Week 10 HW: Imaging and Measurement


𓃠 Week 10 Homework 𓃠
Homework: Final Project
- α-Pinene titer: quantity of our target molecule produced per chassis per carbon source. Measured by GC-MS with dodecane overlay extraction from 96-well deep plates.
- AgPS protein identity and mass: purified AgPS-His6 protein (via Ni-NTA) can be characterized by intact protein LC-MS to confirm correct molecular mass and verify the His6 tag is present.
- Cell growth (OD₆₀₀): optical density at 600nm across all chassis × carbon source conditions. Measured on the PHERAstar FSX plate reader. Tells us if the strain is healthy while producing.
- Gene expression (mRNA level): RT-qPCR on the CFX Opus machine using primers targeting AgGPPS2 and AgPS transcripts. Confirms the genes are being transcribed in vivo.
Homework: Waters Part I — Molecular Weight
The Theoritical Molecular weight of the eGFP Sequence is 28006.60
I have decided to pick two peaks from the Figure 1 that was provided in the homework:
- m/z_n (Right Peak): 903.7148 (Lower charge)
- m/z_{n+1} (Left Peak): 875.4421 (Higher charge)
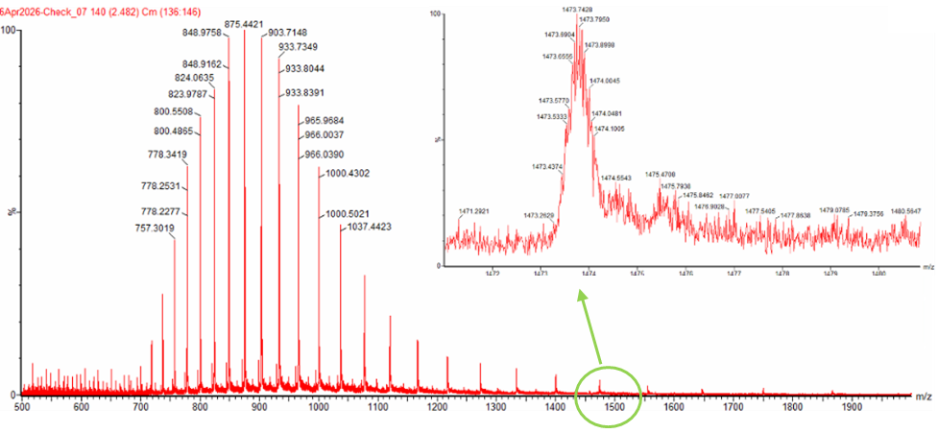

1. Charge State Calculation ($z$) $$ z = \frac{875.4421 - 1}{903.7148 - 875.4421} $$ $$ z = \frac{874.4421}{28.2727} $$ $$ z \approx 30.93 \rightarrow \mathbf{31} $$
2. Experimental Molecular Weight ($MW_{exp}$) $$ MW = 31 \times (903.7148 - 1) $$ $$ MW = 31 \times 902.7148 $$ $$ MW_{exp} = \mathbf{27,984.16 \text{ Da}} $$
3. Mass Accuracy (ppm) $$ \text{Accuracy} = \frac{|27,984.16 - 28,006.60|}{28,006.60} \times 1,000,000 $$ $$ \text{Accuracy} = \frac{22.44}{28,006.60} \times 1,000,000 $$ $$ \text{Accuracy} \approx \mathbf{801 \text{ ppm}} $$
Also the zoomed in part is blurry and unclear but i guess i cant identify the charge state anyways, the more i try to look it seems the numbers are so close together mostly 1473 and 1474 with lots of decimal changes in between
Homework: Waters Part III — Peptide Mapping - primary structure
MVSKGEELFTG VVPILVELDG DVNGHKFSVS GEGEGDATYG KLTLKFICTT GKLPVPWPTL VTTLTYGVQC FSRYPDHMKQ HDFFKSAMPE GYVQERTIFF KDDGNYKTRA EVKFEGDTLV NRIELKGIDF KEDGNILGHK LEYNYNSHNV YIMADKQKNG IKVNFKIRHN IEDGSVQLAD HYQQNTPIGD GPVLLPDNHY LSTQSALSKD PNEKRDHMVL LEFVTAAGIT LGMDELYKLE HHHHHH
The eGFP sequence has 19 Lysines (K) and 4 Arginines (R)
19 peptides will be generated from the tryptic digestion of eGFP.
i drew a blue line to mimic the >10% relative abundance but that is probably not exactly 10% XD but anyways, it gives exactly 19 peptides like what was predicted before so they match but if i did it exactly 10%, it would probably not match and they are more peptide fragments here than predicted
The most abundant peak is The peak eluting at 2.78 minutes and has an $m/z$ of 525.76, the spacing between isotopes is mostly 0.5 so the charge will be $1/0.5 = 2$
Single Charged Mass $$ \text{[M+H]}^+ = (z \times m/z) - (z - 1) $$ $$ \text{[M+H]}^+ = (2 \times 525.76) - 1 $$ $$ \text{[M+H]}^+ = 1051.52 - 1 $$ $$ \text{[M+H]}^+ = \mathbf{1050.52 \text{ Da}} $$
Experimental Mass = 1050.52 Da
The peptide perfect match would be 1050.5214 FEGDTLVNR
Mass Accuracy (ppm) $$ \text{Accuracy} = \frac{|MW_{\text{experiment}} - MW_{\text{theory}}|}{MW_{\text{theory}}} \times 1,000,000 $$ $$ \text{Accuracy} = \frac{|1050.52 - 1050.5214|}{1050.5214} \times 1,000,000 $$ $$ \text{Accuracy} = \frac{0.0014}{1050.5214} \times 1,000,000 $$ $$ \text{Accuracy} \approx \mathbf{1.33 \text{ ppm}} $$
Accuracy $\approx$ 1.33 ppm
88% of the eGFP sequence is confirmed by peptide mapping in Figure 6
Homework: Waters Part IV — Oligomers
| Polypeptide Subunit Name | Subunit Mass |
|---|---|
| 7FU | 340 kDa |
| 8FU | 400 kDa |
first we need to calculate the Mass of each of our oligomeric species
Decamer is 10 so didecamer is 20 and 3 decamer is 30 etc
- A) 7FU Decamer: Mass = 10 x 340 kDa = 3400 kDa (3.4 Megadaltons (MDa))
- B) 8FU Didecamer: Mass = 20 x 400 kDa = 8000 kDa (8 Megadaltons (MDa))
- C) 8FU 3-Decamer: Mass = 30 x 400 kDa = 3400 kDa (12 Megadaltons (MDa))
- D) 8FU 4-Decamer: Mass = 40 x 400 kDa = 3400 kDa (16 Megadaltons (MDa))
Homework: Waters Part V — Did I make GFP?
| Theoritical | Observed | PPM Mass Error |
|---|---|---|
| 28006.60 | 27,984.16 | 801 |