Week 6 HW: Genetic Circuits Pt.1
DNA Assembly
- What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
The components of the Phusion High-Fidelity PCR Master Mix are the following:
- Phusion DNA Polymerase, incorporates nucleotides to “fill in” the gaps in the annealed DNA fragments. it is a hot-start, proofreading PCR enzyme, enabling generation of PCR amplicons with high sequence accuracy, sensitivity, and specificity. Phusion DNA Polymerase is a thermostable polymerase that possesses 5´→ 3´ polymerase activity, 3´→ 5´ exonuclease activity and will generate blunt-ended products.
- nucleotides: building blocks for new DNA strands during amplification.
- Buffer: it provides the optimal pH, ionic strength, and Mg²⁺ concentration (1.5 mM final) required for Phusion DNA polymerase to bind primers, extend DNA efficiently, and maintain its high fidelity
- What are some factors that determine primer annealing temperature during PCR?
- the specific primer annealing temperature depends on specific length and sequence of the primers.
- it depends also on melting temperature of the primers and therefore GC content TM = 4(G + C) + 2(A+T)
- There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
Restriction Enzyme Digest
- It is a process in which DNA is cut at specific sites, dictated by the surrounding DNA sequence.
- is accomplished by incubation of the target DNA molecule with restriction enzymes - enzymes that recognize and bind specific DNA sequences and cleave at specific nucleotides either within the recognition sequence or outside of the recognition sequence.
- Restriction digestion can result in the production of blunt ends (ends of a DNA molecule that end with a base pair) or sticky ends
- Restriction digestion is usually used to prepare a DNA fragment for subsequence molecular cloning
- The results of a restriction digestion can be evaluated by gel electrophoresis, in which the products of the digestion are separated by molecule length
- The components of a typical restriction digestion reaction include the DNA template, the restriction enzyme of choice, a buffer and sometimes BSA protein. The reaction is incubated at a specific temperature required for optimal activity of the restriction enzyme and terminated by heat.
- reaction mixing, incubation at specific temperatures and time
PCR
- is a method for amplifying DNA. millions of copies of a DNA sequence can be generated from a single copy or just a few copies of DNA
- PCR protocols consist of assembling a PCR reaction mix containing Taq polymerase (a thermostable DNA polymerase that can withstand the high temperatures required for thermal cycling), primers (short DNA sequences that define the target region for amplification), deoxynucleotide triphosphates (dNTPs, the building blocks of DNA) and MgCl2 (Taq polymerase co-factor) in a buffered solution. The reaction mixture undergoes three basic thermal cycling steps including (1) denaturation (usually at 95°C, (2) annealing (usually lowest primer melting temperature - 5°C, (3) extension (usually at 72°C).
- components mixing , 2. denaturation step where the double stranded DNA denatures into single strands, annealing for primers annealing then extension where DNA polymerase extends the DNA creating million copies of required DNA fragment
- How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
- after generating dna fragments by pcr, run agarose gel to check for size and yield
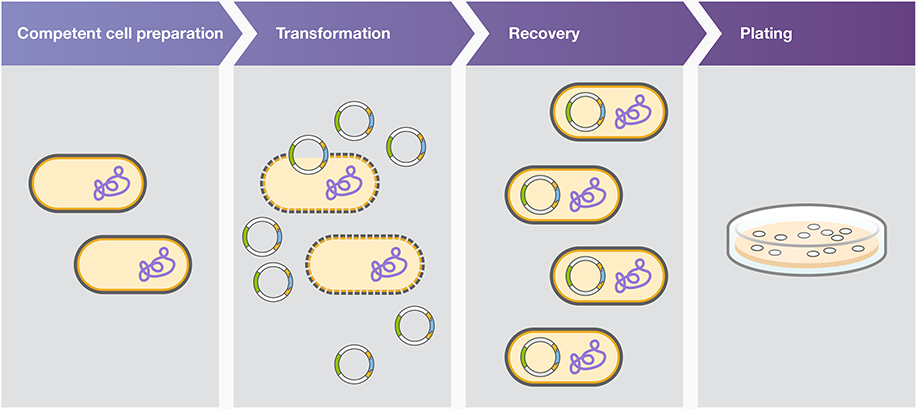
How does the plasmid DNA enter the E. coli cells during transformation?
Heat shock transformation is also known as chemical transformation and calcium chloride transformation. This method involves subjecting the cells to a sudden increase in temperature, often achieved by briefly immersing them in hot water or placing them in a heating block, followed by a rapid decrease in temperature through incubation on ice. The heat shock causes the cell membrane to become more permeable, facilitating the uptake of exogenous DNA.
Plasmid uptake by chemically competent cells is facilitated by heat shock, and plasmid uptake by electrocompetent cells is facilitated by electroporation.

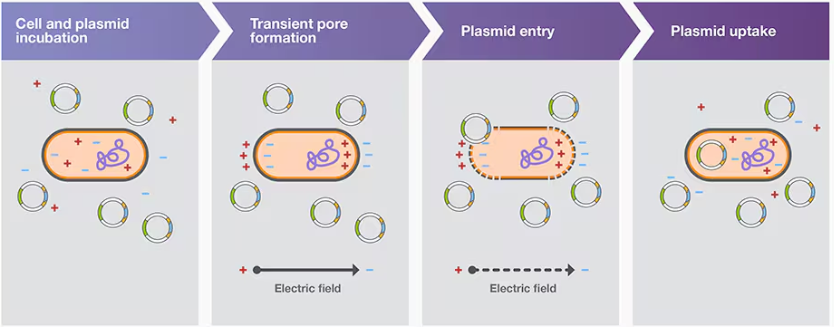
Electroporation transformation
Bacterial transformation aided by electroporation is called electroporation transformation; electroporation involves using an electroporator to subject competent cells and the plasmid carrying DNA construct to a brief pulse of a high-voltage electric field (Figure 3B). This treatment induces transient pores in cell membranes, which permits plasmid entry into the cells
One of the main issues with electroporation is arcing, or electric discharge, which may lower cell viability and transformation efficiency. Arcing often results from electroporation in conductive buffers, such as those containing MgCl2 and phosphates.
- it enters through creating pores in bacterial membrane, these pores can be created by either heat shock

- Describe another assembly method in detail (such as Golden Gate Assembly)
- Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
- Model this assembly method with Benchling or Asimov Kernel! https://www.snapgene.com/guides/golden-gate-assembly
References
- https://www.neb.com/en/products/m0531-phusion-high-fidelity-pcr-master-mix-with-hf-buffer?srsltid=AfmBOopKylwQ43HL-LppGa9GH2B6iMXIrXuwRITgpYMEq3PQqljrIYko
- https://www.thermofisher.com/eg/en/home/life-science/cloning/cloning-learning-center/invitrogen-school-of-molecular-biology/pcr-education/pcr-reagents-enzymes/pcr-cycling-considerations.html
- https://www.genscript.com/what-is-restriction-digestion.html
- https://www.genscript.com/what-is-pcr.html
- https://www.addgene.org/protocols/gibson-assembly/
- https://www.thermofisher.com/eg/en/home/life-science/cloning/cloning-learning-center/invitrogen-school-of-molecular-biology/molecular-cloning/transformation/bacterial-transformation-workflow.html
Asimov Kernel
- Create a Repository for your work
- Create a blank Notebook entry to document the homework and save it to that Repository
- Explore the devices in the Bacterial Demos Repo to understand how the parts work together by running the Simulator on various examples, following the instructions for the simulator found in the “Info” panel (click the “i” icon on the right to open the Info panel)
- Create a blank Construct and save it to your Repository
- Recreate the Repressilator in that empty Construct by using parts from the Characterized Bacterial Parts repository
- Search the parts using the Search function in the right menu
- Drag and drop the parts into the Construct
- Confirm it works as expected by running the Simulator (“play” button) and compare your results with the Repressilator Construct found in the Bacterial Demos repository
- Document all of this work in your Notebook entry - you can copy the glyph image and the simulator graphs, and paste them into your Notebook
- Build three of your own Constructs using the parts in the Characterized Bacterials Parts Repo
- Explain in the Notebook Entry how you think each of the Constructs should function
- Run the simulator and share your results in the Notebook Entry
- If the results don’t match your expectations, speculate on why and see if you can adjust the simulator settings to get the expected outcome
References