Week 4 HW: Protein Design Part i


𓃠 Week 4 Homework 𓃠
Part A. Conceptual Questions
1. How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons)
Proteins make about 20%-30% of the composition of meat, so we can take 25% as the average, this means 500 x 0.25 = 125g protein, to determine molar mass, 1 Dalton is approximately equal to 1 gram per mole (g/mol), so an average amino acid weighs 100 g/mol which when divided by the molar mass of 100 daltons gives 1.25 moles of amino acids, then we can multiply this by Avogadro's number (6.022x1023), we get 7.5275 x 1023 molecules of amino acids in the 500 grams of meat (this is extremely huge wow XD)
2. Why do humans eat beef but do not become a cow, eat fish but do not become fish?
Because when we eat them, we do not inherit their DNA or Genomic composition, we just digest them and use the amino acids and other nutrients from their catabolism to build our own self using our own Genome instructions
3. Why are there only 20 natural amino acids?
The currently settled on 20 Amino acids provide a sweet spot between the most chemical diversity with the fewest components to a build functioning complex system, however we dont seem to be only stuck with 20, as there are 2 other natural amino acids Selenocysteine is one of them, it is used by humans and bacteria for precise antioxidant enzymes, the other is Pyrrolysine which is found strictly in ancient methane-producing archaea.
4. Can you make other non-natural amino acids? Design some new amino acids.
Yes, you definetly can, actually non natural amino acids have been created before, usually by chemically modifying the “R-group” side chain to create proteins with other powers like heat resistance, fluorescence, or adhesive properties. Maybe we can design new amino acids that can glow different colors depending on how powerful radiation is in a certain place.
5. Where did amino acids come from before enzymes that make them, and before life started?
(Had to use AI for this one) Amino acids were already being mass produced by high energy physics and geology, forming spontaneously whenever simple carbon compounds interact with energy, some common sources were Atmospheric Sparks (Lightning), Cosmic Delivery (Meteorites), and Deep-Sea Vents (Geothermal Heat).
6. If you make an α-helix using D-amino acids, what handedness (right or left) would you expect?
it will form a Left-Handed helix, since D-amino Acids are mirrors of the L-amino acids that make Right handed helices
8. Why are most molecular helices right-handed?
Because more than 99% of the amino acids in the body are in the L-form (Left-handed), as they are the exclusive building blocks for proteins and enzymes, so most of the produced helices will be right handed.
9. Why do β-sheets tend to aggregate? What is the driving force for β-sheet aggregation?
Unlike alpha helices, which are self contained cylinders, a single beta strand is structurally incomplete. Its edges have exposed hydrogen bond donors and acceptors that act like open Velcro hooks. If these edges are not protected by the protein’s own fold, they will inevitably recruit strands from neighboring proteins to satisfy these bonds, triggering a stacking event. The primary driving force for beta sheet aggregation is the thermodynamic stability gained from intermolecular hydrogen bonding combined with the hydrophobic effect.
10. Why do many amyloid diseases form β-sheets? Can you use amyloid β-sheets as materials?
Proteins normally fold into delicate 3D shapes to do their jobs, but keeping those shapes takes constant effort. If a protein gets damaged, it can collapse into a “β-sheet”, a misfolded shape that acts like a microscopic zipper. These sheets lock together so tightly and squeeze out so much water that the body’s natural recycling system literally cannot break them apart. That extreme, indestructible stability is exactly why they clump up in the brain and cause diseases.
However, that same indestructible nature makes them an incredible raw material. Because these microscopic β-sheets are stronger than steel and completely waterproof, scientists are now programming cells to grow harmless, synthetic versions of them in the lab. We can harvest these lab grown amyloids to build strong biofibers.
Part B: Protein Analysis and Visualization
Briefly describe the protein you selected and why you selected it.
crtI (phytoene desaturase), it is a bacterial enzyme that handles the multi-step conversion of colorless phytoene into the bright red pigment lycopene. i mainly chose it because i was working on a metabolic engineering project recently on lycopene production so i thought it would be cool to pick crtI and explore it more
Identify the amino acid sequence of your protein.
MKPTTVIGAGFGGLALAIRLQAAGIPVLLLEQRDKPGGRAYVYEDQGFTFDAGPTVITDPSAIEELFALAGKQLKEYVELLPVTPFYRLCWESGKVFNYDNDQTRLEAQIQQFNPRDVEGYRQFLDYSRAVFKEGYLKLGTVPFLSFRDMLRAAPQLAKLQAWRSVYSKVASYIEDEHLRQAFSFHSLLVGGNPFATSSIYTLIHALEREWGVWFPRGGTGALVQGMIKLFQDLGGEVVLNARVSHMETTGNKIEAVHLEDGRRFLTQAVASNADVVHTYRDLLSQHPAAVKQSNKLQTKRMSNSLFVLYFGLNHHHDQLAHHTVCFGPRYRELIDEIFNHDGLAEDFSLYLHAPCVTDSSLAPEGCGSYYVLAPVPHLGTANLDWTVEGPKLRDRIFAYLEQHYMPGLRSQLVTHRMFTPFDFRDQLNAYHGSAFSVEPVLTQSAWFRPHNRDKTITNLYLVGAGTHPGAGIPGVIGSAKATAGLMLEDLI
The length of the protein is 492 aminoacids, with Leucine (L) being the most common amino acid which appears 57 times.
Uniprot ID
P21685and it has 242 homologs.
Identify the structure page of your protein in RCSB
- The structure for Phytoene Dehydrogenase was deposited on Jan 26th 2012, it has a good resolution of 2.4 Å
- Apart from the Main Protein Polymer, it has other components including Ligands, Water and Ions.
Open the structure of your protein in any 3D molecule visualization software.
Cartoon View
Ribbon View
Ball & Sticks View
crtI Residues, Blue are hydrophilic and red are hydrophobic, it seems to have more hydrophilic residues than hydrophobic ones
crtI Cavities
Part C. Using ML-Based Protein Design Tools
C1. Protein Language Modeling
- Some Patterns that I identified include that some amino acids in the crtI protein, especially those between positions 4-8 seem to have the lowest scores for mutations possibly highlighting how important or cruical these are for the protein’s stability and function.
- On the other hand, some amino acids show high and positive scores when mutated to other amino acids, potentially showing better stability or favorable changes, such as positions 354 & 365 seem to be having a high score for each differtiated amino acid.
- Position 269 seems to have the highest score when mutated to Valine (V) or Isoleucine (I).
Here is the TSNE map, i hightlighted my crtI protein in black.
I found this neighborhood cluster that hosts many dehydrogenase proteins like Retinal Dehydrogenase & Putative dehydrogenase, and other Reductase proteins too
Here in this cluster i found many microbial cytochrome proteins and some photosystem proteins, and other mitochondiria and ATP related ones
C2. Protein Folding
This is how the normal crtI folded
Here i tried swapping amino acids 10-13 to be all Valine (V) and here is the result
Here i actually picked two of the lowest scores from the mutation scan heatmap and decided to switch positions 4 and 5 which were Threonine (T) and switched them with Glutamic acid (E) which had a very low negative scores of -9.97 and -9.39 for positions 4 and 5 respectively
Here i tried to go a bit crazy and removed 30 amino acids, in sets of 10 randomly across the protein to see what effect this could have on it
C3. Protein Generation
Inverse Folded Protein Sequence:
ALPVAVVDGGAGGLALAIRLKAAGLPVVLLESGXXXXXXXGSVEKDGFIFDTTDLIITDPSPIEALFALAGKKLEDYVKLLKVEPFYRMVFENGRTFDFNQDLAAILAQIAKFNPADVAGFQALMAALRARYAEGYPXXGPVPYLDFDRLLRVAPTLRESPAYKAIHAEIAKYIKDPFLQLALTTFHLLVSGRPXXDTDPYHLISYFTQDWDVYYPEGGYKALVEAMKTLLRDLGGTIVEGARVARFELEGNRVVAVVLEDGRVIPVSAVALPPAXXXXXXXXXXXXXXXXXXXXXXXXXXXRYDLLELFFATDKRYDHLAMYTLVFKPVXXXXXXXXXXXXXLDYSLALVIYNPNVVDPSLAPEGGNSLYVKAPVPALGSANIDWSVWGPEKAEELLAYLEAHLMPGLRASLVTHAIVTPADKLXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXELENLFVIGXXXXXXGGIPGAIAAAFEVADRILAALK
I generated an AA Probaility Heat map and the ESM Fold for the ProteinMPNN produced sequence, it actually doesn’t really hold the same shape as the original one, when i checked the output of the ProteinMPNN on colab it was this:
>T=0.1, sample=0, score=0.8096, seq_recovery=0.4642 ALPVAVVDGGAGGLALAIRLKAAGLPVVLLESGXXXXXXXGSVEKDGFIFDTTDLIITDPSPIEALFALAGKKLEDYVKLLKVEPFYRMVFENGRTFDFNQDLAAILAQIAKFNPADVAGFQALMAALRARYAEGYPXXGPVPYLDFDRLLRVAPTLRESPAYKAIHAEIAKYIKDPFLQLALTTFHLLVSGRPXXDTDPYHLISYFTQDWDVYYPEGGYKALVEAMKTLLRDLGGTIVEGARVARFELEGNRVVAVVLEDGRVIPVSAVALPPAXXXXXXXXXXXXXXXXXXXXXXXXXXXRYDLLELFFATDKRYDHLAMYTLVFKPVXXXXXXXXXXXXXLDYSLALVIYNPNVVDPSLAPEGGNSLYVKAPVPALGSANIDWSVWGPEKAEELLAYLEAHLMPGLRASLVTHAIVTPADKLXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXELENLFVIGXXXXXXGGIPGAIAAAFEVADRILAALK
The sequence recovery was 0.4642, i tried to regenerate it again and got 0.4840, still both are under 50% so that explains why the produced Fold is quite different
Part D. Group Brainstorm on Bacteriophage Engineering
By: 2026a-nourelden-rihan, 2026a-ritika-saha, 2026a-rahul-yaji, 2026a-keerthana-gunaretnam
Goals and Strategy
Primary Goal: Increase the structural stability of the MS2 bacteriophage lysis protein (L) while preserving its ability to lyse bacterial cells.
Secondary Goal: Decrease the protein’s reliance on the host chaperone DnaJ, which could allow the lysis protein to function more efficiently and independently in engineered systems
Design Focus: The strategy involves stabilizing the transmembrane and oligomerization regions, protecting essential functional motifs (such as the L48–S49 motif), and modifying the N-terminal region to bypass DnaJ dependence
Computational Pipeline
The project utilizes a multi-step computational protein engineering pipeline to rationally design mutations:
Homolog Discovery (BLAST): Identifying related lysis proteins to find evolutionarily conserved residues and natural sequence variations
Multiple Sequence Alignment (Clustal Omega): Mapping essential structural regions and differentiating between highly conserved zones (to be protected) and mutable sites
In Silico Mutagenesis (ESM): Using protein language models to generate mutation heatmaps and rationally select amino acid substitutions that improve protein fitness and stability
Structure Prediction (ESMFold): Modeling the 3D structures of promising mutants to ensure the essential transmembrane helix is not distorted
Complex Prediction (AlphaFold Multimer): Evaluating whether mutated proteins can successfully form the required oligomeric pore complex (>10 subunits) and assessing if N-terminal mutations successfully reduce interactions with DnaJ
Expected Outcomes and Applications
The pipeline is expected to yield MS2 L variants with enhanced structural stability, proper transmembrane insertion, lower aggregation risks, and reduced DnaJ dependency
These optimized proteins have potential downstream applications in synthetic phage engineering, antimicrobial protein development, and bacterial ghost cell production
Challenges and Future Validation
Key computational challenges include the limited training data for small transmembrane toxins (as models primarily focus on globular proteins), the poor database annotation of single-gene lysis proteins (amurins), and the risk of over-stabilizing the protein, which could impede proper membrane insertion or functional oligomerization
Additionally, mutations might inadvertently expose protease cleavage sites, reducing stability inside the cell
Future steps will involve experimentally expressing the computationally identified mutants in E. coli to validate protein stability, lysis timing, and DnaJ independence
Schematic
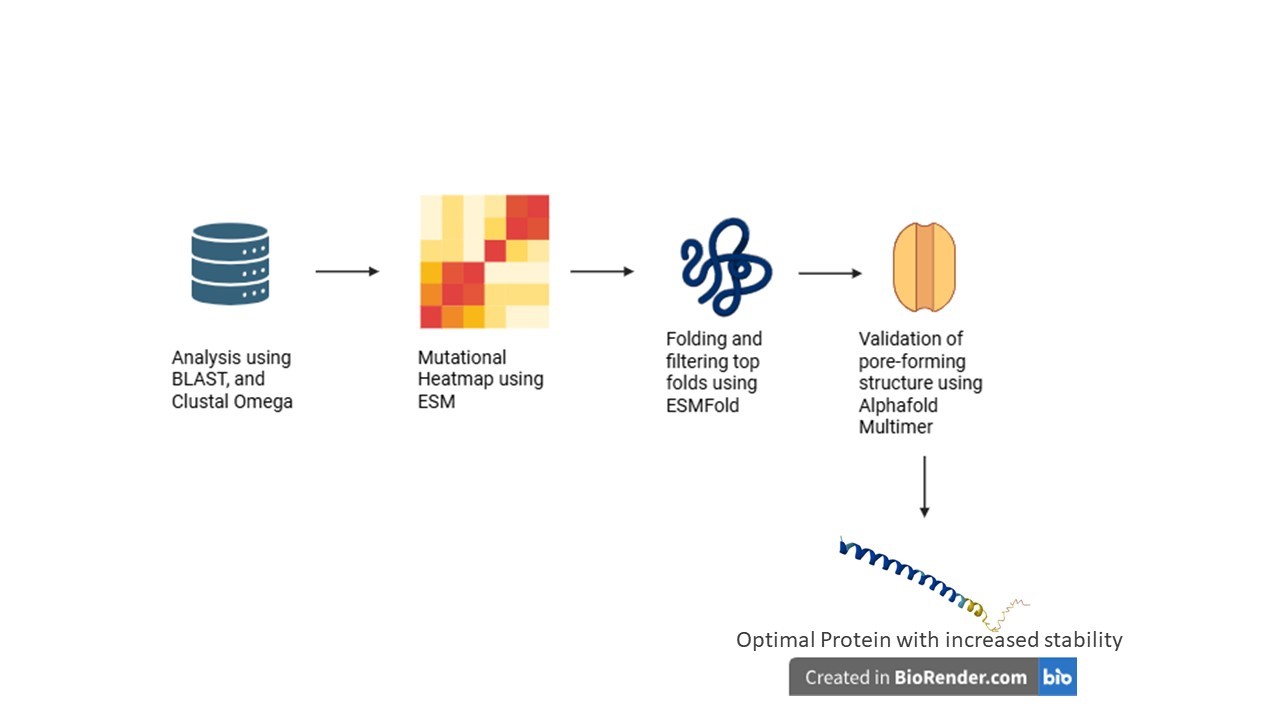

You can check out the fully detailed Project Proposal here