Homework
Weekly homework submissions:
Week 1 HW: Principles and Practices
Class Assignment Describe a biological engineering application or tool you want to develop and why. The project aims to develop a tool to promote Parkinson’s disease phenotype manifestation in human brain organoids by controllable induction of alpha-synuclein protein expression in dopaminergic neurons. The tool is a genetic construct containing switches and regulators to produce alpha-synuclein beyond normal levels in a subpopulation of cells in patient-derived brain organoids for the investigation of patient-specific pathogenic mechanisms, pathways, and phenotypes.
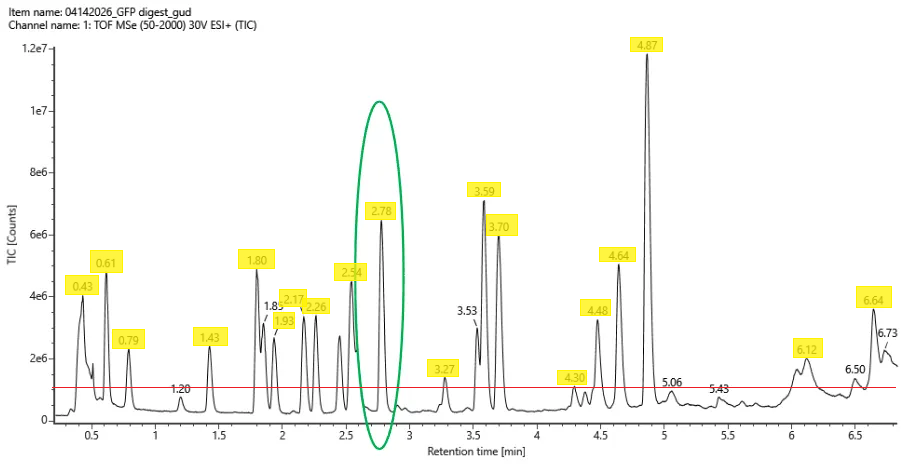
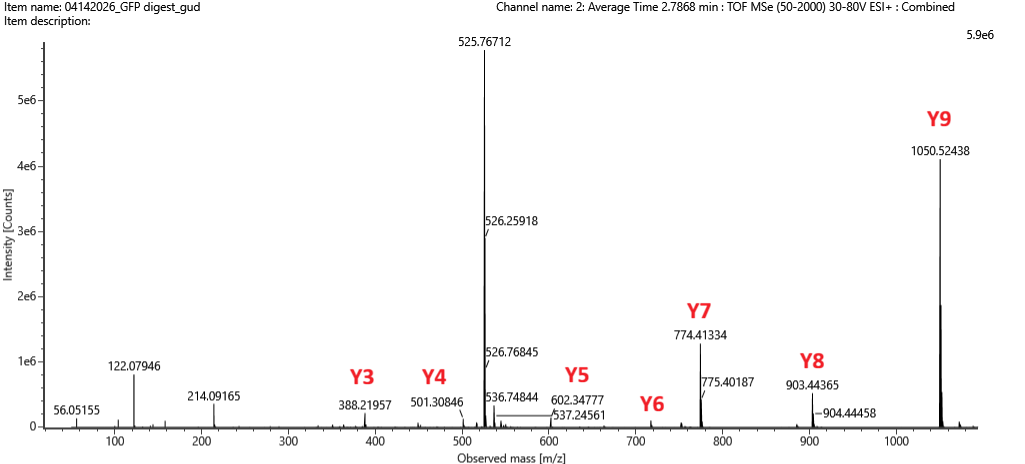
Week 10 HW: Imaging and Measurement (Mass Spectrometry)
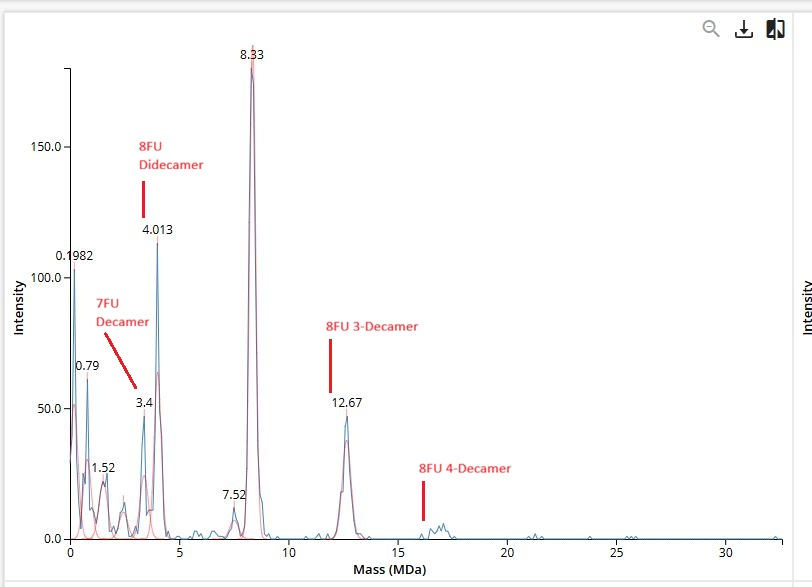
Waters Part I — Molecular Weight Theoretical molecular weight Based on the 247 aa sequence (including the initiator methionine, linker, and His-tag), the calculated average molecular weight is 28006.60 Da. Chromophore maturation (an autocatalytic post-translational modification where residues T66-Y67-G68 undergo cyclization resulting in −18.02) Da and oxidation (resulting in −2.02 Da) contribute to a 20.04 Da mass loss. The corrected predicted mass would therefore be 27,986.56 Da.
Week 2 HW: DNA Read, Write, and Edit
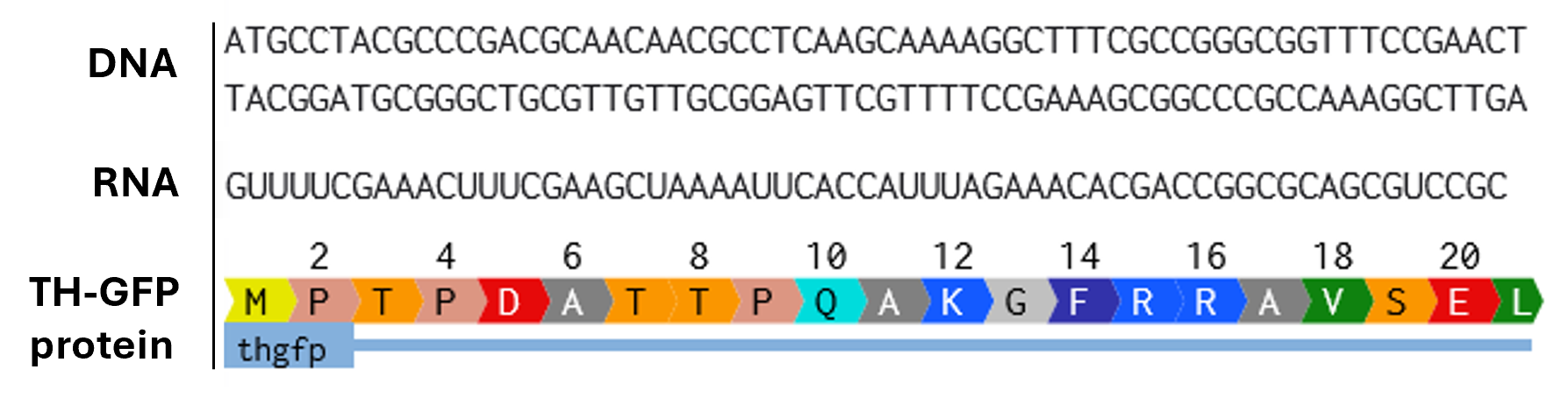
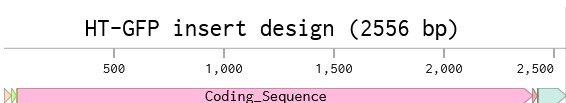
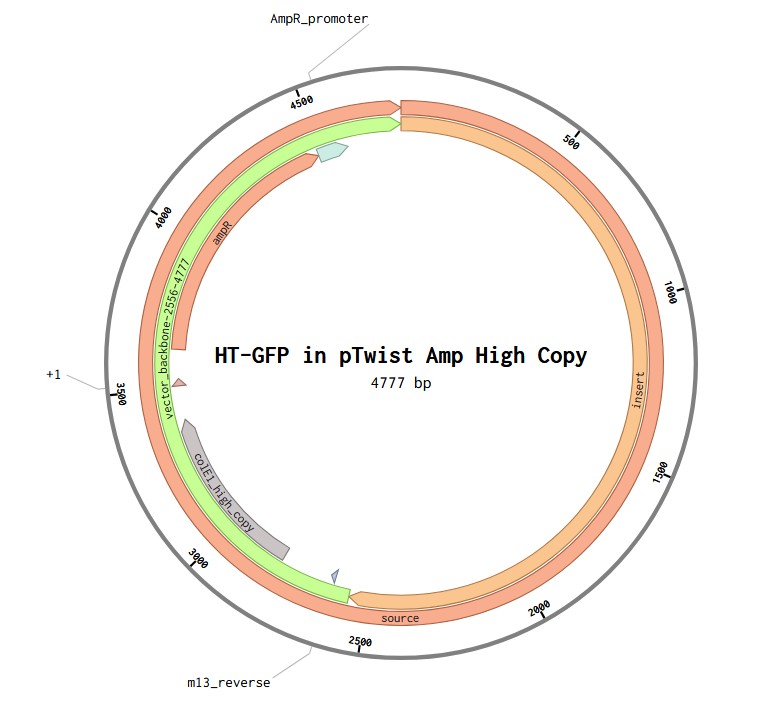
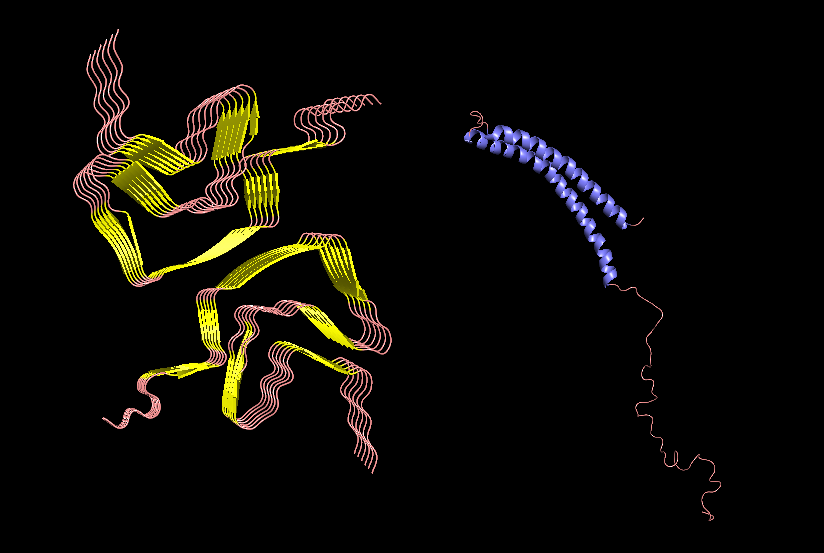
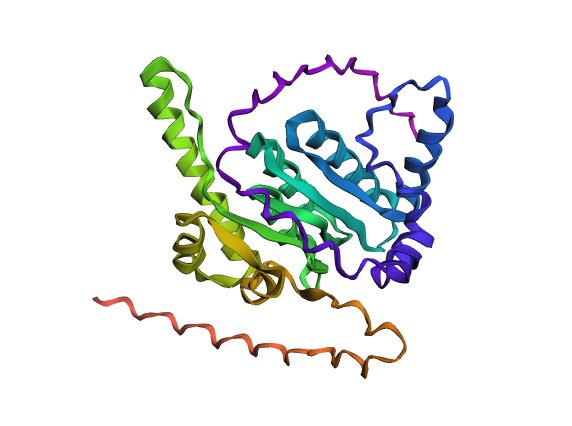
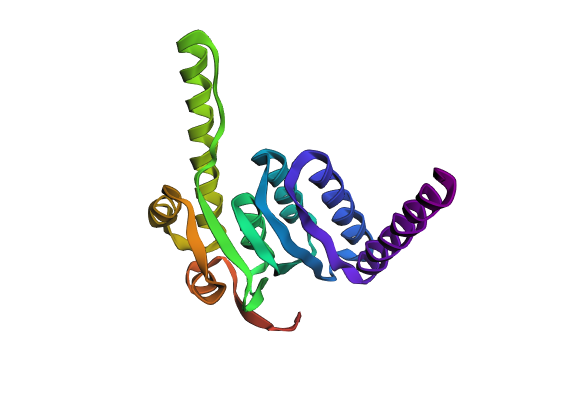
Part 1: Benchling & In-silico Gel Art This virtual digest of phage Lambda DNA was performed in Benchling. The enzymes used to process the DNA are listed below each column. Part 3: DNA Design Challenge I chose to practice designing a fluorescent-tagged human tyrosine hydroxylase (TH), relevant for my project on Parkinson’s disease. Tyrosine Hydroxylase converts tyrosine to dopamine and is an essential marker for my target cell population, dopaminergic neurons. Although in real applications, GFP under the TH promoter is used to trace dopaminergic neurons, and GFP fused to large (~56 kDa) TH can disrupt tetramerization, enzymatic activity, and folding, I chose to design a TH-GFP construct for training purposes.
Part 1: Opentrons Artwork This design was generated using the GUI at opentrons-art.rcdonovan.com and can be accessed through https://opentrons-art.rcdonovan.com/?id=1s7h4g7m1kn174o Coordinates mrfp1_points = [(27.5, 25.3),(25.3, 23.1),(23.1, 18.7),(20.9, 16.5),(18.7, 14.3),(36.3, 5.5),(38.5, 5.5),(27.5, 3.3),(29.7, 3.3),(31.9, 3.3),(34.1, 3.3),(36.3, 3.3),(20.9, 1.1),(23.1, 1.1),(25.3, 1.1),(16.5, -1.1),(18.7, -1.1)] mscarlet_i_points = [(29.7, 25.3),(27.5, 23.1),(29.7, 23.1),(25.3, 20.9),(27.5, 20.9),(25.3, 18.7),(23.1, 16.5),(20.9, 14.3),(16.5, 12.1),(18.7, 12.1),(16.5, 9.9),(14.3, 7.7),(38.5, 7.7),(12.1, 5.5),(34.1, 5.5),(9.9, 3.3),(38.5, 3.3),(14.3, -1.1)] electra2_points = [(31.9, 23.1),(29.7, 20.9),(31.9, 20.9),(34.1, 20.9),(27.5, 18.7),(29.7, 18.7),(31.9, 18.7),(25.3, 16.5),(27.5, 16.5),(29.7, 16.5),(23.1, 14.3),(25.3, 14.3),(27.5, 14.3),(20.9, 12.1),(23.1, 12.1),(18.7, 9.9),(20.9, 9.9),(16.5, 7.7),(18.7, 7.7),(14.3, 5.5),(16.5, 5.5),(12.1, 3.3),(9.9, 1.1),(9.9, -1.1)] mturquoise2_points = [(34.1, 18.7),(-5.5, 16.5),(31.9, 16.5),(34.1, 16.5),(36.3, 16.5),(29.7, 14.3),(31.9, 14.3),(34.1, 14.3),(25.3, 12.1),(27.5, 12.1),(29.7, 12.1),(23.1, 9.9),(25.3, 9.9),(27.5, 9.9),(20.9, 7.7),(23.1, 7.7),(1.1, 5.5),(18.7, 5.5),(14.3, 3.3),(16.5, 3.3),(12.1, 1.1),(23.1, -14.3),(7.7, -16.5),(9.9, -16.5),(12.1, -16.5),(14.3, -16.5),(-7.7, -18.7),(-5.5, -18.7),(-3.3, -18.7),(-1.1, -18.7),(1.1, -18.7),(3.3, -18.7),(5.5, -18.7),(7.7, -18.7),(9.9, -18.7),(12.1, -18.7),(16.5, -18.7),(-9.9, -20.9),(-7.7, -20.9),(-5.5, -20.9),(-3.3, -20.9),(-1.1, -20.9),(1.1, -20.9),(3.3, -20.9),(5.5, -20.9),(7.7, -20.9),(9.9, -20.9),(12.1, -20.9),(14.3, -20.9),(18.7, -20.9),(-12.1, -23.1),(-9.9, -23.1),(-7.7, -23.1),(-5.5, -23.1),(-3.3, -23.1),(-1.1, -23.1),(1.1, -23.1),(3.3, -23.1),(5.5, -23.1),(7.7, -23.1),(9.9, -23.1),(12.1, -23.1),(14.3, -23.1),(16.5, -23.1),(-14.3, -25.3),(-12.1, -25.3),(-9.9, -25.3),(-7.7, -25.3),(-5.5, -25.3),(-3.3, -25.3),(-1.1, -25.3),(1.1, -25.3),(3.3, -25.3),(5.5, -25.3),(-16.5, -27.5),(-14.3, -27.5),(-12.1, -27.5),(-9.9, -27.5)] azurite_points = [(-5.5, 14.3),(-3.3, 14.3),(-5.5, 12.1),(-1.1, 12.1),(-5.5, 9.9),(-3.3, 9.9),(-1.1, 9.9),(1.1, 9.9),(-7.7, 7.7),(-5.5, 7.7),(-3.3, 7.7),(-1.1, 7.7),(3.3, 7.7),(-7.7, 5.5),(-5.5, 5.5),(-3.3, 5.5),(-1.1, 5.5),(3.3, 5.5),(5.5, 5.5),(-7.7, 3.3),(-5.5, 3.3),(-3.3, 3.3),(-1.1, 3.3),(1.1, 3.3),(5.5, 3.3),(7.7, 3.3),(-7.7, 1.1),(-5.5, 1.1),(-3.3, 1.1),(-1.1, 1.1),(1.1, 1.1),(5.5, 1.1),(7.7, 1.1),(-9.9, -1.1),(-7.7, -1.1),(-5.5, -1.1),(-3.3, -1.1),(-1.1, -1.1),(1.1, -1.1),(3.3, -1.1),(5.5, -1.1),(7.7, -1.1),(-9.9, -3.3),(-7.7, -3.3),(-5.5, -3.3),(-3.3, -3.3),(-1.1, -3.3),(1.1, -3.3),(3.3, -3.3),(5.5, -3.3),(7.7, -3.3),(9.9, -3.3),(12.1, -3.3),(14.3, -3.3),(9.9, -5.5),(12.1, -5.5),(14.3, -5.5),(16.5, -5.5),(-12.1, -7.7),(-9.9, -7.7),(-7.7, -7.7),(-5.5, -7.7),(-3.3, -7.7),(-1.1, -7.7),(1.1, -7.7),(3.3, -7.7),(5.5, -7.7),(7.7, -7.7),(9.9, -7.7),(12.1, -7.7),(14.3, -7.7),(16.5, -7.7),(18.7, -7.7),(-12.1, -9.9),(-9.9, -9.9),(-7.7, -9.9),(-5.5, -9.9),(-3.3, -9.9),(-1.1, -9.9),(1.1, -9.9),(3.3, -9.9),(5.5, -9.9),(7.7, -9.9),(9.9, -9.9),(14.3, -9.9),(16.5, -9.9),(18.7, -9.9),(20.9, -9.9),(-12.1, -12.1),(-9.9, -12.1),(-7.7, -12.1),(-5.5, -12.1),(-3.3, -12.1),(-1.1, -12.1),(1.1, -12.1),(3.3, -12.1),(5.5, -12.1),(7.7, -12.1),(9.9, -12.1),(12.1, -12.1),(14.3, -12.1),(16.5, -12.1),(18.7, -12.1),(20.9, -12.1),(23.1, -12.1),(-12.1, -14.3),(-9.9, -14.3),(-7.7, -14.3),(-5.5, -14.3),(-3.3, -14.3),(-1.1, -14.3),(1.1, -14.3),(3.3, -14.3),(5.5, -14.3),(7.7, -14.3),(9.9, -14.3),(12.1, -14.3),(16.5, -14.3),(18.7, -14.3),(20.9, -14.3),(-12.1, -16.5),(-9.9, -16.5),(-7.7, -16.5),(-5.5, -16.5),(-3.3, -16.5),(-1.1, -16.5),(1.1, -16.5),(3.3, -16.5),(5.5, -16.5),(16.5, -16.5),(18.7, -16.5),(20.9, -16.5),(-12.1, -18.7),(-9.9, -18.7),(14.3, -18.7),(18.7, -18.7),(-14.3, -20.9),(-12.1, -20.9),(16.5, -20.9),(-14.3, -23.1),(-16.5, -25.3)] sfgfp_points = [(36.3, 14.3),(31.9, 12.1),(34.1, 12.1),(36.3, 12.1),(29.7, 9.9),(31.9, 9.9),(25.3, 7.7),(27.5, 7.7),(20.9, 5.5),(23.1, 5.5),(18.7, 3.3),(14.3, 1.1)] venus_points = [(34.1, 9.9),(36.3, 9.9),(29.7, 7.7),(25.3, 5.5),(20.9, 3.3),(16.5, 1.1)] mko2_points = [(-36.3, 12.1),(-38.5, 9.9),(-36.3, 9.9),(-34.1, 9.9),(38.5, 9.9),(-38.5, 7.7),(-36.3, 7.7),(-34.1, 7.7),(-31.9, 7.7),(31.9, 7.7),(34.1, 7.7),(36.3, 7.7),(-34.1, 5.5),(-31.9, 5.5),(-29.7, 5.5),(27.5, 5.5),(29.7, 5.5),(31.9, 5.5),(-29.7, 3.3),(-27.5, 3.3),(-25.3, 3.3),(23.1, 3.3),(25.3, 3.3),(-25.3, 1.1),(-23.1, 1.1),(-20.9, 1.1),(18.7, 1.1),(-20.9, -1.1),(-18.7, -1.1),(-16.5, -1.1),(12.1, -1.1),(-14.3, -3.3)] mjuniper_points = [(-3.3, 12.1),(1.1, 7.7),(3.3, 3.3),(3.3, 1.1),(-9.9, -5.5),(-7.7, -5.5),(-5.5, -5.5),(-3.3, -5.5),(-1.1, -5.5),(1.1, -5.5),(3.3, -5.5),(5.5, -5.5),(7.7, -5.5),(12.1, -9.9),(14.3, -14.3)] Part 2: Post-Lab Questions Part 3: Final Project Ideas Project 1: Tunable Induction of Alpha-Synuclein Expression for Modeling Parkinson’s Disease Aim:
Part A. Conceptual Questions Q1: How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons) 1 Da equals 1.66053906892(52)×10−27 kg, so 1 aa is 1.66053906892(52)×10−25 kg. The average protein fraction in meat is ~20%. Therefore, the total amount of protein is 100g. The number of aa in 100g = 0.1kg of protein is ~6×10²³ or Avogadro number, 1 mole.
Week 5: Protein design, part II
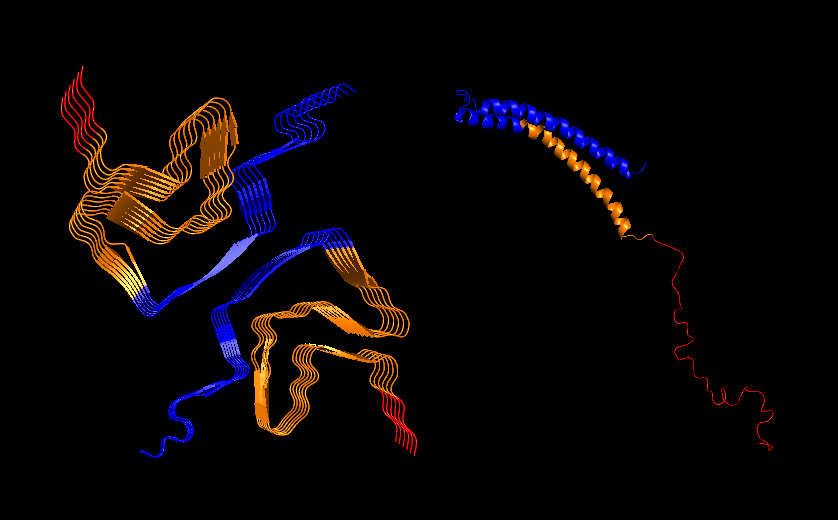
Part C: Final Project: L-Protein Mutants Original Sequence Soluble N-terminal domain C-terminal domain METRFPQQSQQTPASTNRRRPFKHEDYPCRRQQRSST LYVLIFLAIFLS KFTNQLLLSLLEAVIRTVTTLQQLLT Variable sites identified aligning BLAST results in ClustalOmega (8 in the N-terminus and 4 in the transmembrane domain highlighted): METRFPQQSQQTPASTNRRRPFKHEDYPCRRQQRSST LYVLIFLAIFLS KFTNQLLLSLLEAVIRTVTTLQQLLT Mutated Sequence 1 METRFPRQSQETLESTNRRRPRKHEDYPCRRQQRSST LYVLIFLAIFLSKFTNQLLLSLLEAVIRTVTTLQQLLT For this mutant, I modified the N-terminal domain, aiming to stabilize the disordered domain. I introduced as many charged pairs as possible in the variable sites (changed 4 out of 8 in the N-terminal domain), and additionally changed one conserved site on the left side of the 2nd pair. Summary of mutations
.png)