Week 5: Protein design, part II
Part C: Final Project: L-Protein Mutants
Original Sequence
Soluble N-terminal domain C-terminal domain METRFPQQSQQTPASTNRRRPFKHEDYPCRRQQRSST LYVLIFLAIFLS KFTNQLLLSLLEAVIRTVTTLQQLLT
Variable sites identified aligning BLAST results in ClustalOmega (8 in the N-terminus and 4 in the transmembrane domain highlighted):
METRFPQQSQQTPASTNRRRPFKHEDYPCRRQQRSST LYVLIFLAIFLS KFTNQLLLSLLEAVIRTVTTLQQLLT
Mutated Sequence 1
METRFPRQSQETLESTNRRRPRKHEDYPCRRQQRSST LYVLIFLAIFLSKFTNQLLLSLLEAVIRTVTTLQQLLT
For this mutant, I modified the N-terminal domain, aiming to stabilize the disordered domain. I introduced as many charged pairs as possible in the variable sites (changed 4 out of 8 in the N-terminal domain), and additionally changed one conserved site on the left side of the 2nd pair.
Summary of mutations
Conserved site changed: 13P->L
Variable sites changed: (7Q->R, 11Q->E, 14A->E, 22F->R)
Pairs introduced by changing the 4 variable sites: Pair 1 (R7–E11), Pair 2 (E14–R18), Pair 3 (R22–D26)
Soluble N-terminal domain C-terminal domain
METRFPQQSQQTPASTNRRRPFKHEDYPCRRQQRSST LYVLIFLAIFLS KFTNQLLLSLLEAVIRTVTTLQQLLT (Original Sequence)
R---E LE--- R--- (Mutated Sites)
V V CV V (Conserved / Variable)
METRFPRQSQETLESTNRRRPRKHEDYPCRRQQRSST LYVLIFLAIFLS KFTNQLLLSLLEAVIRTVTTLQQLLT (Mutated Sequence 1)
Mutated Sequence 2
METRFPRQSQETLRSTNERRPRKHEDYPCRRQQRSST LYVLIFLAIFLSKFTNQLLLSLLEAVIRTVTTLQQLLT
For this mutant, I modified the previous sequence (Mutated Sequence 1), aiming to further stabilize the disordered domain.
I introduced 1 more mutation to a variable site to invert the second pair.
Summary of mutations
Conserved site changed: 13P->L
Variable sites changed: (7Q->R, 11Q->E, 14A->R, 18A->E, 22R->E)
Pairs introduced by changing the 5 variable sites: Pair 1 (R7–E11), Pair 2 (R14–E18), Pair 3 (R22–D26)
Soluble N-terminal domain C-terminal domain
METRFPQQSQQTPASTNRRRPFKHEDYPCRRQQRSST LYVLIFLAIFLS KFTNQLLLSLLEAVIRTVTTLQQLLT (Original Sequence)
R---E LR---E R--- (Mutated Sites)
V V CV V V (Conserved / Variable)
METRFPRQSQETLRSTNERRPRKHEDYPCRRQQRSST LYVLIFLAIFLS KFTNQLLLSLLEAVIRTVTTLQQLLT (Mutated Sequence 2)
AlphafoldServer was used to fold the monomers of Mutated Sequence 1 and Mutated Sequence 2. alfafold2_multimer_v2 was used to fold the multimers. alfafold2_multimer_v2 parameters used:
num_relax: 0
template_mode: none
msa_mode: mmseqs2_uniref_env
Pair mode: paired
num_recycles: 3
recycle_early_stop_tolerance: auto
relax_max_iterations: 200
pairing_strategy: greedy
max_msa: auto
num_seeds: 1
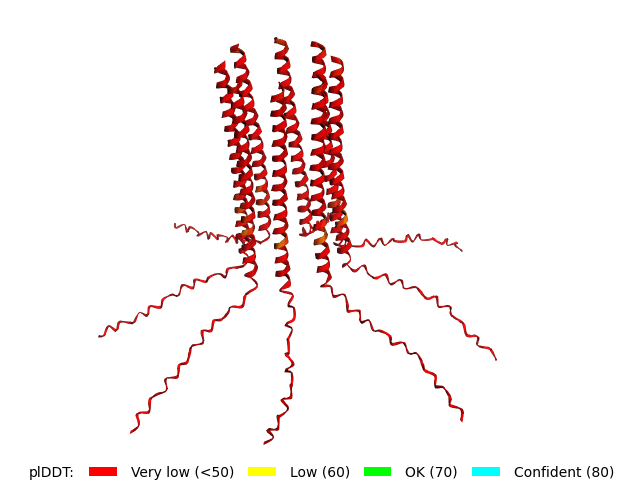 Original Sequence Multimer | 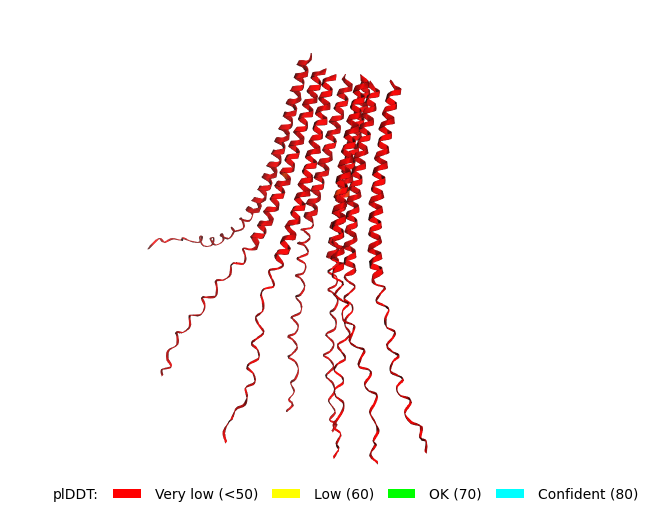 Mutant 1 pLDDT=37.6, pTM-0.189, ipTM = 0.127. 3 pairs/bridges introduced, 1 conserved site changed (13P->L), RRR site kept, (1 conserved and 4 variable sites changed) | 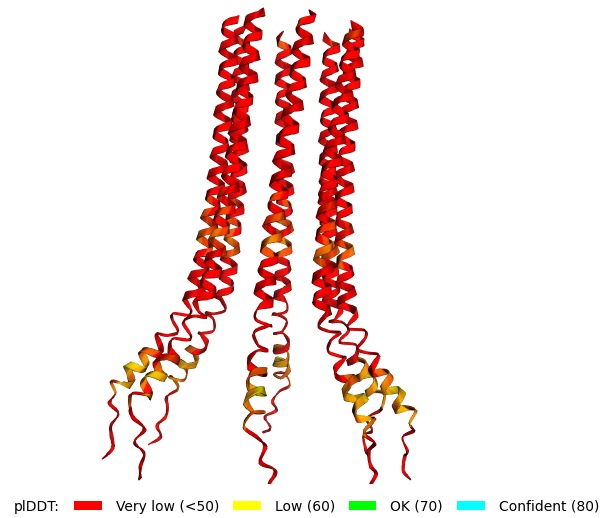 Mutant 2 pLDDT=45.8, pTM-0.187, ipTM = 0.126. 3 pairs/bridges introduced, 1 conserved site changed (13P->L), 2nd pair inverted, no RRR site (1 conserved and 5 variable sites changed) |
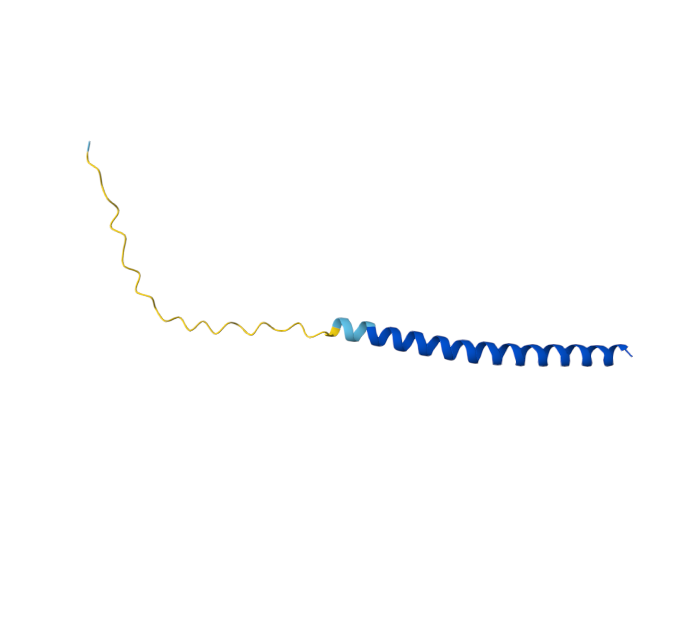 Original Sequence AlphaFold ipTM = -pTM = 0.44 | 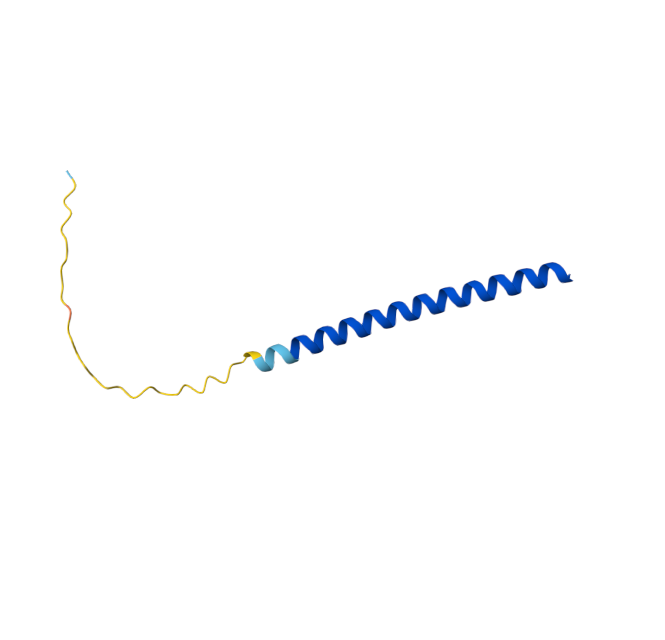 Mutant 1 AlphaFold ipTM = -pTM = 0.43 |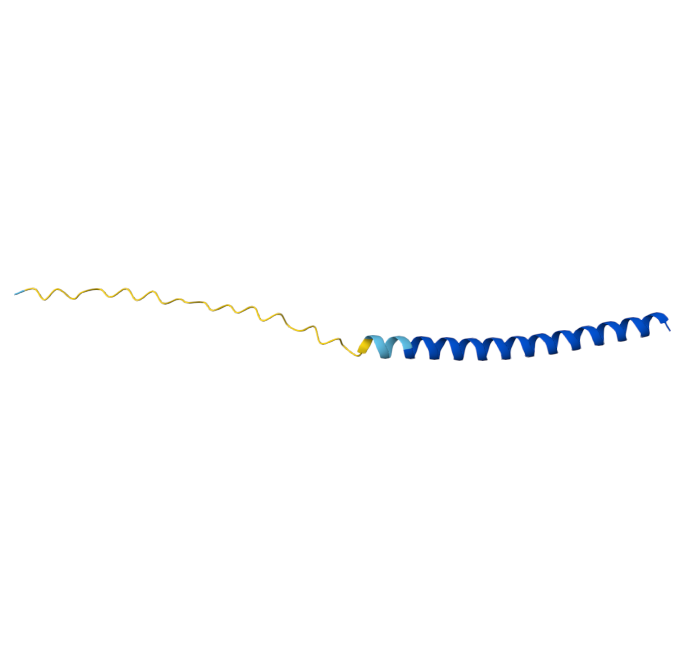  Mutant 2 AlphaFold ipTM = - , pTM = 0.44 |
Mutated Sequence 3
METRFPRQSQETLESTNRRRPRKHEDYPCRRQQRSST LYVLIFLAIFLSKFTNQLLLSLLEAVIRTVTTLQQLLT
This sequence was designed to explore whether changing the conserved site (13P->L) was required to achieve the same structure as that of the Mutated Sequence 1. For that, the mutated conserved site of the Sequence 1 was changed back to the original (13L->P).
Summary of mutations
Conserved site changed: None
Variable sites changed (as in Mutated Sequence 1): (7Q->R, 11Q->E, 14A->E, 22F->R)
Pairs introduced by changing the 4 variable sites (as in Mutated Sequence 1): Pair 1 (R7–E11), Pair 2 (E14–R18), Pair 3 (R22–D26)
Soluble N-terminal domain C-terminal domain
METRFPQQSQQTPASTNRRRPFKHEDYPCRRQQRSST LYVLIFLAIFLS KFTNQLLLSLLEAVIRTVTTLQQLLT (Original Sequence)
R---E LE--- R--- (Mutated Sites)
V V CV V (Conserved / Variable)
METRFPRQSQETLESTNRRRPRKHEDYPCRRQQRSST LYVLIFLAIFLS KFTNQLLLSLLEAVIRTVTTLQQLLT (Mutated Sequence 1)
L (Reverted site)
C (Conserved / Variable)
METRFPRQSQETLESTNRRRPRKHEDYPCRRQQRSST LYVLIFLAIFLS KFTNQLLLSLLEAVIRTVTTLQQLLT (Mutated Sequence 3)
Mutated Sequence 4
METRFPRQSQETLESTNRRRPRKHEDYPCRRQQRSST LYVLIFLAIFLSKFTNQLLLSLLEAVIRTVTTLQQLLT
This sequence was designed to explore whether changing the conserved site (13P->L) was required to achieve the helix as in the Mutated Sequence 2. For that, the mutated conserved site of the Sequence 2 was changed back to the original (13L->P).
Summary of mutations
Conserved site changed: None
Variable sites changed (as in Mutated Sequence 1): (7Q->R, 11Q->E, 14A->E, 22F->R)
Pairs introduced by changing the 4 variable sites (as in Mutated Sequence 2): Pair 1 (R7–E11), Pair 2 (E14–R18), Pair 3 (R22–D26)
Soluble N-terminal domain C-terminal domain
METRFPQQSQQTPASTNRRRPFKHEDYPCRRQQRSST LYVLIFLAIFLS KFTNQLLLSLLEAVIRTVTTLQQLLT (Original Sequence)
R---E LR---E R--- (Mutated Sites)
V V CV V V (Conserved / Variable)
METRFPRQSQETLRSTNERRPRKHEDYPCRRQQRSST LYVLIFLAIFLS KFTNQLLLSLLEAVIRTVTTLQQLLT (Mutated Sequence 2)
L (Reverted site)
C (Conserved / Variable)
METRFPRQSQETLRSTNERRPRKHEDYPCRRQQRSST LYVLIFLAIFLS KFTNQLLLSLLEAVIRTVTTLQQLLT (Mutated Sequence 4)
alfafold2_multimer_v2 was used to fold the multimers of Mutated Sequence 3 and Mutated Sequence 4. alfafold2_multimer_v2 parameters used:
num_relax: 0
template_mode: none
msa_mode: mmseqs2_uniref_env
Pair mode: paired
num_recycles: 3
recycle_early_stop_tolerance: auto
relax_max_iterations: 200
pairing_strategy: greedy
max_msa: auto
num_seeds: 1
 Mutant 1 pLDDT=37.6, pTM-0.189, ipTM = 0.127. 3 pairs/bridges introduced, 1 conserved site changed (13P->L), RRR site kept, (1 conserved and 4 variable sites changed) | 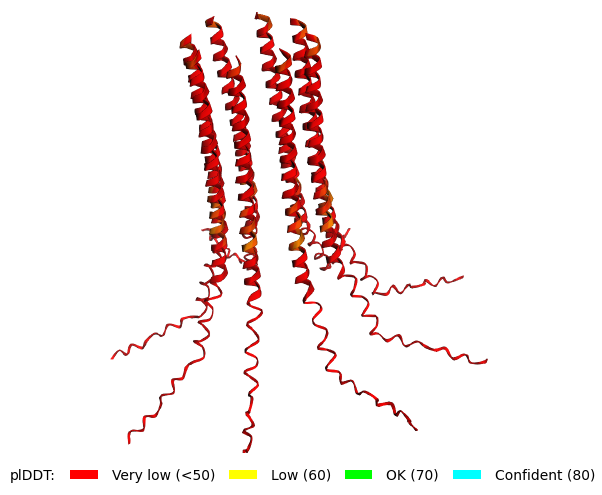 Mutant 3 pLDDT=43.3, pTM-0.188, ipTM = 0.127. Mutant 1 -> the conserved site mutation reverted (13L->P) (4 variable sites of the Original Sequence changed) |
 Mutant 2 pLDDT=45.8, pTM-0.187, ipTM = 0.126. 3 pairs/bridges introduced, 1 conserved site changed (13P->L), 2nd pair inverted, no RRR site (1 conserved and 5 variable sites changed) | 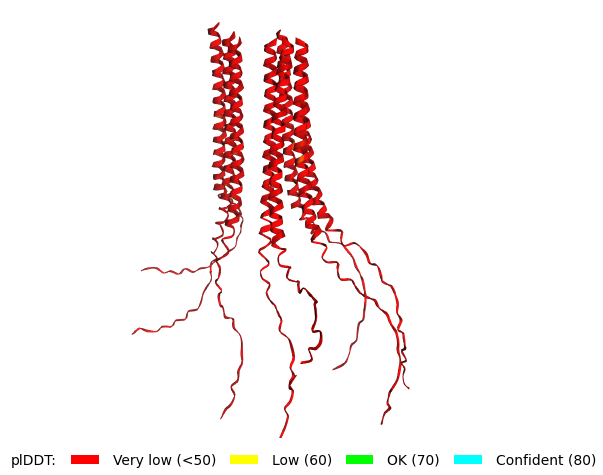 Mutant 4 pLDDT=37, pTM-0.189, ipTM = 0.127. Mutant 2 -> the conserved site mutation reverted (13L->P) (5 variable sites of the Original Sequence changed) |