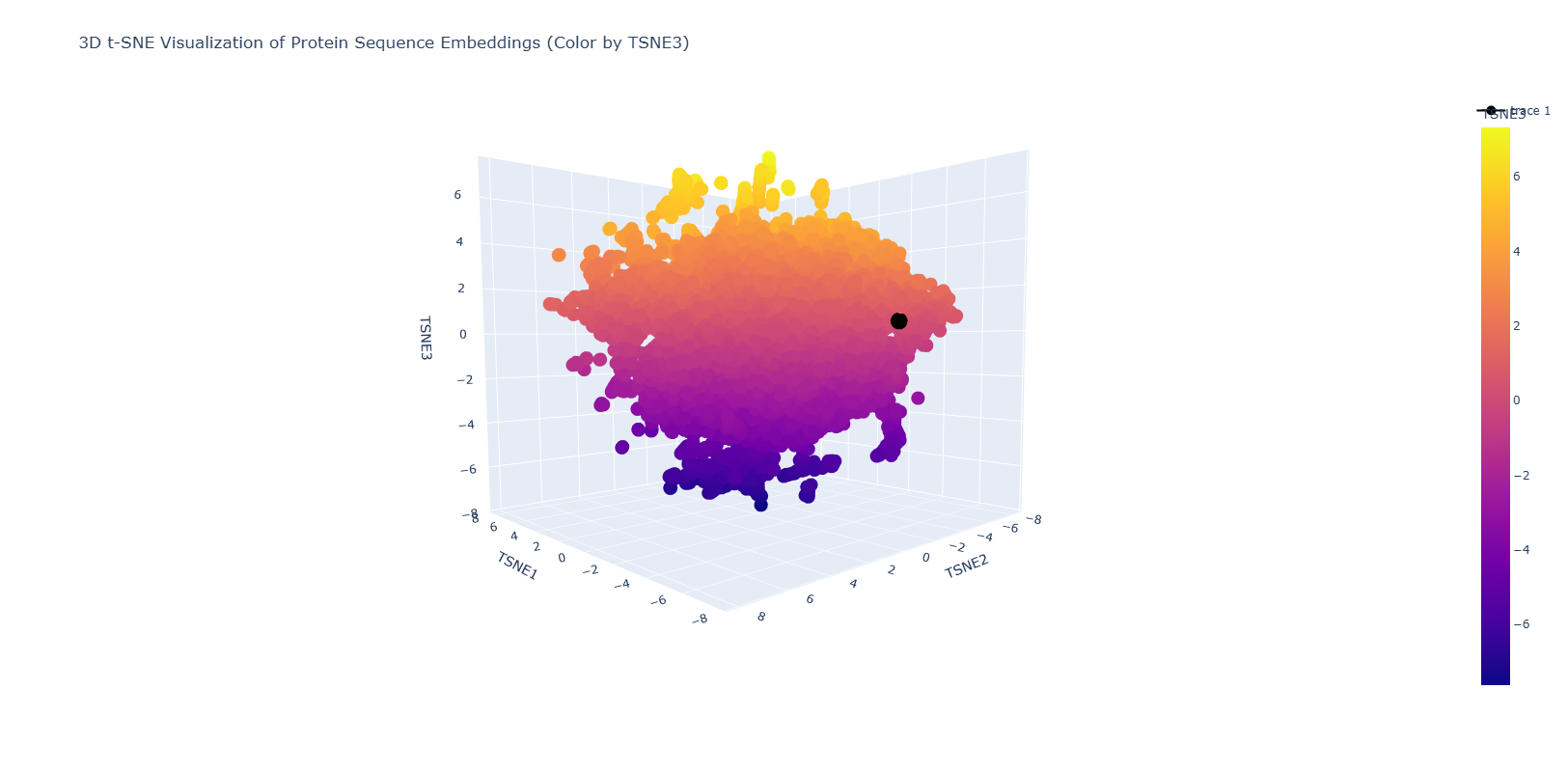

C2. Protein Folding
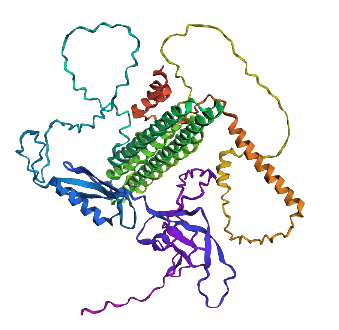
- Fold your protein with ESMFold. Do the predicted coordinates match your original structure?
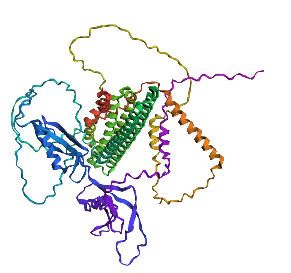

No, they don’t match it
- Try changing the sequence, first try some mutations, then large segments. Is your protein structure resilient to mutations? Mutation done: replacing position 22 (L) by K
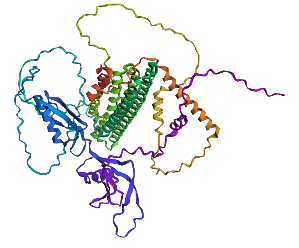

Mutation: replacing positions 20 to 29 LPLLLPLLLL with NNNNNNNNNN

C3. Protein Generation
- Analyze the predicted sequence probabilities and compare the predicted sequence vs the original one.
- Input this sequence into ESMFold and compare the predicted structure to your original.
Part D: Group Brainstorm on Bacteriophage Engineering
[x] Find a group of ~3–4 students [ ] Read through the Phage Reading material listed under “Reading & Resources” below.
References colab link: https://colab.research.google.com/drive/16VrQUyOY0s-a7m07FFV2H-4UhvRD3eje?authuser=2#scrollTo=ySOWXRjTja9D https://huggingface.co/blog/AmelieSchreiber/mutation-scoring