Week 2 HW: DNA Read, Write, and Edit
Part 1: Benchling & In-silico Gel Art
.png)
.png)
This virtual digest of phage Lambda DNA was performed in Benchling. The enzymes used to process the DNA are listed below each column.
Part 3: DNA Design Challenge
I chose to practice designing a fluorescent-tagged human tyrosine hydroxylase (TH), relevant for my project on Parkinson’s disease. Tyrosine Hydroxylase converts tyrosine to dopamine and is an essential marker for my target cell population, dopaminergic neurons. Although in real applications, GFP under the TH promoter is used to trace dopaminergic neurons, and GFP fused to large (~56 kDa) TH can disrupt tetramerization, enzymatic activity, and folding, I chose to design a TH-GFP construct for training purposes.
3.1. Choose your protein.
For TH, UniProt P07101 (Tyrosine 3-monooxygenase, Tyrosine 3-hydroxylase (TH)), I chose TH isoform 1 of 528 amino acids as it’s the canonical and most common isoform in the brain.
UniProt P07101-1, 528 aa: MPTPDATTPQAKGFRRAVSELDAKQAEAIMVRGQGAPGPSLTGSPWPGTAAPAASYTPTPRSPRFIGRRQSLIEDARKEREAAVAAAAAAVPSEPGDPLEAVAFEEKEGKAVLNLLFSPRATKPSALSRAVKVFETFEAKIHHLETRPAQRPRAGGPHLEYFVRLEVRRGDLAALLSGVRQVSEDVRSPAGPKVPWFPRKVSELDKCHHLVTKFDPDLDLDHPGFSDQVYRQRRKLIAEIAFQYRHGDPIPRVEYTAEEIATWKEVYTTLKGLYATHACGEHLEAFALLERFSGYREDNIPQLEDVSRFLKERTGFQLRPVAGLLSARDFLASLAFRVFQCTQYIRHASSPMHSPEPDCCHELLGHVPMLADRTFAQFSQDIGLASLGASDEEIEKLSTLYWFTVEFGLCKQNGEVKAYGAGLLSSYGELLHCLSEEPEIRAFDPEAAAVQPYQDQTYQSVYFVSESFSDAKDKLRSYASRIQRPFSVKFDPYTLAIDVLDSPQAVRRSLEGVQDELDTLAHALSAIG
For GFP, I chose EGFP variant GenBank AAB02572 of 239 aa, because of its higher-intensity emission and optimization for 37°C.
GenBank: AAB02572.1 239 aa: MVSKGEELFT GVVPILVELD GDVNGHKFSV SGEGEGDATY GKLTLKFICT TGKLPVPWPT LVTTLTYGVQ CFSRYPDHMK QHDFFKSAMP EGYVQERTIF FKDDGNYKTR AEVKFEGDTL VNRIELKGID FKEDGNILGH KLEYNYNSHN VYIMADKQKN GIKVNFKIRH NIEDGSVQLA DHYQQNTPIG DGPVLLPDNH YLSTQSALSK DPNEKRDHMV LLEFVTAAGI TLGMDELYK
As a linker, I chose a flexible (GGGGS)3 linker of 15 aa, as this type of linker is used in design in recombinant fusion proteins to increase spatial separation between domains, which could be useful for fusing GFP with a large protein. Since both N and C domains of TH are functional, with the N-terminal domain containing a phosphorylation site needed for the enzyme activation and the C-terminal domain allowing tetramerization, and the fused domain functionality can only be tested empirically, I chose to add a flexible linker to the C-terminal domain, as this linker can allow some spatial freedom for tetramerization.
Linker sequence: GGGGSGGGGSGGGGS
The full sequence (TH-linker-GFP): MPTPDATTPQAKGFRRAVSELDAKQAEAIMVRGQGAPGPSLTGSPWPGTAAPAASYTPTPRSPRFIGRRQSLIEDARKEREAAVAAAAAAVPSEPGDPLEAVAFEEKEGKAVLNLLFSPRATKPSALSRAVKVFETFEAKIHHLETRPAQRPRAGGPHLEYFVRLEVRRGDLAALLSGVRQVSEDVRSPAGPKVPWFPRKVSELDKCHHLVTKFDPDLDLDHPGFSDQVYRQRRKLIAEIAFQYRHGDPIPRVEYTAEEIATWKEVYTTLKGLYATHACGEHLEAFALLERFSGYREDNIPQLEDVSRFLKERTGFQLRPVAGLLSARDFLASLAFRVFQCTQYIRHASSPMHSPEPDCCHELLGHVPMLADRTFAQFSQDIGLASLGASDEEIEKLSTLYWFTVEFGLCKQNGEVKAYGAGLLSSYGELLHCLSEEPEIRAFDPEAAAVQPYQDQTYQSVYFVSESFSDAKDKLRSYASRIQRPFSVKFDPYTLAIDVLDSPQAVRRSLEGVQDELDTLAHALSAIG GGGGSGGGGSGGGGS MVSKGEELFT GVVPILVELD GDVNGHKFSV SGEGEGDATY GKLTLKFICT TGKLPVPWPT LVTTLTYGVQ CFSRYPDHMK QHDFFKSAMP EGYVQERTIF FKDDGNYKTR AEVKFEGDTL VNRIELKGID FKEDGNILGH KLEYNYNSHN VYIMADKQKN GIKVNFKIRH NIEDGSVQLA DHYQQNTPIG DGPVLLPDNH YLSTQSALSK DPNEKRDHMV LLEFVTAAGI TLGMDELYK
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
atgccgaccccggatgcgaccaccccgcaggcgaaaggctttcgccgcgcggtgagcgaa ctggatgcgaaacaggcggaagcgattatggtgcgcggccagggcgcgccgggcccgagc ctgaccggcagcccgtggccgggcaccgcggcgccggcggcgagctataccccgaccccg cgcagcccgcgctttattggccgccgccagagcctgattgaagatgcgcgcaaagaacgc gaagcggcggtggcggcggcggcggcggcggtgccgagcgaaccgggcgatccgctggaa gcggtggcgtttgaagaaaaagaaggcaaagcggtgctgaacctgctgtttagcccgcgc gcgaccaaaccgagcgcgctgagccgcgcggtgaaagtgtttgaaacctttgaagcgaaa attcatcatctggaaacccgcccggcgcagcgcccgcgcgcgggcggcccgcatctggaa tattttgtgcgcctggaagtgcgccgcggcgatctggcggcgctgctgagcggcgtgcgc caggtgagcgaagatgtgcgcagcccggcgggcccgaaagtgccgtggtttccgcgcaaa gtgagcgaactggataaatgccatcatctggtgaccaaatttgatccggatctggatctg gatcatccgggctttagcgatcaggtgtatcgccagcgccgcaaactgattgcggaaatt gcgtttcagtatcgccatggcgatccgattccgcgcgtggaatataccgcggaagaaatt gcgacctggaaagaagtgtataccaccctgaaaggcctgtatgcgacccatgcgtgcggc gaacatctggaagcgtttgcgctgctggaacgctttagcggctatcgcgaagataacatt ccgcagctggaagatgtgagccgctttctgaaagaacgcaccggctttcagctgcgcccg gtggcgggcctgctgagcgcgcgcgattttctggcgagcctggcgtttcgcgtgtttcag tgcacccagtatattcgccatgcgagcagcccgatgcatagcccggaaccggattgctgc catgaactgctgggccatgtgccgatgctggcggatcgcacctttgcgcagtttagccag gatattggcctggcgagcctgggcgcgagcgatgaagaaattgaaaaactgagcaccctg tattggtttaccgtggaatttggcctgtgcaaacagaacggcgaagtgaaagcgtatggc gcgggcctgctgagcagctatggcgaactgctgcattgcctgagcgaagaaccggaaatt cgcgcgtttgatccggaagcggcggcggtgcagccgtatcaggatcagacctatcagagc gtgtattttgtgagcgaaagctttagcgatgcgaaagataaactgcgcagctatgcgagc cgcattcagcgcccgtttagcgtgaaatttgatccgtataccctggcgattgatgtgctg gatagcccgcaggcggtgcgccgcagcctggaaggcgtgcaggatgaactggataccctg gcgcatgcgctgagcgcgattggcggcggcggcggcagcggcggcggcggcagcggcggc ggcggcagcatggtgagcaaaggcgaagaactgtttaccggcgtggtgccgattctggtg gaactggatggcgatgtgaacggccataaatttagcgtgagcggcgaaggcgaaggcgat gcgacctatggcaaactgaccctgaaatttatttgcaccaccggcaaactgccggtgccg tggccgaccctggtgaccaccctgacctatggcgtgcagtgctttagccgctatccggat catatgaaacagcatgatttttttaaaagcgcgatgccggaaggctatgtgcaggaacgc accattttttttaaagatgatggcaactataaaacccgcgcggaagtgaaatttgaaggc gataccctggtgaaccgcattgaactgaaaggcattgattttaaagaagatggcaacatt ctgggccataaactggaatataactataacagccataacgtgtatattatggcggataaa cagaaaaacggcattaaagtgaactttaaaattcgccataacattgaagatggcagcgtg cagctggcggatcattatcagcagaacaccccgattggcgatggcccggtgctgctgccg gataaccattatctgagcacccagagcgcgctgagcaaagatccgaacgaaaaacgcgat catatggtgctgctggaatttgtgaccgcggcgggcattaccctgggcatggatgaactg tataaataa
3.3. Codon optimization
Codon optimization is needed to adapt the DNA sequence to the different frequencies of tRNAs in the target organism in which expression is desired. It’s necessary because the same amino acid can be encoded by multiple synonymous codons, while different organisms have different preferences for which codons they use most frequently. By choosing codons that match the most abundant tRNAs in that organism, translation becomes more efficient.
HT-GFP sequence optimised for E.coli:
ATGCCTACGCCCGACGCAACAACGCCTCAAGCAAAAGGCTTTCGCCGGGCGGTTTCCGAACTGGATGCCAAGCAGGCGGAAGCCATTATGGTTCGTGGACAAGGCGCACCGGGTCCCAGCCTTACGGGTAGCCCTTGGCCGGGTACTGCGGCACCTGCTGCTAGCTACACGCCTACTCCTCGCTCACCCCGTTTTATAGGACGTCGTCAATCTCTCATAGAAGATGCTCGCAAAGAACGCGAAGCAGCAGTTGCAGCAGCGGCAGCAGCGGTACCTTCCGAGCCCGGAGACCCTTTAGAGGCTGTTGCATTTGAGGAAAAAGAAGGTAAAGCAGTTCTGAATTTGCTTTTCTCTCCTCGTGCGACAAAACCTTCGGCACTGTCACGGGCTGTCAAGGTTTTCGAAACTTTCGAAGCTAAAATTCACCATTTAGAAACACGACCGGCGCAGCGTCCGCGTGCCGGGGGGCCTCACTTGGAGTACTTCGTGCGTCTGGAGGTTCGACGTGGCGACCTTGCTGCTCTGTTGAGCGGTGTGCGCCAGGTTTCCGAAGATGTTCGTAGTCCTGCCGGACCTAAAGTACCATGGTTTCCGCGCAAAGTTTCCGAATTGGATAAGTGTCATCATCTTGTGACGAAATTTGATCCGGATCTTGACCTCGACCATCCGGGGTTCTCTGATCAGGTGTATCGTCAGCGTCGCAAACTCATTGCAGAGATTGCTTTTCAATATCGCCATGGCGACCCGATTCCCCGCGTAGAGTATACCGCTGAAGAAATAGCTACTTGGAAAGAAGTGTACACAACCCTGAAGGGCTTATATGCTACACACGCGTGTGGCGAACATTTAGAAGCCTTTGCTCTTCTCGAACGTTTCTCAGGTTATAGAGAGGACAACATTCCACAGTTAGAGGACGTTTCCCGATTTCTCAAAGAACGTACCGGCTTTCAGCTGAGACCCGTGGCCGGTTTATTGTCTGCTCGTGATTTCCTGGCATCACTGGCCTTTAGAGTATTCCAGTGTACTCAGTATATTCGCCATGCTTCCTCGCCAATGCACTCACCCGAACCAGATTGTTGCCATGAGTTACTTGGACATGTACCAATGCTCGCAGATCGAACATTTGCGCAATTCTCTCAAGATATCGGCCTGGCTAGTTTAGGCGCTTCAGATGAAGAAATTGAAAAGCTGTCCACACTGTACTGGTTCACCGTAGAATTTGGACTGTGCAAACAGAATGGCGAGGTTAAAGCGTACGGTGCCGGGCTTCTGTCCAGCTATGGTGAATTACTGCACTGTCTGTCAGAGGAGCCGGAGATTCGCGCATTTGATCCTGAAGCAGCCGCCGTCCAGCCATATCAAGATCAGACGTACCAGTCTGTGTATTTTGTTTCCGAAAGCTTTTCAGATGCCAAGGATAAGTTGCGCTCTTACGCTTCACGTATCCAACGCCCGTTTTCTGTAAAGTTCGACCCGTATACGCTGGCCATTGACGTCCTGGATAGCCCACAGGCAGTGCGCAGAAGTCTTGAAGGGGTTCAAGATGAGCTCGATACACTTGCCCATGCCCTTTCCGCTATAGGCGGGGGTGGTGGCTCTGGCGGTGGAGGTAGTGGAGGGGGTGGGAGCATGGTTTCAAAAGGGGAGGAGTTGTTTACTGGCGTGGTCCCAATCCTGGTAGAGTTAGACGGAGATGTTAACGGGCACAAATTCAGCGTTAGTGGTGAAGGGGAAGGCGACGCTACATATGGTAAACTGACACTGAAATTTATTTGTACCACCGGTAAGCTCCCAGTGCCCTGGCCGACTTTGGTTACCACGTTGACATATGGTGTACAATGTTTCTCCCGCTATCCTGACCACATGAAACAACATGATTTTTTCAAATCTGCTATGCCGGAAGGATATGTACAGGAACGTACGATCTTCTTCAAAGATGATGGCAACTATAAAACACGTGCCGAGGTTAAATTTGAGGGTGATACGCTGGTGAATCGCATTGAGTTAAAAGGAATAGACTTTAAGGAGGATGGGAATATTCTTGGCCACAAACTGGAGTACAATTACAATTCTCATAATGTGTATATCATGGCTGATAAACAGAAAAATGGTATCAAGGTTAACTTCAAAATCCGTCATAATATCGAGGATGGTTCTGTTCAGCTTGCTGATCATTATCAGCAAAATACGCCAATCGGTGATGGACCAGTCCTGTTGCCTGATAATCATTACCTCTCTACACAGTCAGCGCTGTCCAAAGACCCAAATGAGAAACGAGATCATATGGTATTGCTGGAATTCGTTACCGCTGCCGGAATTACACTTGGCATGGATGAATTATACAAATAA
3.4. You have a sequence! Now what?
Protein synthesis in cells. A double-stranded DNA first needs to be resolved by helicases to obtain a single strand accessible for RNA polymerase. RNA polymerase binds to a promoter on a 3’ to 5’ ‘template’ strand and produces a 5 to 3 strand of mRNA. In eukaryotic cells, mRNA gets modified right during transcription and just after that. It gets stabilised, protected from endonucleases, prepared for export to the cytoplasm, recognition by ribosomes, and translation initiation, i.e., a cap (a triphosphate link and a modified G) is added on its 5’ end, mRNA is spliced, and a poly-A tail sequence is added on its 3’ end. Mature mRNA then leaves the nucleus for translation in the cytoplasm. mRNA is read in 5’ to 3’ direction, in codons, within a ribosome, a protein-RNA complex or ribozyme, with a catalytic domain in ribosomal RNA. In a ribosome, protein synthesis is a series of processes (initiation, elongation, and termination) with ribosomal proteins acting as initiation factors, terminating factors, and mediating mRNA and tRNA meeting and release between ribosomal subunits. tRNAs recognise codons with their anti-codon and carry the corresponding amino acid, which binds with the previous one with the catalytic activity of the ribosome. If a protein is produced in bacteria, no preprocessing of mRNA occurs (no splicing like in eukaryotes as bacteria lack introns, but there is 5’end protection and addition of poly-A tail for degradation), and translation is coupled with transcription within the cytoplasm, in ribosomes that are smaller than eukaryotic (70S vs 80S). After synthesis, proteins undergo modifications, which are much more complex in eukaryotic cells than in bacteria. Post-translational modifications in eucaryotes occur in membrane organelles, endoplasmic reticulum, and Golgi apparatus, and include a variety of modifications (including phosphorylation, acetylation, formation of disulfide bonds, glycosylation, ubiquitination, most well-studied). In bacteria, these modifications are much simpler and happen in the cytoplasm.
Cell-free protein synthesis. A cell extract (eucaryotic or bacterial) or a PURE system is added to a DNA, and the extract/mix contains all enzymes and factors needed for transcription and translation, including energy sources and helicases.
3.5. How does it work in nature/biological systems?
- Describe how a single gene codes for multiple proteins at the transcriptional level.
During transcription in eukaryotes, RNA (pre-mRNA) gets spliced, and introns, which are non-coding sequences, are cut out while exons are joined. mRNA goes through either regular or alternative splicing in a spliceosome. In regular splicing, all introns are removed, and all exons are joined together, while in alternative splicing, some exons are removed and some are included. Either one or another splicing occurs, depending on the gene, cell type, and conditions (including developmental stage), instructing specific regulatory proteins mediating splicing. There are variations in how alternative splicing occurs (either an exon is skipped, an intron is left in the sequence, or a specific site is selected in the sequence). The splicing process is regulated by proteins that bind to pre-mRNA sites near exons and introns and either enhance or repress splicing (their relative concentration can also be a regulating factor, not just absence or presence). These proteins regulate the assembly of spliceosomes (complexes of specialized proteins and small RNA) on pre-mRNA and instruct the sites of spliceosome assembly and the efficiency of its binding. A spliceosome cuts out and joins fragments of pre-mRNA. The process overall allows for a variety of proteins that are synthesized from a single gene.
During transcription in bacteria, almost no splicing occurs, but still, different proteins can be produced from a single gene. RNA polymerase can be directed to different sequences related to a single gene by transcription factors, and so different transcripts are produced. Also, a ribosome can shift a reading frame due to specific patterns in mRNA sequence.
- Try aligning the DNA sequence, the transcribed RNA, and also the resulting translated Protein!!! See example below.
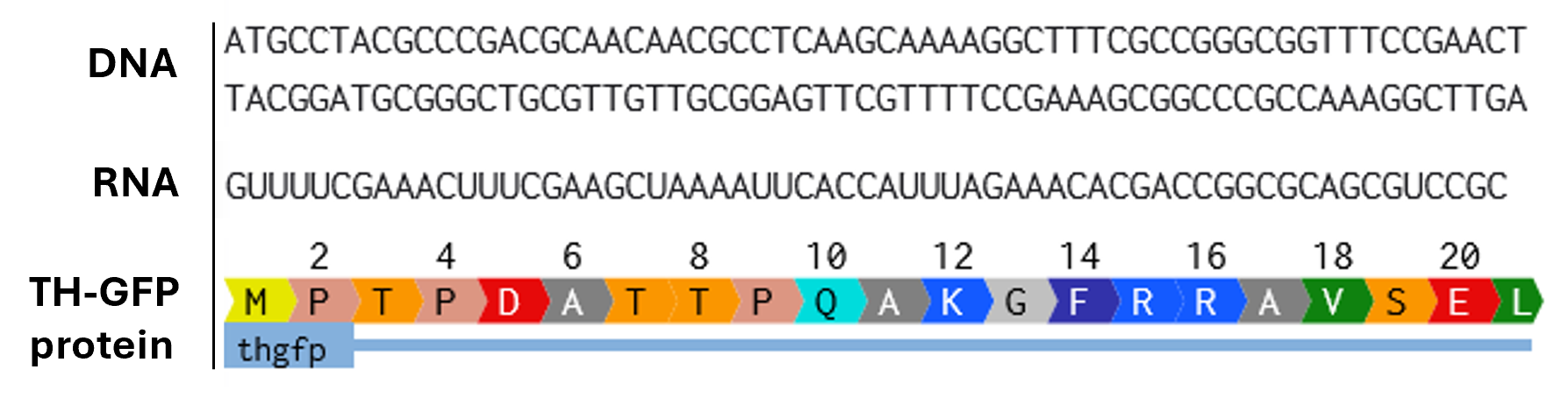

Rearranged snapshot of TH-GFP protein information flow from DNA to RNA to Protein. Captured from my Benchling and stitched together in a ppt.
4.1. Create a Twist account and a Benchling account
4.2. Build Your DNA Insert Sequence
A sequence for expression in E.coli:
Promoter (e.g. BBa_J23106): TTTACGGCTAGCTCAGTCCTAGGTATAGTGCTAGC RBS (e.g. BBa_B0034 with spacers for optimal expression): CATTAAAGAGGAGAAAGGTACC Start Codon: ATG Coding Sequence (codon optimized HT_GFP DNA): ATGCCTACGCCCGACGCAACAACGCCTCAAGCAAAAGGCTTTCGCCGGGCGGTTTCCGAACTGGATGCCAAGCAGGCGGAAGCCATTATGGTTCGTGGACAAGGCGCACCGGGTCCCAGCCTTACGGGTAGCCCTTGGCCGGGTACTGCGGCACCTGCTGCTAGCTACACGCCTACTCCTCGCTCACCCCGTTTTATAGGACGTCGTCAATCTCTCATAGAAGATGCTCGCAAAGAACGCGAAGCAGCAGTTGCAGCAGCGGCAGCAGCGGTACCTTCCGAGCCCGGAGACCCTTTAGAGGCTGTTGCATTTGAGGAAAAAGAAGGTAAAGCAGTTCTGAATTTGCTTTTCTCTCCTCGTGCGACAAAACCTTCGGCACTGTCACGGGCTGTCAAGGTTTTCGAAACTTTCGAAGCTAAAATTCACCATTTAGAAACACGACCGGCGCAGCGTCCGCGTGCCGGGGGGCCTCACTTGGAGTACTTCGTGCGTCTGGAGGTTCGACGTGGCGACCTTGCTGCTCTGTTGAGCGGTGTGCGCCAGGTTTCCGAAGATGTTCGTAGTCCTGCCGGACCTAAAGTACCATGGTTTCCGCGCAAAGTTTCCGAATTGGATAAGTGTCATCATCTTGTGACGAAATTTGATCCGGATCTTGACCTCGACCATCCGGGGTTCTCTGATCAGGTGTATCGTCAGCGTCGCAAACTCATTGCAGAGATTGCTTTTCAATATCGCCATGGCGACCCGATTCCCCGCGTAGAGTATACCGCTGAAGAAATAGCTACTTGGAAAGAAGTGTACACAACCCTGAAGGGCTTATATGCTACACACGCGTGTGGCGAACATTTAGAAGCCTTTGCTCTTCTCGAACGTTTCTCAGGTTATAGAGAGGACAACATTCCACAGTTAGAGGACGTTTCCCGATTTCTCAAAGAACGTACCGGCTTTCAGCTGAGACCCGTGGCCGGTTTATTGTCTGCTCGTGATTTCCTGGCATCACTGGCCTTTAGAGTATTCCAGTGTACTCAGTATATTCGCCATGCTTCCTCGCCAATGCACTCACCCGAACCAGATTGTTGCCATGAGTTACTTGGACATGTACCAATGCTCGCAGATCGAACATTTGCGCAATTCTCTCAAGATATCGGCCTGGCTAGTTTAGGCGCTTCAGATGAAGAAATTGAAAAGCTGTCCACACTGTACTGGTTCACCGTAGAATTTGGACTGTGCAAACAGAATGGCGAGGTTAAAGCGTACGGTGCCGGGCTTCTGTCCAGCTATGGTGAATTACTGCACTGTCTGTCAGAGGAGCCGGAGATTCGCGCATTTGATCCTGAAGCAGCCGCCGTCCAGCCATATCAAGATCAGACGTACCAGTCTGTGTATTTTGTTTCCGAAAGCTTTTCAGATGCCAAGGATAAGTTGCGCTCTTACGCTTCACGTATCCAACGCCCGTTTTCTGTAAAGTTCGACCCGTATACGCTGGCCATTGACGTCCTGGATAGCCCACAGGCAGTGCGCAGAAGTCTTGAAGGGGTTCAAGATGAGCTCGATACACTTGCCCATGCCCTTTCCGCTATAGGCGGGGGTGGTGGCTCTGGCGGTGGAGGTAGTGGAGGGGGTGGGAGCATGGTTTCAAAAGGGGAGGAGTTGTTTACTGGCGTGGTCCCAATCCTGGTAGAGTTAGACGGAGATGTTAACGGGCACAAATTCAGCGTTAGTGGTGAAGGGGAAGGCGACGCTACATATGGTAAACTGACACTGAAATTTATTTGTACCACCGGTAAGCTCCCAGTGCCCTGGCCGACTTTGGTTACCACGTTGACATATGGTGTACAATGTTTCTCCCGCTATCCTGACCACATGAAACAACATGATTTTTTCAAATCTGCTATGCCGGAAGGATATGTACAGGAACGTACGATCTTCTTCAAAGATGATGGCAACTATAAAACACGTGCCGAGGTTAAATTTGAGGGTGATACGCTGGTGAATCGCATTGAGTTAAAAGGAATAGACTTTAAGGAGGATGGGAATATTCTTGGCCACAAACTGGAGTACAATTACAATTCTCATAATGTGTATATCATGGCTGATAAACAGAAAAATGGTATCAAGGTTAACTTCAAAATCCGTCATAATATCGAGGATGGTTCTGTTCAGCTTGCTGATCATTATCAGCAAAATACGCCAATCGGTGATGGACCAGTCCTGTTGCCTGATAATCATTACCTCTCTACACAGTCAGCGCTGTCCAAAGACCCAAATGAGAAACGAGATCATATGGTATTGCTGGAATTCGTTACCGCTGCCGGAATTACACTTGGCATGGATGAATTATACAAATAA 7x His Tag (Let’s add a 7×His tag at the C-terminus of the protein to enable protein purification from E. coli): CATCACCATCACCATCATCAC Stop Codon: TAA Terminator (e.g. BBa_B0015): CCAGGCATCAAATAAAACGAAAGGCTCAGTCGAAAGACTGGGCCTTTCGTTTTATCTGTTGTTTGTCGGTGAACGCTCTCTACTAGAGTCACACTGGCTCACCTTCGGGTGGGCCTTTCTGCGTTTATA
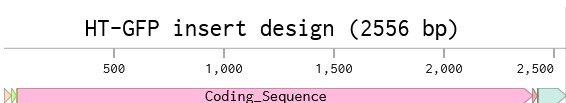

4.6. Choose Your Vector
Vector: pTwist Amp High Copy
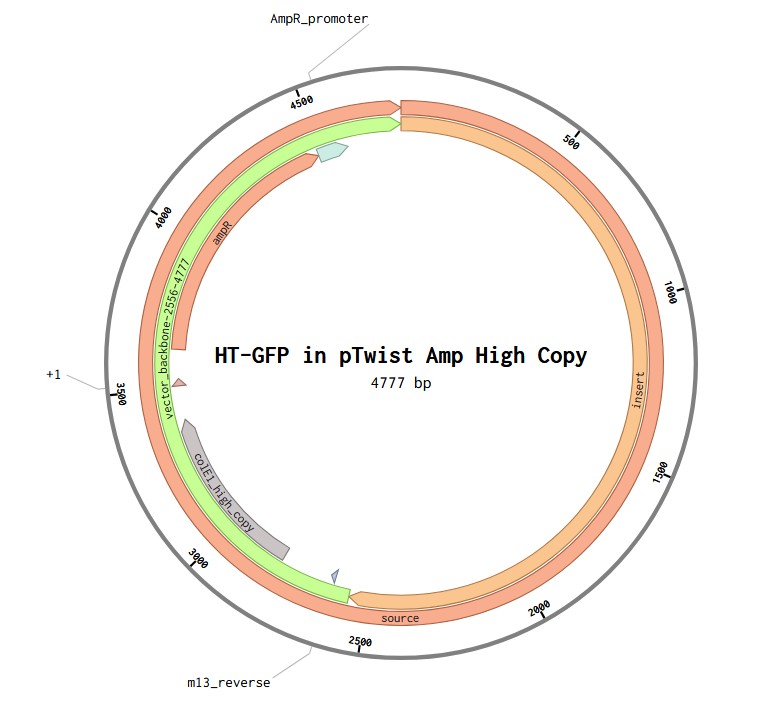

TH-GFP Plasmid
Part 5: DNA Read/Write/Edit
5.1 DNA Read
- What DNA would you want to sequence (e.g., read) and why?
It could be useful to sequence mitochondrial DNA of dopaminergic neurons and other cells of PD patients. As mutant mtDNA accumulates with age, mitochondrial dysfunction is a common phenotype observed in brain tissue, and the link between mtDNA maintenance and neurodegeneration is fairly recognised. Sequencing may reveal patterns of mitochondrial dysfunction and mt genome instability useful for predicting vulnerability to PD, neurodegenerative, and non-neurodegenerative conditions, as well as investigating fundamental aspects of aging in general. Concentrating on mtDNA involves a large patient population. However, to be a practical diagnostic tool (mtDNA biomarker), more accessible mtDNA in CSF or blood, or peripheral mtDNA has to be included, as tissue decline during aging is heterogeneous, and peripheral mtDNA may also contain information on the vulnerability of neuronal mtDNA to PD. It could be useful to aim at sequencing all 37 mitochondrial genes.
- In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?