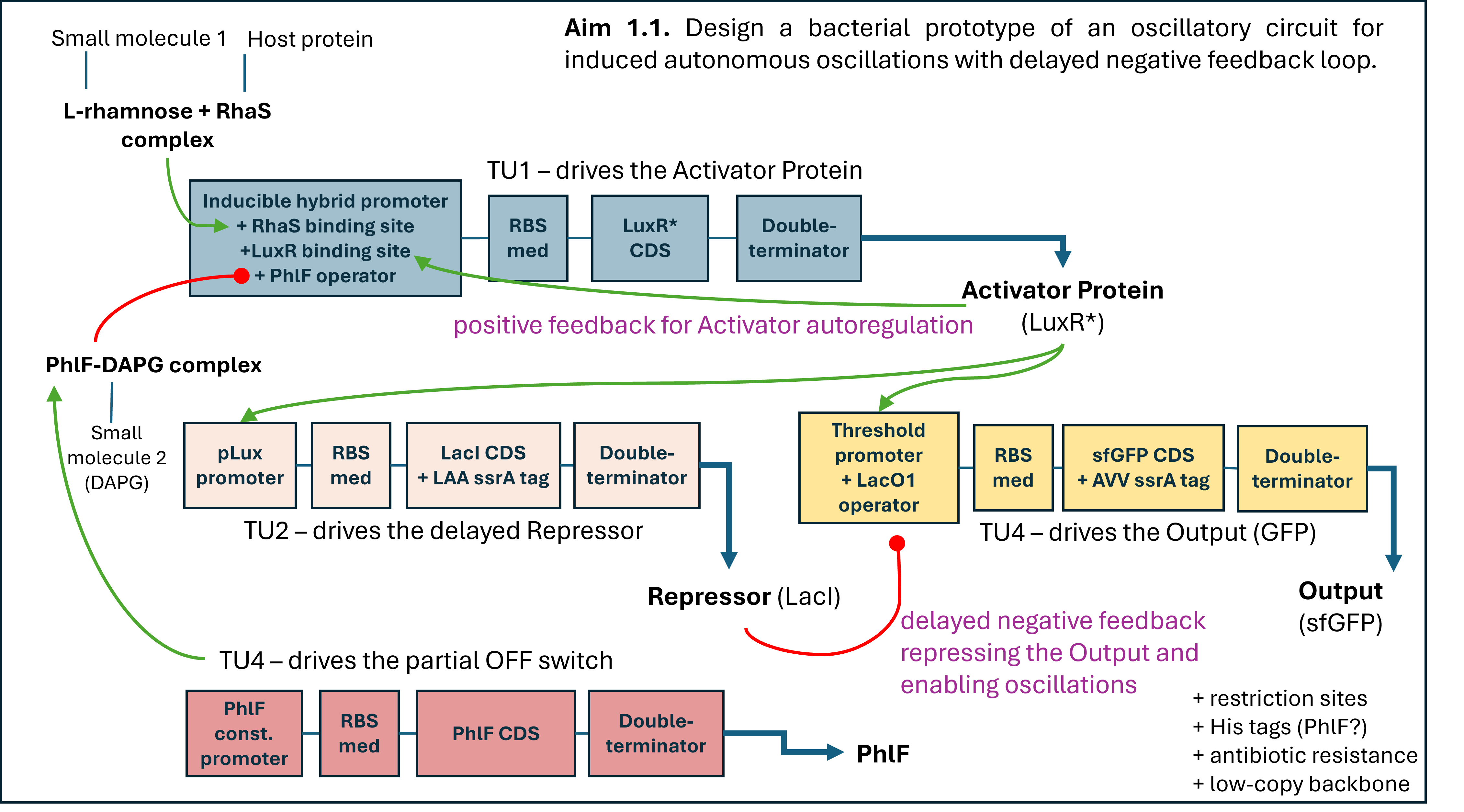
Week 3 HW: Lab Automation
Part 1: Opentrons Artwork This design was generated using the GUI at opentrons-art.rcdonovan.com and can be accessed through https://opentrons-art.rcdonovan.com/?id=1s7h4g7m1kn174o
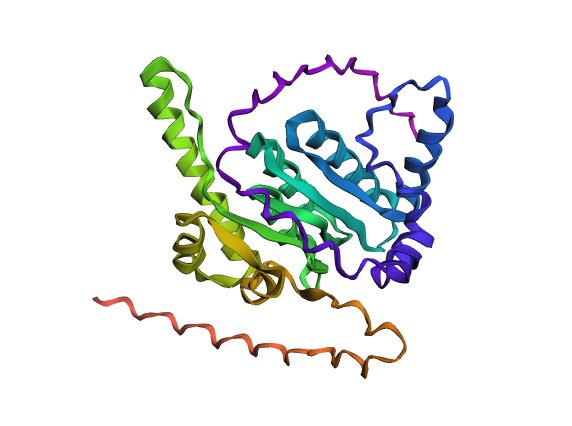
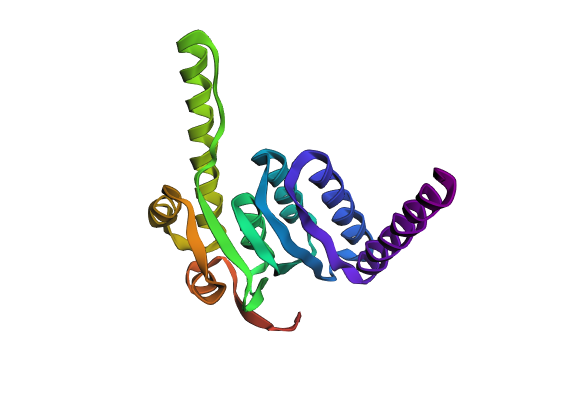
Coordinates mrfp1_points = [(27.5, 25.3),(25.3, 23.1),(23.1, 18.7),(20.9, 16.5),(18.7, 14.3),(36.3, 5.5),(38.5, 5.5),(27.5, 3.3),(29.7, 3.3),(31.9, 3.3),(34.1, 3.3),(36.3, 3.3),(20.9, 1.1),(23.1, 1.1),(25.3, 1.1),(16.5, -1.1),(18.7, -1.1)] mscarlet_i_points = [(29.7, 25.3),(27.5, 23.1),(29.7, 23.1),(25.3, 20.9),(27.5, 20.9),(25.3, 18.7),(23.1, 16.5),(20.9, 14.3),(16.5, 12.1),(18.7, 12.1),(16.5, 9.9),(14.3, 7.7),(38.5, 7.7),(12.1, 5.5),(34.1, 5.5),(9.9, 3.3),(38.5, 3.3),(14.3, -1.1)] electra2_points = [(31.9, 23.1),(29.7, 20.9),(31.9, 20.9),(34.1, 20.9),(27.5, 18.7),(29.7, 18.7),(31.9, 18.7),(25.3, 16.5),(27.5, 16.5),(29.7, 16.5),(23.1, 14.3),(25.3, 14.3),(27.5, 14.3),(20.9, 12.1),(23.1, 12.1),(18.7, 9.9),(20.9, 9.9),(16.5, 7.7),(18.7, 7.7),(14.3, 5.5),(16.5, 5.5),(12.1, 3.3),(9.9, 1.1),(9.9, -1.1)] mturquoise2_points = [(34.1, 18.7),(-5.5, 16.5),(31.9, 16.5),(34.1, 16.5),(36.3, 16.5),(29.7, 14.3),(31.9, 14.3),(34.1, 14.3),(25.3, 12.1),(27.5, 12.1),(29.7, 12.1),(23.1, 9.9),(25.3, 9.9),(27.5, 9.9),(20.9, 7.7),(23.1, 7.7),(1.1, 5.5),(18.7, 5.5),(14.3, 3.3),(16.5, 3.3),(12.1, 1.1),(23.1, -14.3),(7.7, -16.5),(9.9, -16.5),(12.1, -16.5),(14.3, -16.5),(-7.7, -18.7),(-5.5, -18.7),(-3.3, -18.7),(-1.1, -18.7),(1.1, -18.7),(3.3, -18.7),(5.5, -18.7),(7.7, -18.7),(9.9, -18.7),(12.1, -18.7),(16.5, -18.7),(-9.9, -20.9),(-7.7, -20.9),(-5.5, -20.9),(-3.3, -20.9),(-1.1, -20.9),(1.1, -20.9),(3.3, -20.9),(5.5, -20.9),(7.7, -20.9),(9.9, -20.9),(12.1, -20.9),(14.3, -20.9),(18.7, -20.9),(-12.1, -23.1),(-9.9, -23.1),(-7.7, -23.1),(-5.5, -23.1),(-3.3, -23.1),(-1.1, -23.1),(1.1, -23.1),(3.3, -23.1),(5.5, -23.1),(7.7, -23.1),(9.9, -23.1),(12.1, -23.1),(14.3, -23.1),(16.5, -23.1),(-14.3, -25.3),(-12.1, -25.3),(-9.9, -25.3),(-7.7, -25.3),(-5.5, -25.3),(-3.3, -25.3),(-1.1, -25.3),(1.1, -25.3),(3.3, -25.3),(5.5, -25.3),(-16.5, -27.5),(-14.3, -27.5),(-12.1, -27.5),(-9.9, -27.5)] azurite_points = [(-5.5, 14.3),(-3.3, 14.3),(-5.5, 12.1),(-1.1, 12.1),(-5.5, 9.9),(-3.3, 9.9),(-1.1, 9.9),(1.1, 9.9),(-7.7, 7.7),(-5.5, 7.7),(-3.3, 7.7),(-1.1, 7.7),(3.3, 7.7),(-7.7, 5.5),(-5.5, 5.5),(-3.3, 5.5),(-1.1, 5.5),(3.3, 5.5),(5.5, 5.5),(-7.7, 3.3),(-5.5, 3.3),(-3.3, 3.3),(-1.1, 3.3),(1.1, 3.3),(5.5, 3.3),(7.7, 3.3),(-7.7, 1.1),(-5.5, 1.1),(-3.3, 1.1),(-1.1, 1.1),(1.1, 1.1),(5.5, 1.1),(7.7, 1.1),(-9.9, -1.1),(-7.7, -1.1),(-5.5, -1.1),(-3.3, -1.1),(-1.1, -1.1),(1.1, -1.1),(3.3, -1.1),(5.5, -1.1),(7.7, -1.1),(-9.9, -3.3),(-7.7, -3.3),(-5.5, -3.3),(-3.3, -3.3),(-1.1, -3.3),(1.1, -3.3),(3.3, -3.3),(5.5, -3.3),(7.7, -3.3),(9.9, -3.3),(12.1, -3.3),(14.3, -3.3),(9.9, -5.5),(12.1, -5.5),(14.3, -5.5),(16.5, -5.5),(-12.1, -7.7),(-9.9, -7.7),(-7.7, -7.7),(-5.5, -7.7),(-3.3, -7.7),(-1.1, -7.7),(1.1, -7.7),(3.3, -7.7),(5.5, -7.7),(7.7, -7.7),(9.9, -7.7),(12.1, -7.7),(14.3, -7.7),(16.5, -7.7),(18.7, -7.7),(-12.1, -9.9),(-9.9, -9.9),(-7.7, -9.9),(-5.5, -9.9),(-3.3, -9.9),(-1.1, -9.9),(1.1, -9.9),(3.3, -9.9),(5.5, -9.9),(7.7, -9.9),(9.9, -9.9),(14.3, -9.9),(16.5, -9.9),(18.7, -9.9),(20.9, -9.9),(-12.1, -12.1),(-9.9, -12.1),(-7.7, -12.1),(-5.5, -12.1),(-3.3, -12.1),(-1.1, -12.1),(1.1, -12.1),(3.3, -12.1),(5.5, -12.1),(7.7, -12.1),(9.9, -12.1),(12.1, -12.1),(14.3, -12.1),(16.5, -12.1),(18.7, -12.1),(20.9, -12.1),(23.1, -12.1),(-12.1, -14.3),(-9.9, -14.3),(-7.7, -14.3),(-5.5, -14.3),(-3.3, -14.3),(-1.1, -14.3),(1.1, -14.3),(3.3, -14.3),(5.5, -14.3),(7.7, -14.3),(9.9, -14.3),(12.1, -14.3),(16.5, -14.3),(18.7, -14.3),(20.9, -14.3),(-12.1, -16.5),(-9.9, -16.5),(-7.7, -16.5),(-5.5, -16.5),(-3.3, -16.5),(-1.1, -16.5),(1.1, -16.5),(3.3, -16.5),(5.5, -16.5),(16.5, -16.5),(18.7, -16.5),(20.9, -16.5),(-12.1, -18.7),(-9.9, -18.7),(14.3, -18.7),(18.7, -18.7),(-14.3, -20.9),(-12.1, -20.9),(16.5, -20.9),(-14.3, -23.1),(-16.5, -25.3)] sfgfp_points = [(36.3, 14.3),(31.9, 12.1),(34.1, 12.1),(36.3, 12.1),(29.7, 9.9),(31.9, 9.9),(25.3, 7.7),(27.5, 7.7),(20.9, 5.5),(23.1, 5.5),(18.7, 3.3),(14.3, 1.1)] venus_points = [(34.1, 9.9),(36.3, 9.9),(29.7, 7.7),(25.3, 5.5),(20.9, 3.3),(16.5, 1.1)] mko2_points = [(-36.3, 12.1),(-38.5, 9.9),(-36.3, 9.9),(-34.1, 9.9),(38.5, 9.9),(-38.5, 7.7),(-36.3, 7.7),(-34.1, 7.7),(-31.9, 7.7),(31.9, 7.7),(34.1, 7.7),(36.3, 7.7),(-34.1, 5.5),(-31.9, 5.5),(-29.7, 5.5),(27.5, 5.5),(29.7, 5.5),(31.9, 5.5),(-29.7, 3.3),(-27.5, 3.3),(-25.3, 3.3),(23.1, 3.3),(25.3, 3.3),(-25.3, 1.1),(-23.1, 1.1),(-20.9, 1.1),(18.7, 1.1),(-20.9, -1.1),(-18.7, -1.1),(-16.5, -1.1),(12.1, -1.1),(-14.3, -3.3)] mjuniper_points = [(-3.3, 12.1),(1.1, 7.7),(3.3, 3.3),(3.3, 1.1),(-9.9, -5.5),(-7.7, -5.5),(-5.5, -5.5),(-3.3, -5.5),(-1.1, -5.5),(1.1, -5.5),(3.3, -5.5),(5.5, -5.5),(7.7, -5.5),(12.1, -9.9),(14.3, -14.3)] Part 2: Post-Lab Questions Part 3: Final Project Ideas Project 1: Tunable Induction of Alpha-Synuclein Expression for Modeling Parkinson’s Disease Aim:
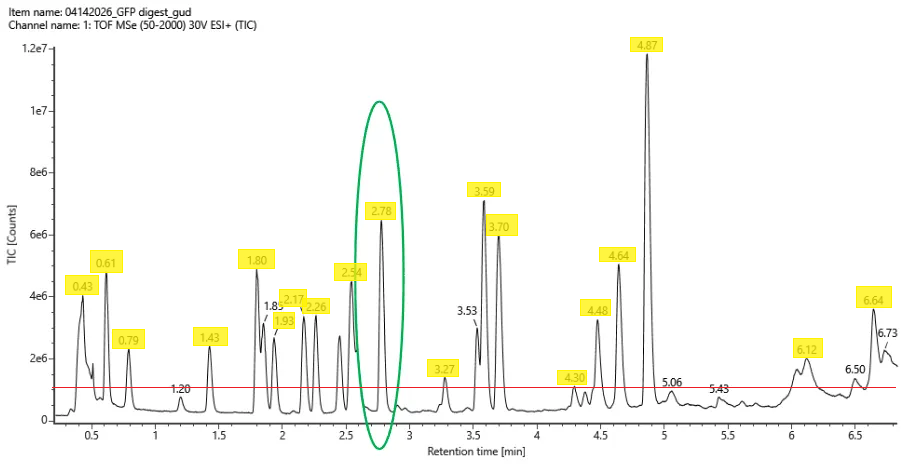
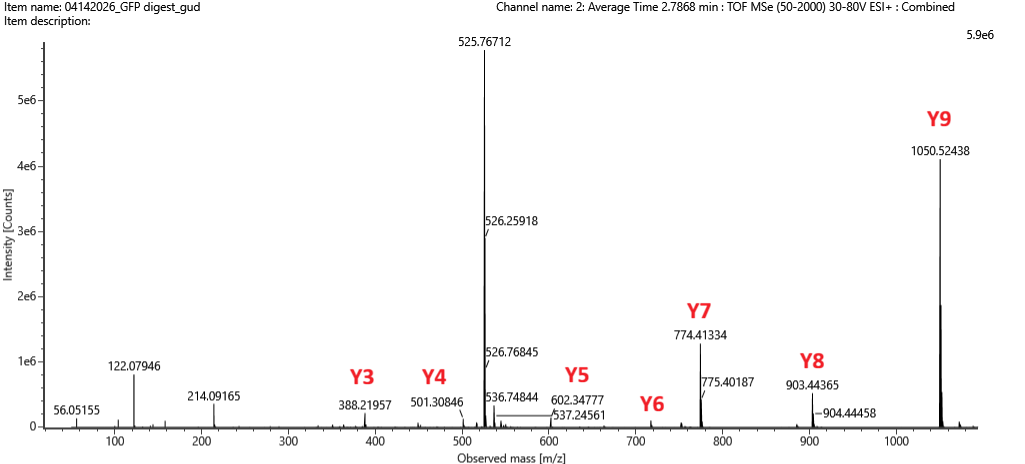
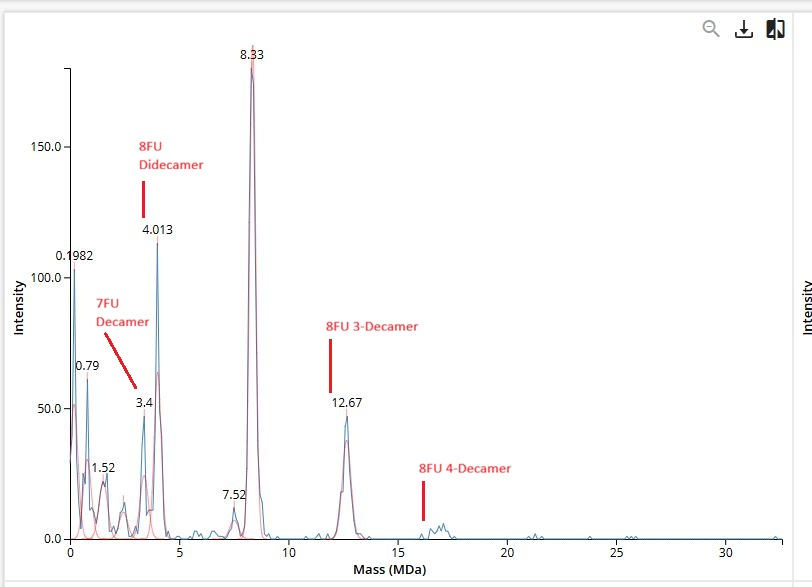
.png)